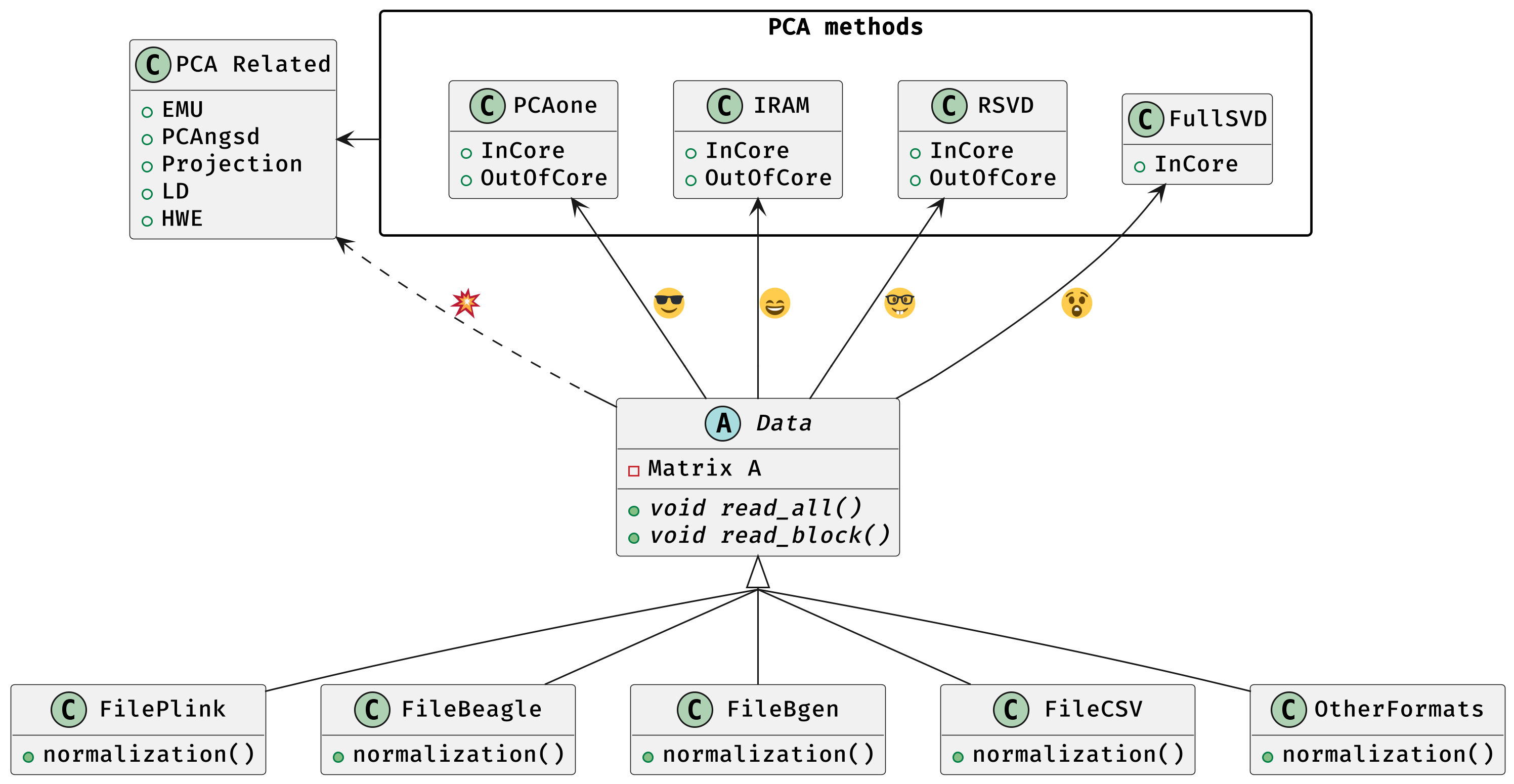
PCAone is a fast and memory efficient PCA tool implemented in C++ aiming at providing comprehensive features and algorithms for different scenarios. PCAone implements 3 fast PCA algorithms for finding the top eigenvectors of large datasets, which are Implicitly Restarted Arnoldi Method (IRAM, –svd 0), single pass Randomized SVD but with power iteration scheme (RSVD, –svd 1, Algorithm1 in paper) and our own RSVD method with window based power iteration scheme (PCAone, –svd 2, Algorithm2 in paper). All have both in-core and out-of-core implementation. Addtionally, full SVD (–svd 3) is supported via in-core mode only. There is also an R package here. PCAone supports multiple different input formats, such as PLINK, BGEN, Beagle genetic data formats and a general comma separated CSV format for other data, such as scRNAs and bulk RNAs. For genetics data, PCAone also implements EMU and PCAngsd algorithm for data with missingness and uncertainty.
- If you find PCAone helpful, please cite our paper on genome reseach Fast and accurate out-of-core PCA framework for large scale biobank data.
- If using the EMU algorithm, please also cite Large-scale inference of population structure in presence of missingness using PCA.
- If using the PCAngsd algorithm, please also cite Inferring Population Structure and Admixture Proportions in Low-Depth NGS Data.
- If using the LD pruning and clumping algorithm, please also cite Measuring linkage disequilibrium and improvement of pruning and clumping in structured populations.
See change log here.
- Has both Implicitly Restarted Arnoldi Method (IRAM) and Randomized SVD (RSVD) with out-of-core implementation.
- Implements our new fast window based Randomized SVD algorithm for tera-scale dataset.
- Quite fast with multi-threading support using high performance library MKL or OpenBLAS as backend.
- Supports the PLINK, BGEN, Beagle genetic data formats.
- Supports a general comma separated CSV format for single cell RNA-seq or bulk RNA-seq data compressed by zstd.
- Supports EMU algorithm for scenario with large proportion of missingness.
- Supports PCAngsd algorithm for low coverage sequencing scenario with genotype likelihood as input.
- Novel LD prunning and clumping method for admixed population.
We can run the following on Linux to have a quick start.
pkg=https://github.com/Zilong-Li/PCAone/releases/latest/download/PCAone-avx2-Linux.zip
wget $pkg
unzip -o PCAone-avx2-Linux.zip
wget http://popgen.dk/zilong/datahub/pca/example.tar.gz
tar -xzf example.tar.gz && rm -f example.tar.gz
# in default calculating top 10 PCs with in-core mode if having enough RAM
./PCAone --bfile example/plink -k 10
R -s -e 'df=read.table("pcaone.eigvecs", h=F);plot(df[,1:2], xlab="PC1", ylab="PC2");'We will find those files in your current directory.
.
├── PCAone # program
├── Rplots.pdf # pca plot
├── example # folder of example data
├── pcaone.eigvals # eigenvalues
├── pcaone.eigvecs # eigenvectors, the PCs you need to plot
├── pcaone.eigvecs2 # eigenvectors with header line
└── pcaone.log # log fileOn Linux platform,
pkg=https://github.com/Zilong-Li/PCAone/releases/latest/download/PCAone-avx2-Linux.zip
wget $pkg || curl -LO $pkg
unzip -o PCAone-avx2-Linux.zipIf the server is too old to support avx2 instruction, one can download the following version.
pkg=https://github.com/Zilong-Li/PCAone/releases/latest/download/PCAone-x64-Linux.zip
wget $pkg || curl -LO $pkg
unzip -o PCAone-x64-Linux.zippkg=http://popgen.dk/zilong/datahub/pca/example.tar.gz
wget $pkg || curl -LO $pkg
tar -xzf example.tar.gz && rm -f example.tar.gzWe should find a fold named example with some example data.
There are 3 ways to install PCAone.
There are compiled binaries provided for both Linux and Mac platform. Check the releases page to download one.
PCAone is also available from bioconda.
conda config --add channels bioconda
conda install pcaone
PCAone --helpPCAone can be running on a normal computer/laptop with x86-64 instruction set architecture. PCAone has been tested on both Linux and MacOS system. To build PCAone from the source code, the following dependencies are required:
- GCC/Clang compiler with C++11 support
- GNU make
- zlib
We recommend building the software from source with MKL as backend to maximize the performance. For MacOS users, we recommend using llvm by brew install llvm instead of the default clang shipped with MacOS. Check out the mac workflow.
Build PCAone dynamically with MKL can maximize the performance since the faster threading layer libiomp5 will be linked at runtime. One can obtain the MKL by one of the following option:
- install
mklby conda
conda install -c conda-forge -c anaconda -y mkl mkl-include intel-openmp
git clone https://github.com/Zilong-Li/PCAone.git
cd PCAone
# if mkl is installed by conda then use ${CONDA_PREFIX} as mklroot
make -j4 MKLROOT=${CONDA_PREFIX}
./PCAone -h- download
mklfrom the website
After having mkl installed, find the mkl root path and replace the path below with your own.
# if libiomp5 is not in the mklroot path, please link it to $MKLROOT/lib folder
make -j4 MKLROOT=/path/to/mklrootAlternatively, for advanced user, modify variables directly in Makefile and run make to use MKL or OpenBlas as backend.
If you don’t want any optimized math library as backend, just run:
git clone https://github.com/Zilong-Li/PCAone.git
cd PCAone
make -j4
./PCAone -hIf this doesn’t work because the server is too outdated, run make clean && make -j4 AVX=0 instead.
\newpage
run ./PCAone --help to see all options. Below are some useful and important options.
Main options:
-h, --help print all options including hidden advanced options
-d, --svd arg (=2) svd method to be applied. default 2 is recommended for big data.
0: the Implicitly Restarted Arnoldi Method (IRAM)
1: the Yu's single-pass Randomized SVD with power iterations
2: the proposed window-based Randomized SVD method
3: the full Singular Value Decomposition.
-b, --bfile arg prefix to PLINK .bed/.bim/.fam files
-B, --binary arg path of binary file (experimental and in-core mode)
-c, --csv arg path of comma seperated CSV file compressed by zstd
-g, --bgen arg path of BGEN file compressed by gzip/zstd
-G, --beagle arg path of BEAGLE file compressed by gzip
-k, --pc arg (=10) top k eigenvalues (PCs) to be calculated
-m, --memory arg (=0) desired RAM usage in GB unit. default [0] uses all RAM
-n, --threads arg (=10) number of threads for multithreading
-o, --out arg (=pcaone) prefix to output files. default [pcaone]
-p, --maxp arg (=40) maximum number of power iterations for RSVD algorithm
-S, --no-shuffle do not shuffle the data if it is already permuted
-v, --verbose verbose message output
-w, --batches arg (=64) number of mini-batches to be used by PCAone --svd 2
-C, --scale arg (=0) do scaling for input file.
0: do just centering
1: do log transformation eg. log(x+0.01) for RNA-seq data
2: do count per median log transformation (CPMED) for scRNAs
--emu uses EMU algorithm for genotype input with missingness
--pcangsd uses PCAngsd algorithm for genotype likelihood input
--maf arg (=0) skip variants with minor allele frequency below maf
-V, --printv output the right eigenvectors with suffix .loadings
--ld output a binary matrix for LD related stuff
--ld-r2 arg (=0) cutoff for ld pruning. A value > 0 activates ld pruning
--ld-bp arg (=1000000) physical distance threshold in bases for ld pruning
--ld-stats arg (=0) statistics for calculating ld-r2. (0: the adj; 1: the std)
--clump arg assoc-like file with target variants and pvalues for clumping
--clump-names arg (=CHR,BP,P) olumn names in assoc-like file for locating chr, pos and pvalue respectively
--clump-p1 arg (=0.0001) significance threshold for index SNPs
--clump-p2 arg (=0.01) secondary significance threshold for clumped SNPs
--clump-r2 arg (=0.5) r2 cutoff for ld clumping
--clump-bp arg (=250000) physical distance threshold in bases for clumping
PCAone is designed to be extensible to accept many different formats. Currently, PCAone can work with SNP major genetic formats to study population structure. such as PLINK, BGEN and Beagle. Also, PCAone supports a comma delimited CSV format compressed by zstd, which is useful for other datasets requiring specific normalization such as single cell RNAs data.
Eigen vectors are saved in file with suffix .eigvecs. Each row represents a sample and each col represents a PC.
Eigen values are saved in file with suffix .eigvals. Each row represents the eigenvalue of corresponding PC.
Features Loadings are saved in file with suffix .loadings. Each row represents a feature and each col represents a PC. need to use --printv option to print it.
PCAone has both in-core and out-of-core mode for 3 different partial SVD algorithms, which are IRAM (--svd 0), Yu+Halko RSVD (--svd 1) and PCAone window-based RSVD (--svd 2). Also, PCAone supports full SVD (--svd 3) but with only in-core mode. Therefore, there are 7 ways in total for doing PCA in PCAone. In default PCAone uses in-core mode with --memory 0, which is the fastest way to do calculation. However, in case the server runs out of memory with in-core mode, the user can trigger out-of-core mode by specifying the amount of memory using --memory option with a value greater than 0.
./PCAone --bfile example/plink --svd 2./PCAone --bfile example/plink --svd 2 -m 2./PCAone --bfile example/plink --svd 1./PCAone --bfile example/plink --svd 1 -m 2./PCAone --bfile example/plink --svd 0 -m 2./PCAone --bfile example/plink --svd 0 -m 2./PCAone --bfile example/plink --svd 3PCAone will automatically apply the standard normalization for genetic data. Additionally, there are 3 different normalization method implemented with --scale option.
- 0: do just centering by substracting the mean
- 1: do log transformation (usually for count data, such as bulk RNA-seq data)
- 2: do count per median log transformation (usually for single cell RNA-seq data)
One should choose proper normalization method for specific type of data.
This is a novel statistics on LD calculation in admixed population. For more details, see our paper.
PCAone -b plink -k 3 --ld-stats 0 --ld-r2 0.8 --ld-bp 1000000If you already done LD prunning with PCAone, then you can find a binary file named .residuals, which will be used by LD clumping here.
# first output a LD matrix
PCAone -b plink -k 3 --ld
# do clumping given the LD matrix and user-defined association results
PCAone -B pcaone.residuals --clump plink.assoc --clump-p1 5e-8 --clump-p2 1e-6 --clump-r2 0.01 --clump-bp 10000000Let’s download the example data first.
wget http://popgen.dk/zilong/datahub/pca/example.tar.gz
tar -xzf example.tar.gz && rm -f example.tar.gzWe want to compute the top 10 PCs for this genotype dataset using 4 threads and only 2GB memory. We will use the proposed window-based RSVD algorithm with default setting --svd 2.
./PCAone --bfile example/plink -k 10 -n 4 -m 2Then, we can make a PCA plot in R.
pcs <- read.table("pcaone.eigvecs",h=F)
fam <- read.table("example/plink.fam",h=F)
pop <- fam[,1]
plot(pcs[,1:2], col=factor(pop), xlab = "PC1", ylab = "PC2")
legend("topright", legend=unique(pop), col=factor(unique(pop)), pch = 21, cex=1.2)Imputation tools usually generate the genotype probabilities or dosages in BGEN format. To do PCA with the imputed genotype probabilities, we can work on BGEN file with --bgen option instead.
./PCAone --bgen example/test.bgen -k 10 -n 4 -m 2Then, we can make a PCA plot in R.
pcs <- read.table("pcaone.eigvecs",h=F)
fam <- read.table("example/plink.fam",h=F)
pop <- fam[,1]
plot(pcs[,1:2], col=factor(pop), xlab = "PC1", ylab = "PC2")
legend("topright", legend=unique(pop), col=factor(unique(pop)), pch = 21, cex=1.2)In this example, we run PCA for the single cell RNAs-seq data using a different input format with a normalization method called count per median log transformation (CPMED).
./PCAone --csv example/BrainSpinalCord.csv.zst -k 10 -n 20 -m 4 --scale 2 --svd 1It should take around 5 minutes.
PCAone use Eigen for linear algebra operation. The IRAM method is based on yixuan/spectra. The bgen lib is ported from jeremymcrae/bgen. The EMU and PCAngsd algorithms are modified from @Jonas packages.