SCRIP
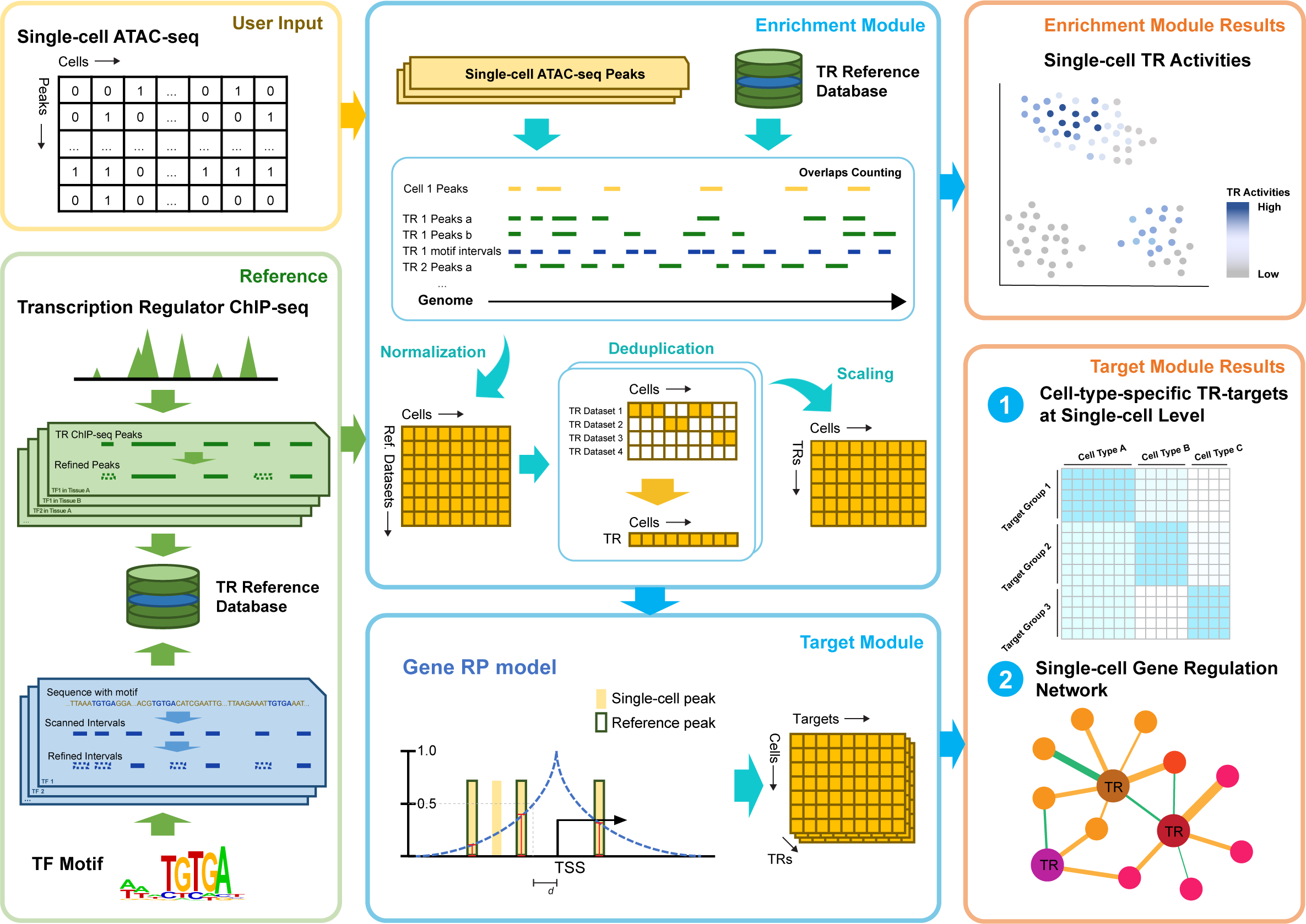
SCRIP (Single Cell Regulatory network Inference using ChIP-seq & motif) is a toolkit for elucidating the gene regulation pattern based on scATAC-seq leveraing a huge amount of bulk ChIP-seq data. It supports (1) evaluating the TR activities in single-cell based on the integration of the scATAC-seq dataset and curated reference; (2) determining the target genes of TR at single-cell resolution; (3) constructing the GRNs in single-cell and identifying cell-specific regulation.
Documentation
For the detailed usage and examples of SCRIP, please refer to the documentation.
For the analysis codes in the paper, please refer to the Notebook.
Installation
Dependency, please install them first:
libpng12-0 tabixInstall SCRIP
git clone git@github.com:wanglabtongji/SCRIP.git
cd SCRIP
python setup.py installThen, please download the reference files and config them with SCRIP config.
Usage
SCRIP all functions
usage: SCRIP [-h] [--version] {enrich,impute,target,config,index} ...
SCRIP
positional arguments:
{enrich,impute,target,config,index}
enrich Main function.
impute Imputation Factor function.
target Calculate targets based on factor peak count.
config Configuration.
index Build index with custom intervals.
optional arguments:
-h, --help show this help message and exit
--version show program's version number and exit
For command line options of each command, type: SCRIP COMMAND -
SCRIP enrich function
usage: SCRIP enrich [-h] -i FEATURE_MATRIX -s {hs,mm} [-p PROJECT] [--min_cells MIN_CELLS] [--min_peaks MIN_PEAKS] [--max_peaks MAX_PEAKS]
[-t N_CORES] [-m {max,mean}] [-y] [--clean]
optional arguments:
-h, --help show this help message and exit
Input files arguments:
-i FEATURE_MATRIX, --input_feature_matrix FEATURE_MATRIX
A cell by peak matrix . REQUIRED.
-s {hs,mm}, --species {hs,mm}
Species. "hs"(human) or "mm"(mouse). REQUIRED.
Output arguments:
-p PROJECT, --project PROJECT
Project name, which will be used to generate output files folder. DEFAULT: Random generate.
Preprocessing paramater arguments:
--min_cells MIN_CELLS
Minimal cell cutoff for features. Auto will take 0.05% of total cell number.DEFAULT: "auto".
--min_peaks MIN_PEAKS
Minimal peak cutoff for cells. Auto will take the mean-3*std of all feature number (if less than 500 is 500). DEFAULT: "auto".
--max_peaks MAX_PEAKS
Max peak cutoff for cells. This will help you to remove the doublet cells. Auto will take the mean+5*std of all feature
number. DEFAULT: "auto".
Other options:
-t N_CORES, --thread N_CORES
Number of cores use to run SCRIP. DEFAULT: 16.
-m {max,mean}, --mode {max,mean}
Deduplicate strategy. DEFAULT: max.
-y, --yes Whether ask for confirmation. DEFAULT: False.
--clean Whether delete tmp files(including bed and search results) generated by SCRIP. DEFAULT: False.
SCRIP target function
usage: SCRIP target [-h] -i FEATURE_MATRIX -s {hs,mm} [-o OUTPUT] [-d DECAY] [-m MODEL]
optional arguments:
-h, --help show this help message and exit
Input files arguments:
-i FEATURE_MATRIX, --input_feature_matrix FEATURE_MATRIX
A cell by peak matrix. h5 or h5ad supported. REQUIRED.
-s {hs,mm}, --species {hs,mm}
Species. "hs"(human) or "mm"(mouse). REQUIRED.
Output arguments:
-o OUTPUT, --output OUTPUT
output h5ad file. DEFAULT: RP.h5ad
Other options:
-d DECAY, --decay DECAY
Range to the effect of peaks. DEFAULT: auto.
-m MODEL, --model MODEL
RP model chosen. DEFAULT: simple.
SCRIP config function
usage: SCRIP config [-h] [--show] [--human_tf_index HUMAN_TF_INDEX] [--human_hm_index HUMAN_HM_INDEX] [--mouse_tf_index MOUSE_TF_INDEX]
[--mouse_hm_index MOUSE_HM_INDEX]
optional arguments:
-h, --help show this help message and exit
--show
--human_tf_index HUMAN_TF_INDEX
--human_hm_index HUMAN_HM_INDEX
--mouse_tf_index MOUSE_TF_INDEX
--mouse_hm_index MOUSE_HM_INDEX
SCRIP index function
usage: SCRIP index [-h] -i INPUT -o OUTPUT
optional arguments:
-h, --help show this help message and exit
-i INPUT, --input INPUT
Path to the folder that includes all your bed files. The bed files should be named in "TRName_ID.bed", e.g. "AR_1.bed".
-o OUTPUT, --output OUTPUT
Path to the output folder.