Structuring of Complex Longitudinal Data into Long Format
Authors: Romain Neugebauer, Noel Pimentel, and Nima Hejazi
The goal of LtAtStructuR is to structure a collection of time-stamped measurements (e.g., electronic health record data) into a standard long format analytic dataset suitable for the evaluation of the effect of multiple time-point interventions in the presence of time-dependent confounding or sources of selection bias.
Download the LtAtStructuR directory and install the package from the command line using
R CMD INSTALL LtAtStructuRor from within R using the package tarball (.tar.gz) with
install.packages("PATH/TO/LtAtStructuR_version.tar.gz", type = "local")where the path to the package tarball and version number must be amended. A tarball may be generated by invoking Rscript -e "devtools::build()" from the root directory of this project.
This is a basic example which shows you how to solve a common problem:
library(LtAtStructuR)
library(data.table)
library(lubridate)
library(future) # optional (for parallel processing)
plan(multiprocess) # optional (for parallel processing)
## Define one cohort dataset, one exposure dataset, and one or more covariate
## datasets
cohort <- setCohort(cohortDT, "ID", "IndexDate", "EOFDate", "EOFtype",
"AMI", c("ageEntry", "sex", "race", "A1c", "eGFR"),
list("ageEntry"=list("categorical"=FALSE,
"impute"=NA,
"impute_default_level"=NA),
"sex"=list("categorical"=TRUE,
"impute"=NA,
"impute_default_level"=NA),
"race"=list("categorical"=TRUE,
"impute"=NA,
"impute_default_level"=NA)) )
exposure <- setExposure(expDT, "ID", "startA", "endA")
covariate1 <- setCovariate(a1cDT, "sporadic", "ID", "A1cDate", "A1c",
categorical = FALSE)
covariate2 <- setCovariate(egfrDT, "sporadic", "ID", "eGFRDate", "eGFR",
categorical = TRUE)
## Gather each input dataset into a single object that specifies the content of
## the output dataset to be constructed
LtAt.specification <- cohort + exposure + covariate1 + covariate2
## Construct the output dataset
LtAt.data <- construct(LtAt.specification, time_unit = 15, first_exp_rule = 1,
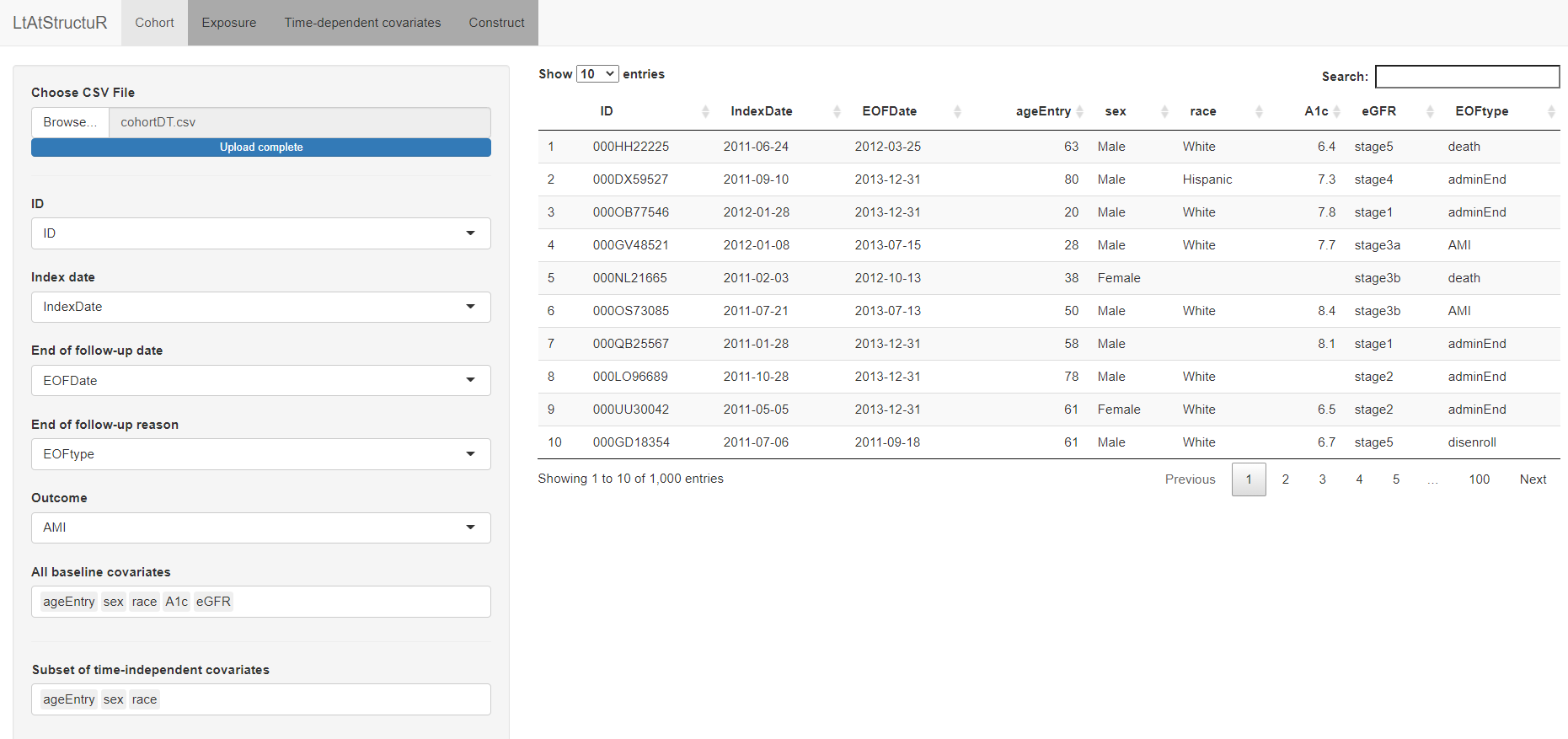
exp_threshold = 0.75)LtAtStructuR can also also be used through an interactive Shiny web application that automates the process of defining the cohort, exposure, and covariate datasets. The shiny application can be executed by running the command:
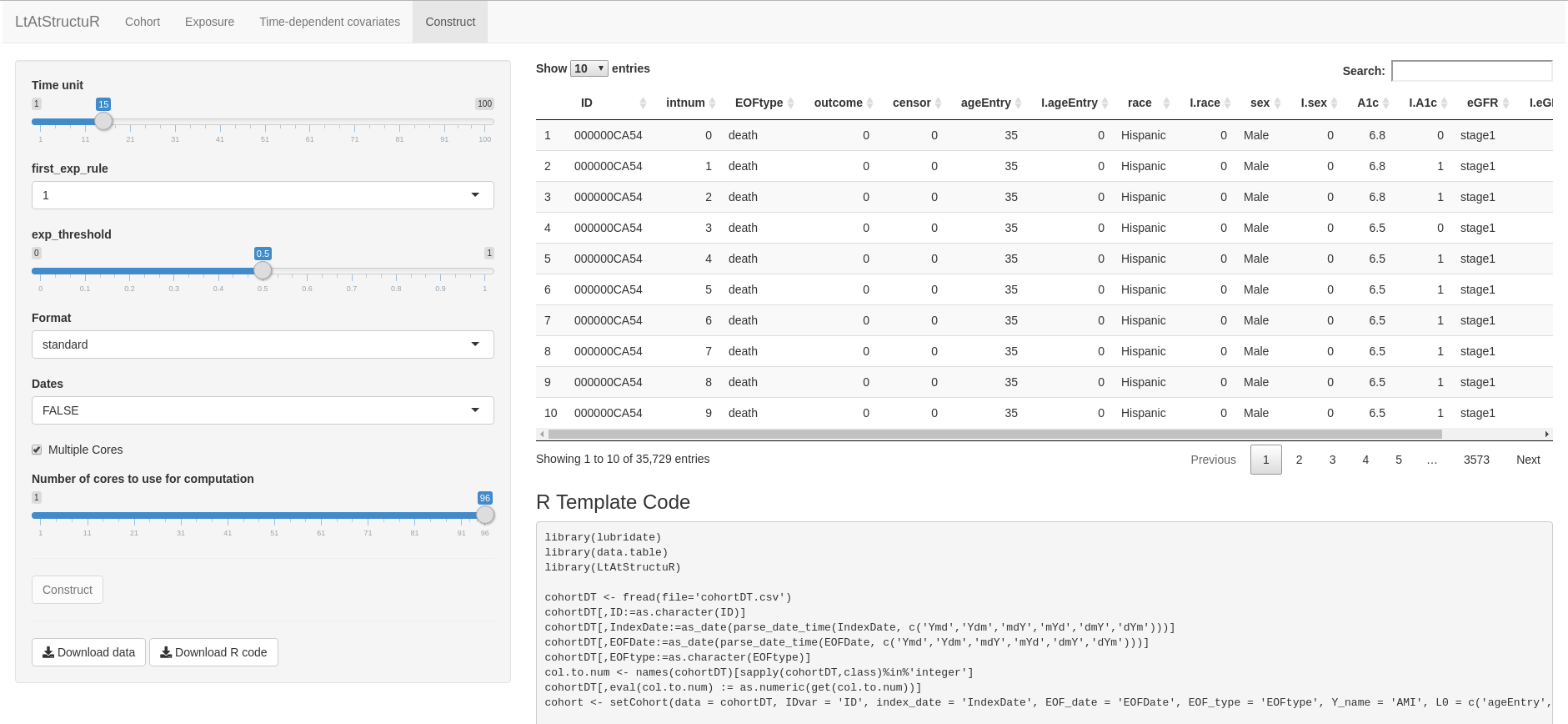
runShiny()The tabs Cohort, Exposure, Covariates, and Construct are analogous to the setCohort, setExposure, setCovariate, and constructfunctions, respectively; each tab offers an interactive user-interace to select input criteria pertaining to each of these functions. As a final step of the application, the user can download the output dataset and R template code specific to the construction of that dataset:
© 2019 Romain S. Neugebauer
The contents of this repository are distributed under the GPL-3 license. See file LICENSE for details.