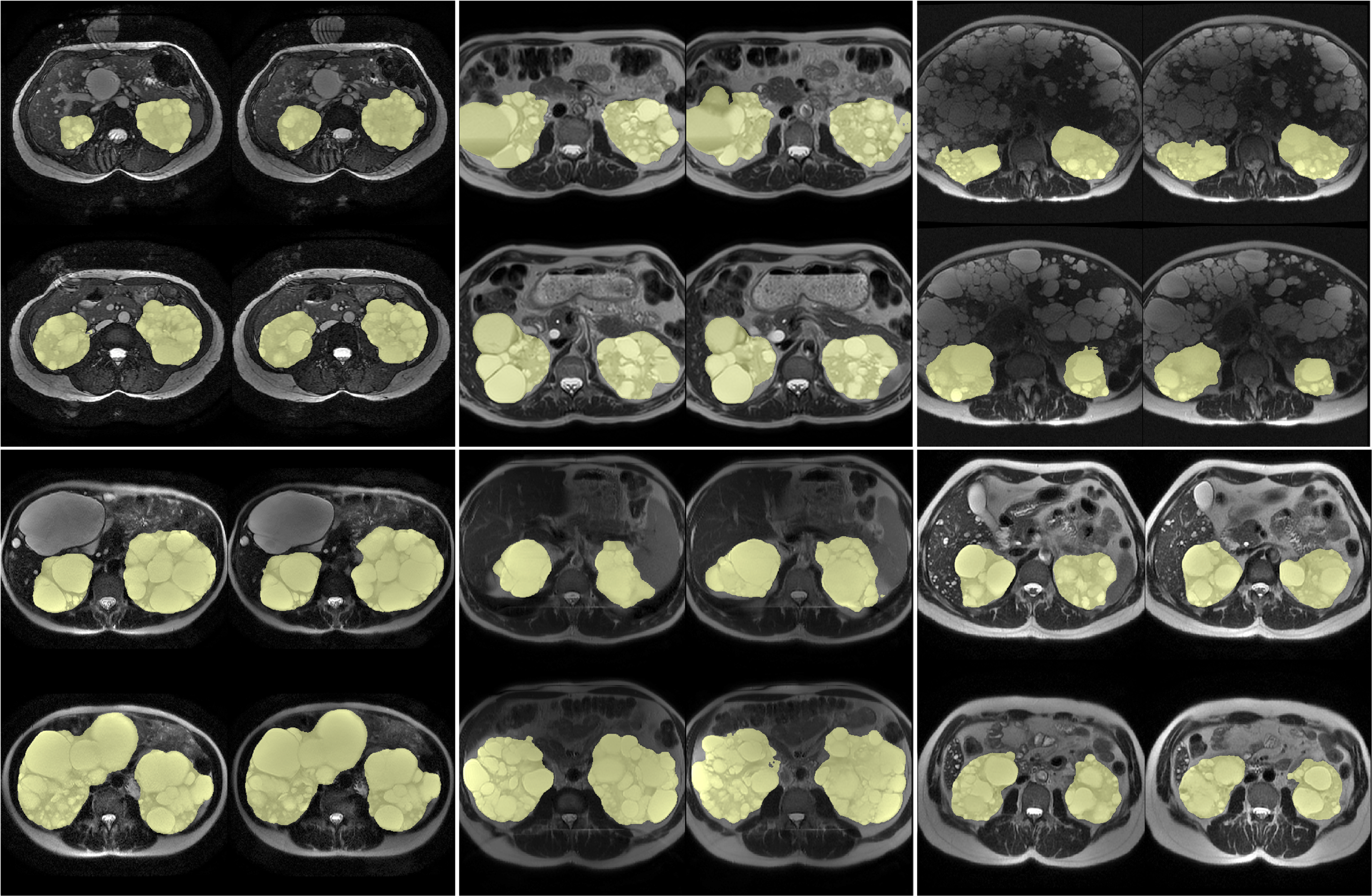
ADPKD-segmentation for determining Total Kidney Volume (TKV)
Autosomal dominant polycystic kidney disease (ADPKD) Segmentation in PyTorch
Project design, data management, and implementation for first version of polycystic kidney work by Akshay Goel, MD.
Follow-up work by researchers at Weill Cornell Medicine and Cornell University.
Published in Radiology: Artificial Intelligence (RSNA) in 2022
Goel A, Shih G, Riyahi S, Jeph S, Dev H, Hu R, et al. Deployed Deep Learning Kidney Segmentation for Polycystic Kidney Disease MRI. Radiology: Artificial Intelligence. p. e210205.
Published Online:Feb 16 2022 https://doi.org/10.1148/ryai.210205
Multiorgan Extention Published in Tomography 2022
Sharbatdaran A, Romano D, Teichman K, Dev H, Raza SI, Goel A, Moghadam MC, Blumenfeld JD, Chevalier JM, Shimonov D, Shih G, Wang Y, Prince MR. Deep Learning Automation of Kidney, Liver, and Spleen Segmentation for Organ Volume Measurements in Autosomal Dominant Polycystic Kidney Disease. Tomography. 2022; 8(4):1804-1819. https://doi.org/10.3390/tomography8040152. URL: https://www.mdpi.com/1723226
See additional README files for more info on:
- Training/Validation data
- Inference input data
- Saved inference output files
- Addition ensemble extension
- Argmax ensemble and multisequence extension
Preliminary Results Presented as Abstract at SIIM 2020
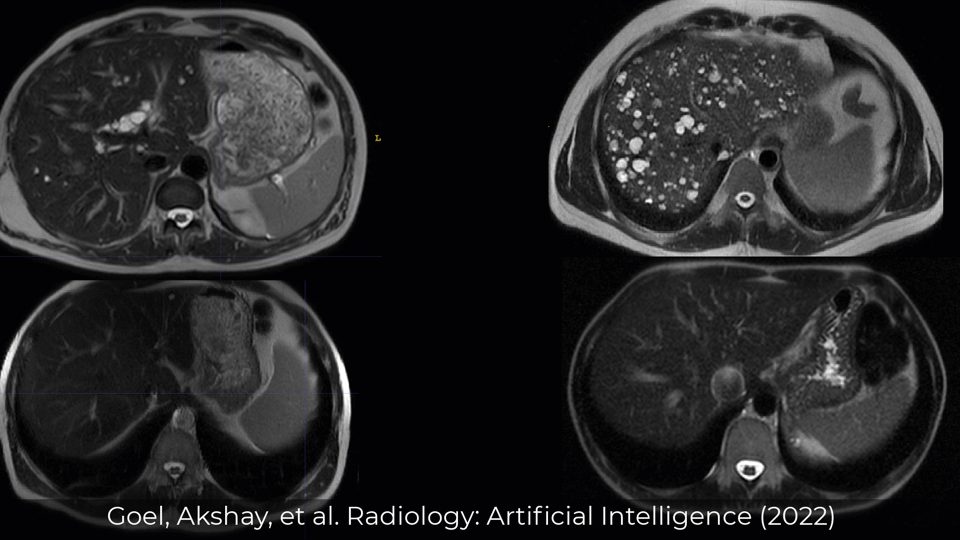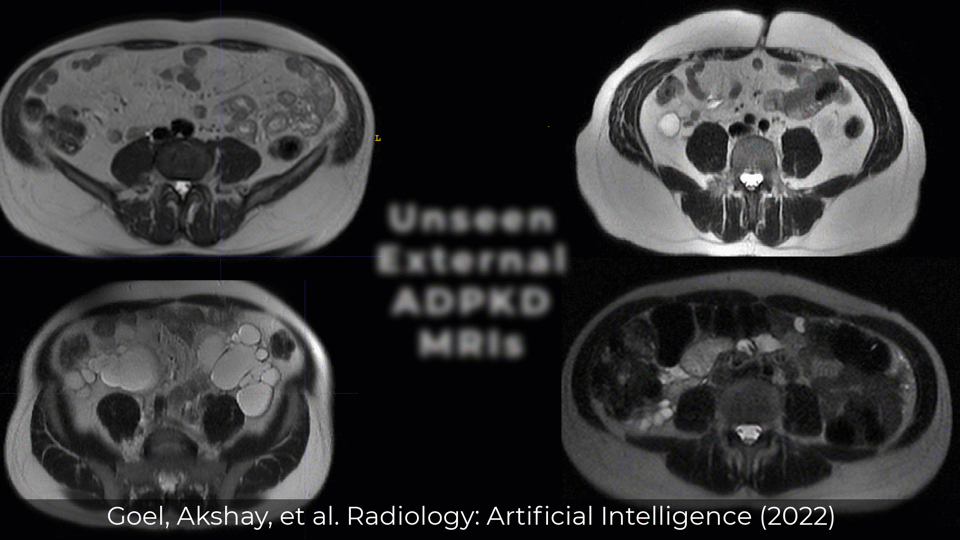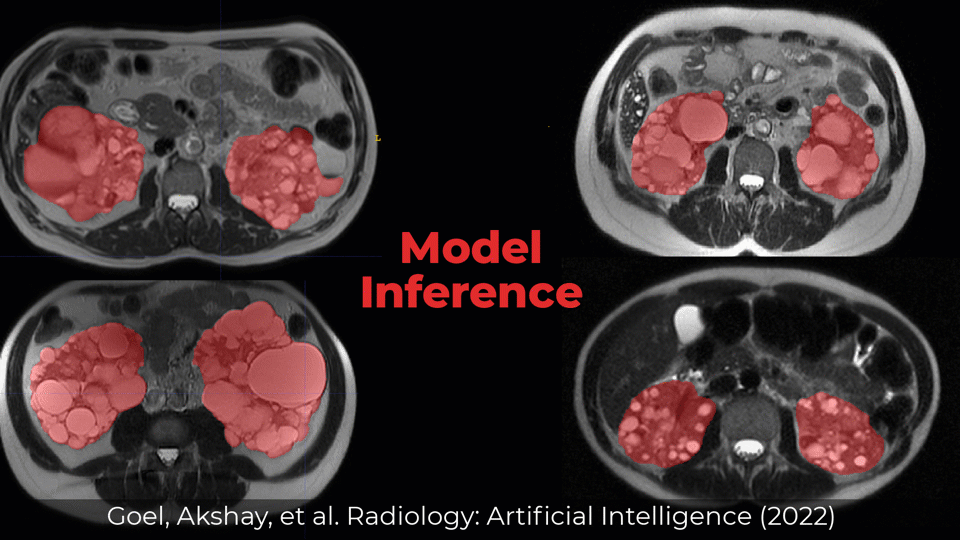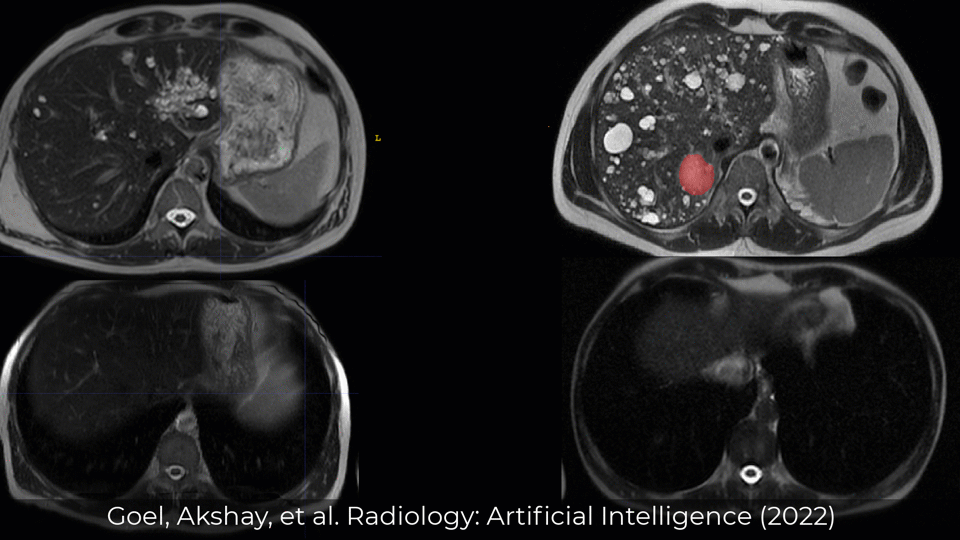
Examples of Performance on Unseen Multi-institute External Data
Inference was performed by checkpoints/inference.yml with checkpoint (checkpoints/best_val_checkpoint.pth)
Steps to run:
Sidenote on Configs and Experiments
- All experimental objects are defined by YAML configuration files.
- Configuration files are instantiated via config_utils.py.
- Select prior experiments can be reproduced via the coresponding config files (see experiments/configs).
- Tensboard runs and Model Checkpoints for these experiments are saved (see experiments/runs).
Inference
1. Install requirements.txt and adpkd-segmentation package from source.
pip install -e . -f https://download.pytorch.org/whl/torch_stable.html
A sidenote on installation
requirements.txtandadpkd-segmentationis supported forpython 3.8- Specifically, the pipeline has been tested on
python 3.8.4andpython 3.8.5 - For best results, we recommend installing all packages in a
python 3.8environment. - Once
python 3.8.4or3.8.5is installed, create a virtual environment withvirtualenv - You may want to reverence the virtual environment documentation:
- Basic Virtual Environment Tutorial
- Package installation for an environment) However, we provide a short example here:
(Windows)
working_dir>pip install virtualenv
working_dir>py -3.8.4 -m Drive:\path\to\environment_name\
Powershell Activation:
working_dir>Drive\path\to\environment_name\Scripts\activate.ps1
Windows Command Line:
working_dir>Drive\path\to\environment_name\Scripts\activate
working_dir>python setup.py install
(Unix/MacOS)
$pip install virtualenv
$python3.8.4 -m venv /path/to/environment_name
$source /path/to/environment_name/bin/activate
$python
2. Select an inference config file.
- To build the model for our best results use checkpoints/inference.yml which points to coresponding checkpoint checkpoints/best_val_checkpoint.pth
3. Run inference script:
$ python adpkd_segmentation/inference/inference.py -h
usage: inference.py [-h] [--config_path CONFIG_PATH] [-i INFERENCE_PATH] [-o OUTPUT_PATH]
optional arguments:
-h, --help show this help message and exit
--config_path CONFIG_PATH
path to config file for inference pipeline
-i INFERENCE_PATH, --inference_path INFERENCE_PATH
path to input dicom data (replaces path in config file)
-o OUTPUT_PATH, --output_path OUTPUT_PATH
path to output location
Training Pipeline
1. Install pip packages.
Install from requirements.txt (inside some virtual env):
pip install -e . -f https://download.pytorch.org/whl/torch_stable.html
2. Set up data as described here.
Note: Depending on the dataloader you may need to create a train / validation / test json file to indicate splits.
3. Select (or create) a config file. See examples at experiments/configs
4. Run training:
$ python -m adpkd_segmentation.train --config path_to_config_yaml --makelinks
- If using a specific GPU (e.g. device 2):
$ CUDA_VISIBLE_DEVICES=2 python -m adpkd_segmentation.train --config path_to_config_yaml --makelinks
The makelinks flag is optional and needed only once to create symbolic links to the data.
5. Evaluate:
$ python -m adpkd_segmentation.evaluate --config path_to_config_yaml --makelinks
If using a specific GPU (e.g. device 2):
$ CUDA_VISIBLE_DEVICES=2 python -m adpkd_segmentation.evaluate --config path_to_config_yaml --makelinks
For generating TKV calculations:
$ python -m adpkd_segmentation.evaluate_patients --config path_to_config_yaml --makelinks --out_path output_csv_path
Misc:
multi_train.pycan be used to run multiple training runs in a sequence.create_eval_configs.pyis a utility script to create evaluation configs from the starting training config. Also done automatically insidetrain.py.
Contact
For questions or comments please feel free to email me at akshay.k.goel@gmail.com.
Citing
@misc{Goel:2021,
Author = {Akshay Goel},
Title = {ADPKD Segmentation in PyTorch},
Year = {2021},
Publisher = {GitHub},
Journal = {GitHub repository},
Howpublished = {\url{https://github.com/aksg87/adpkd-segmentation-pytorch}}
}
License
Project is distributed under MIT License
Acknowledgement
Model architecture utilized from Segmentation Models Pytorch by Pavel Yakubovskiy.
Linters and Formatters
Please apply these prior to any PRs to this repository.
If you use VSCode you can add these to your settings as follows:
"python.formatting.provider": "black",
"python.linting.flake8Enabled": true,
"python.formatting.blackArgs": [
"--experimental-string-processing",
"--line-length",
"79",
],