#phydynR: Coalescent simulation and likelihood for phylodynamic inference This project is meant to replace the Rforge rcolgem project, but is in alpha stage. The rcolgem package is currently more stable and reliable.
Exponential growth example
Let's model a population which is growing exponentially with fixed per-capita birth rate beta and death rate gamma.
Once we've defined the model, we can simulate trajectories and then simulate genealogies if sampling lineages at specific times.
Finally, we'll see how to infer birth and/or death rates if the tree is observed (e.g. reconstructed from genetic sequence data).
First, load the package and define the rates:
require(phydynR)births <- c(I = 'parms$beta * I' )
deaths <- c(I = 'parms$gamma * I' )The rates are specified as a named vector of strings; names correspond to the names of the demes (I in this case).
The strings are interpreted as R expressions, so should be written just as you would write other R code.
The keywod parms that appears in the rate equations is a list of parameters, which can be set as we will see below.
Now build the demographic process like so:
dm <- build.demographic.process(births=births
, deaths = deaths
, parameterNames=c('beta', 'gamma')
, rcpp=FALSE
, sde = TRUE)Note the following
- You must specify the names of parameters that appear in
parms. - The
sdeoption allows you to speficfy if the model is stochastic (stochastic differential equations) or deterministic (ordinary differential equations) - the
rcppoption says if the equations are written as R code or as C code; we used R code in this case.
Once the model is specified, we can simulate and visualise trajectories:
show.demographic.process( dm
, theta = list( beta = 1.5, gamma = 1 )
, x0 = c( I = 1 )
, t0 = 0
, t1 = 10
) 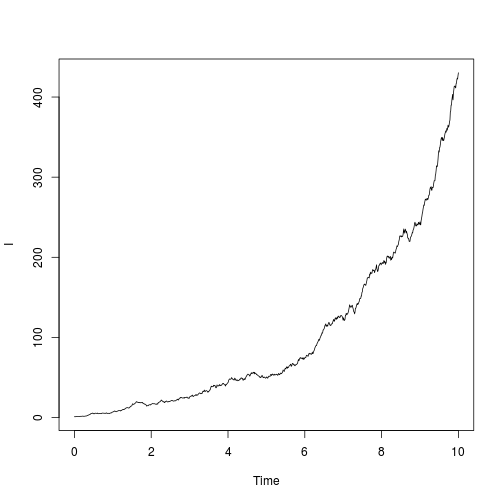 The variables
The variables t0 and t1 determine the time limits of integration. x0 provides a named vector of initial condition; note well that names must correspond to those used to specify the rates. theta is a synonym for parms and provides the parameter list.
You can also directly simulate the model using
dm(theta, x0, t0, t1, res = 1000, integrationMethod = 'adams')
Here integrationMethod is based to the ODE solver and specifies which method to use (e.g. euler or adams). res is the number of time steps to use; larger values will be more accurate at expense of more computation.
We can alternatively specify the equations as C code, in which case the model will be compiled using the Rcpp package.
In this case, simulating the model will be very fast, but it will take a few seconds to compile the model.
Let's compile a deterministic version of the model:
dm.det <- build.demographic.process(births = c(I = 'beta * I')
, deaths = c(I = 'gamma * I')
, parameterNames=c('beta', 'gamma')
, rcpp=TRUE
, sde = FALSE)## [1] "Thu Feb 25 13:24:18 2016 Compiling model..."
## [1] "Thu Feb 25 13:24:25 2016 Model complete"
Note that when we use C code, the parms keyword is not used.
Now let's simulate a coalescent tree conditioning on this demographic process:
tre <- sim.co.tree( list( beta = 1.5, gamma = 1 )
, dm.det
, x0 = c(I = 1 )
, t0 = 0
, sampleTimes = seq(10, 15, length.out=50)
, res = 1000
) This is self-explanotory except for the sampleTimes argument, which is required.
This specifes the times relative to t0 that each lineage is sampled.
The length of this vector determines the sample size.
This can be a named vector, in which case the taxon labels are retained in the returned tree.
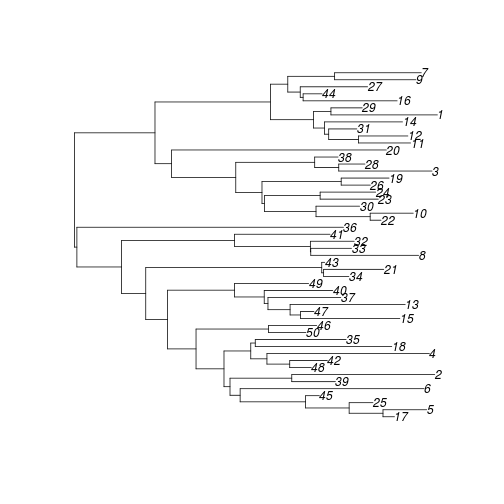
The returned tree is a DatedTree object which subclasses ape::phylo. So, most of the functions in the ape package will also work with the simulated tree. Let's plot it:
plot( ladderize( tre ))Finally, we will see how to compute a likelihood of parameters describing the demographic process given the tree as data.
The colik function can be used to compute this likelihood, and model fitting can be done in a variety of ways; a Bayesian analysis could be done using the mcmc package, or maximum likelihood can be done using the bbmle package.
Note that this returns the log likelihood.
Here is an example invocation of the likelihood function using the simulated tree as data:
colik(tre
, list( beta = 1.5, gamma = 1)
, dm.det
, x0 = c( I = 1 )
, t0 = -1
, res = 1e3
, timeOfOriginBoundaryCondition = FALSE
, AgtYboundaryCondition = TRUE
)## [1] -219.7076
Note the following options:
treis theDatedTreeobject that we fit tox0is the initial conditions for the demographic processt0is the time of origin of the process; you should choose a value that occurs before the root of the treeresis the number of time steps used in the demographic processtimeOfOriginBoundaryCondition(defaultFALSE) : ifTRUE, the function returns-Infif the time of origin occurs before the root of the treeAgtYboundaryCondition(defaultTRUE) : ifTRUE, the function returns-Infif the number of extant lineages in the tree is less than the simulated population size; this should usually be left asTRUE
The phydynR package also includes a convenient wrapper of R's native optim routines which enables easy maximum likelihood inference. Here is an example:
optim.colik(tre
, dm.det
, start = c( beta = 2, gamma = 1, I = 1, t0 = -1)
, est_pars = c( 'beta', 'I')
, ic_pars = c('I')
, t0 = -1
, parm_lowerBounds = c( I = 0, beta = 0)
, timeOfOriginBoundaryCondition = FALSE
, AgtYboundaryCondition = TRUE
, control = list()
) -> fit The return value is the same as returned by optim, so fit$par contains the parameter estimate:
fit## $par
## beta I
## 1.5193956 0.4499639
##
## $value
## [1] 199.8921
##
## $counts
## function gradient
## 137 NA
##
## $convergence
## [1] 0
##
## $message
## NULL
Note the following options:
startcontains the default parameter values including the initial conditions; this may optionally includet0which will specify the time of originest_parsis a vector of parameter names that will be fitted; all remaining parameter values will be fixed at the values instart- You can also fix the time of origin with the
t0argument. - Some parameter values must fit within some pre-defined bounds; for example, the initial population size can not be less than zero, so the
parm_lowerBoundsis a named vector supplying the lower limits of these boundary conditions;parm_upperBoundsprovides the upper limits. controlis a list of options foroptim; see?optimfor more information.