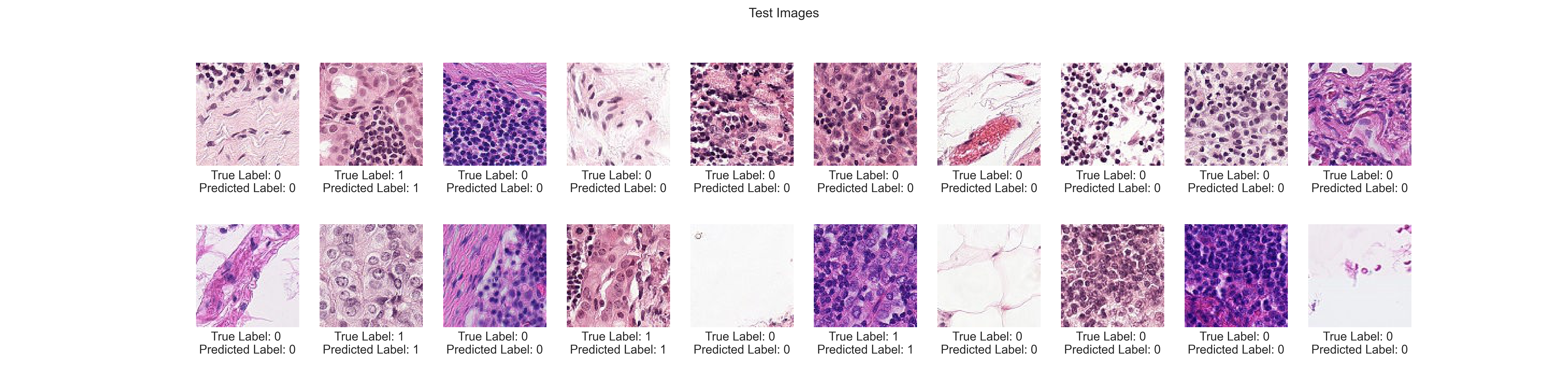
In this project, we tackled the problem of classifying cancer in histopathologic scans of lymph node sections, based on a kaggle challenge1. Check out demo.ipynb for quick a demonstration.
| File/Folder | Description |
|---|---|
dataset.py |
Contains the custom dataset class. |
data_loading.py |
Loads the data from the dataset class to create training and test set. Transforms the data according to project specifications. |
train.py |
Creates and trains a model using command line arguments. |
test.py |
Loads a trained model and evaluates on the test set using command line arguments. |
main.py |
Combination of train.py and test.py. First creates a model, then evaluates it. |
requirements.txt |
Lists all packages used for the project. Designed to be used with pip. |
architecture |
Folder containing all the models that can be used. |
data |
This folder is reserved for the image and label files. |
trained_models |
This folder is reserved for the trained model files (*.pth). |
trained_models_data |
This folder contains training and testing stats. |
imgs |
This folder contains images displayed in this file. |
hyperparameter_tuning |
This folder contains logic and models for hyperparameter tuning. |
README.md |
This file. |
project_report.pdf |
The project report. |
demo.ipynb |
A jupyter notebook demonstrating the classification process with examples. |
| Name | Description |
|---|---|
cnn_1.py |
A simple CNN, referenced in the paper as "Baseline CNN" |
cnn_2.py |
An extension of the simple CNN, referenced in the paper as "Extended CNN" |
resnet18.py |
ResNet18, using the pytorch implementation |
densenet121.py |
DenseNet101, using the pytorch implementation |
mlp_mixer.py |
A custom implementation of the MLP-Mixer architecture proposed by Google 2,3. |
- Python 3.10.6
- packages mentioned in requirements.txt
- PatchCamelyon dataset (Kaggle, Direct Link)
- Trained model files can be downloaded from Google Drive
- clone histopathology-cancer-detection
- cd into histopathology-cancer-detection
- create and activate a custom python virtual environment
- install packages from requirements.txt
$ python -m pip install -r requirements.txt- create a folder
datain the root folder, having the following structure:
histopathology-cancer-detection
└───data
│ │ sample_submissions.csv
│ │ train_labels.csv
│ │
│ └───train
│ | │ 0000d563d5cfafc4e68acb7c9829258a298d9b6a.tif
│ | │ 0000da768d06b879e5754c43e2298ce48726f722.tif
│ | │ ...
│ │ |
| └───test
│ | │ 0000ec92553fda4ce39889f9226ace43cae3364e.tif
│ | │ 000c8db3e09f1c0f3652117cf84d78aae100e5a7.tif
│ | │ ...
- put all trained model files (*.pth) in the
trained_modelsfolder
| Name | Description | Required | Available for Files |
|---|---|---|---|
--name |
How the output files will be named. | Yes | train.py, test.py, main.py |
--model_name |
Which model to be used for training. Can be one of the following: cnn_1, cnn_2, resnet18, densenet121, mlp_mixer. Corrensponds to the file names in architecture. |
Yes | train.py, test.py, main.py |
--lr |
Determine the learning rate. Default is 0.001. | No | train.py, main.py |
--epochs |
Determine the number of epochs. Default is 10. | No | train.py, main.py |
Example:
$ python .\train.py --name baseline_cnn --model_name cnn_1 --lr 0.003 --epochs 5Hyperparameter tuning is seperated from the other files. It has no command line arguments. Instead, it must be adjusted in the main_optuna.py file. The default setting is set to optimize densenet.
The folder hyperparameter_tuning contains two files and one folder:
main_optuna.pyis the starting point for the hyperparameter tuning and contains the tuning logictrain_optuna.pycontains the training logicarchitecturecontains model files adepted for hyperparameter tuning
- csv file containing training metrics and loss function values for all training batches.
- PyTorch .pth file containing the state dict of the trained model.
- csv file containing metrics calculated on the test set.
- csv file containing training metrics and loss function values for all training batches.
- PyTorch .pth file containing the state dict of the trained model.
- csv file containing metrics calculated on the test set.
- pkl file containing all details about the hyperparameter search. Can be loaded by optuna.
- csv file containing all details about the hyperparameter search in human-readable format.
| Model | #Parameters | Learning Rate | Epochs | Batch Size | BCE Loss | Training Accuracy | Test Accuracy | Training Recall | Test Recall | Training F1-Score | Test F1-Score |
|---|---|---|---|---|---|---|---|---|---|---|---|
| CNN_1 | 5.8K | 0.001 | 10 | 64 | 0.1840 | 0.9260 | 0.9281 | 0.8940 | 0.8987 | 0.9070 | 0.9108 |
| CNN_2 | 108.6K | 0.001 | 10 | 64 | 0.1980 | 0.9230 | 0.9195 | 0.8870 | 0.8832 | 0.9020 | 0.8989 |
| AlexNet Transfer Learning | 12.2M | 0.001 | 10 | 64 | 0.6743 | 0.5969 | 0.5948 | 0.0000 | 0.0000 | 0.0000 | 0.0000 |
| MLP Mixer | 17.4M | 0.001 | 10 | 64 | 0.0853 | 0.9688 | 0.9153 | 0.9547 | 0.9069 | 0.9610 | 0.8967 |
| MLP Mixer + Dropout | 17.4M | 0.001 | 10 | 64 | 0.1378 | 0.9481 | 0.9109 | 0.9312 | 0.9171 | 0.9354 | 0.8930 |
| ResNet18 | 11.2M | 0.001 | 10 | 64 | 0.0657 | 0.9752 | 0.9528 | 0.9684 | 0.9281 | 0.9693 | 0.9409 |
| DenseNet121 | 6.9M | 0.001 | 10 | 64 | 0.0793 | 0.9717 | 0.9569 | 0.9605 | 0.9369 | 0.9648 | 0.9463 |
1 https://www.kaggle.com/competitions/histopathologic-cancer-detection/
2 https://arxiv.org/pdf/2105.01601.pdf
3 https://github.com/google-research/vision_transformer/blob/linen/vit_jax/models_mixer.py