YipCat package can realize some of the more commonly used analysis requirements of transcription, such as trajectory analysis,heatmap, cell interaction,imputeweight,cellphonedb,agescore etc , you can view the function description inside the package. Mainly the implementation style of the diagram is more advanced to look at, in particular, the mass spectrometry streaming data is also integrated and can achieve the interaction of cellranger cloupe files, very convenient.
you can install this package via command:
install.packages("yipCat_1.0.1.tar.gz")require(Seurat)
require(dplyr)
require(ggplot2)
require(S4Vectors)
require(nabor)
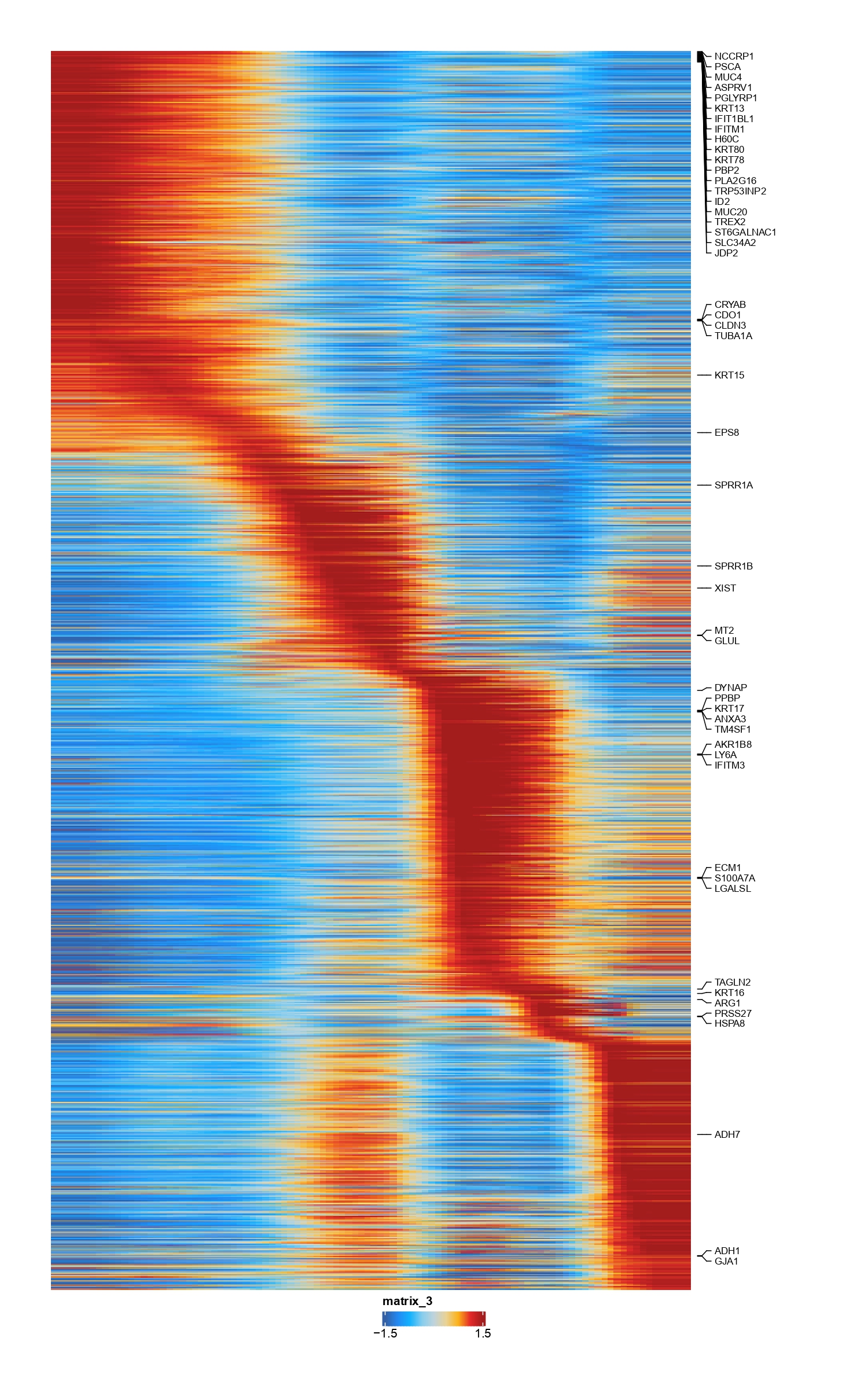
...The implementation of spline trajectory, the specific method description mainly draws on part of the ArchR method, the diagram is as follows:
you can caculate trajecory by follwing commands,for example
seurat<-addSeuratTrajectory(object=seurat,trajectory=c("memory B","Naive B","Plasma"),groupBy="label_fine",embedding="pca") # caculate trajecory
se<-getSeuratTrajectory(seurat) # get trajectory object
ArchR::plotTrajectoryHeatmap(se) # plot trajectory heatmapcomputes imputations weights that describe each cell as a linear combination of many cells based on a MAGIC diffusion matrix.
for example
seurat<-ImputeWeights(seurat,reducedDims="pca",nRep=2,sampleCells=5000) # caculate impute weight
weight<-getImputeWeight(seurat) # return impute weight
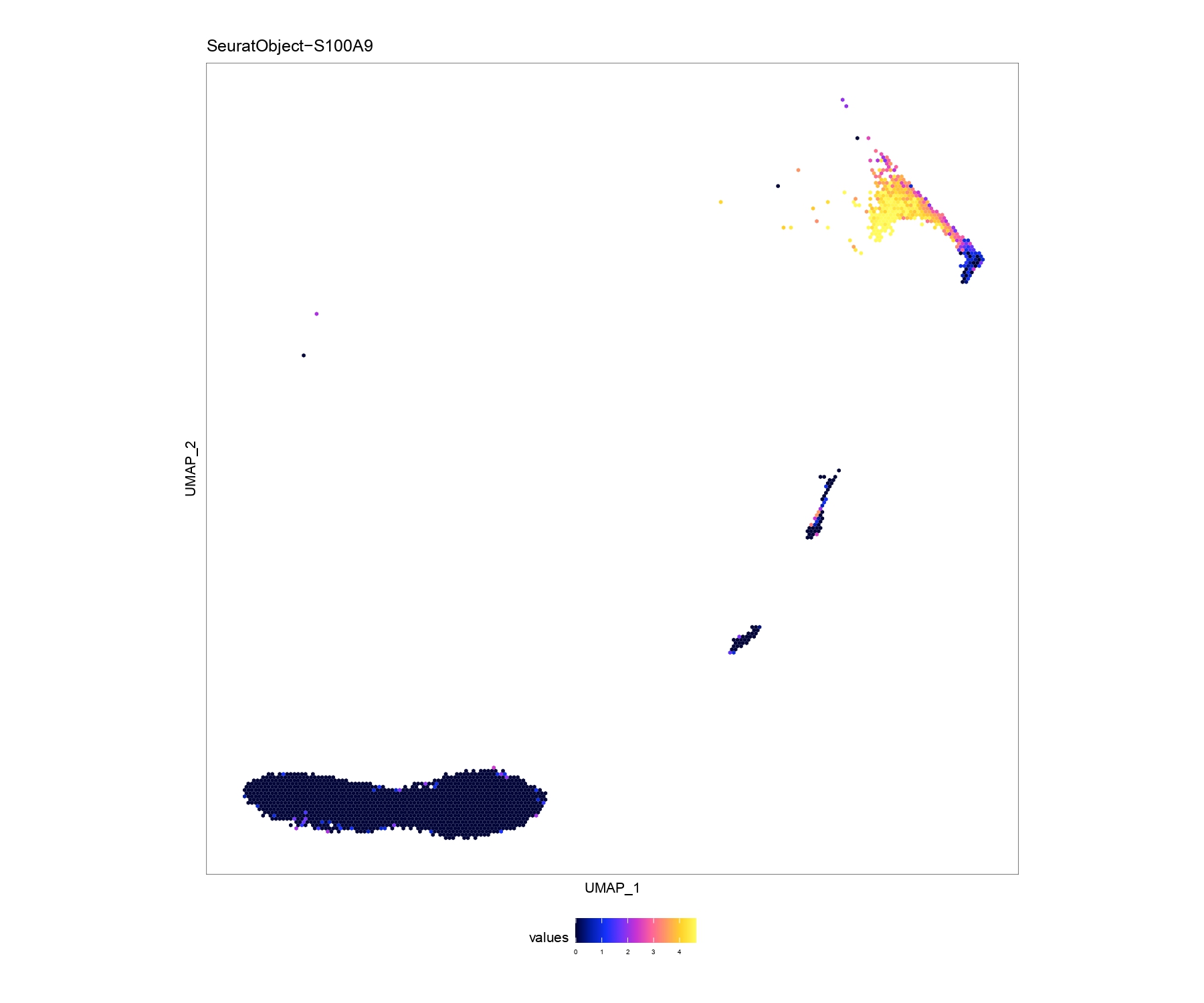
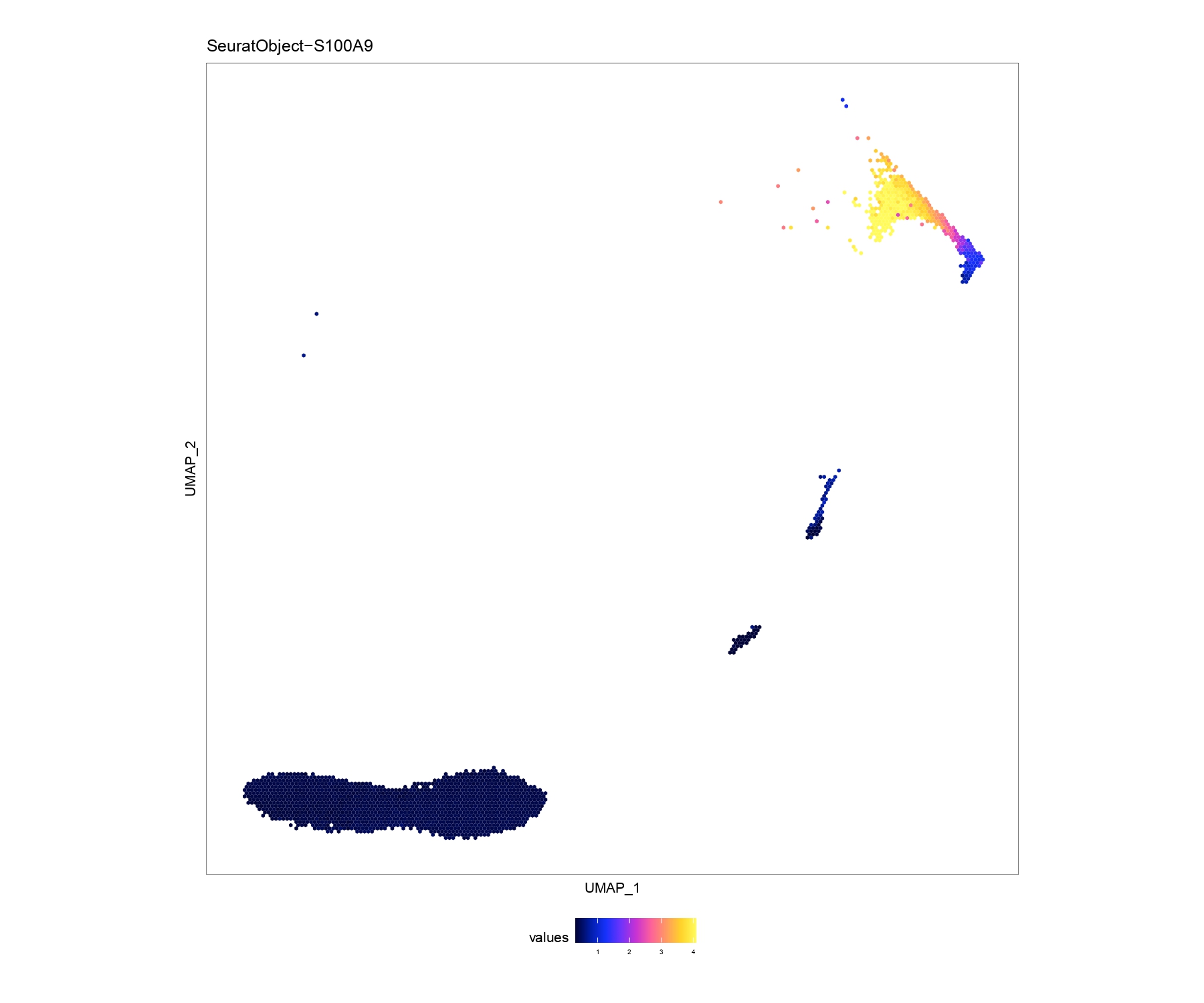
EmbPlot(seurat,colorBy="matrix",features="S100A9",embedding="umap",imputeWeights=NULL) # first plot
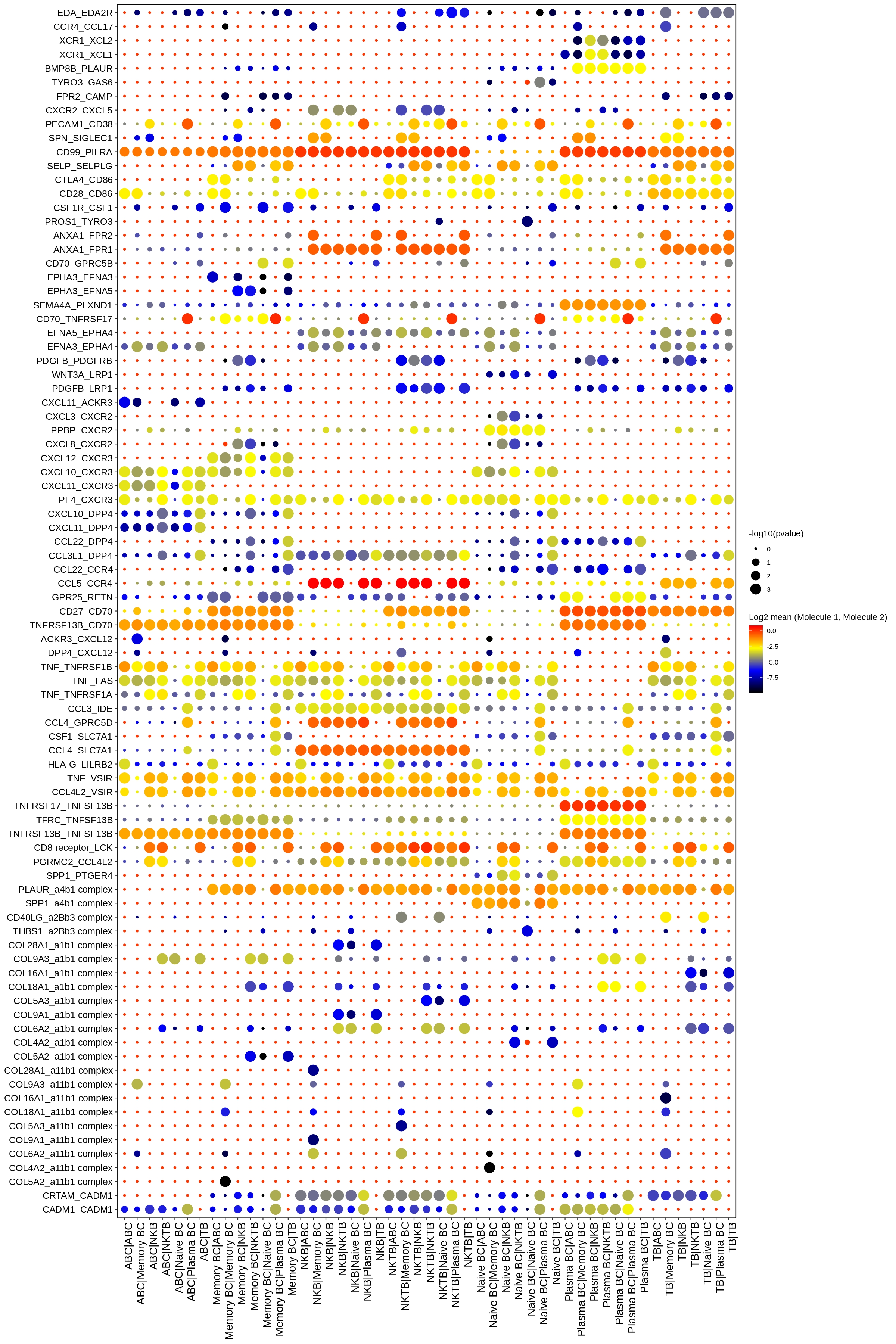
EmbPlot(seurat,colorBy="matrix",features="S100A9",embedding="umap",imputeWeights=weight) # second plotthis package also wrap cellphonedb ,you can choose relative functions to realize what you want for example
exportCellPhoneDB(seurat,cells=cells,features=features,selectCol="label_fine",runCPDB=TRUE) # selectCol : which column to caculate cell-cell interaction,runCPDB=TRUE,run cellphonedb backgroup
CPDBDotplot # cellphonedb result dotplot
CPDBHeatmaps # cellphonedb result heatmapCustomInteractionScore function can caculate custom cell-cell interaction
interaction_csv<-system.file("extdata", "interaction.csv", package = "yipCat")
df<-CustomInteractionScore(seurat,column=column,idents=c("DC","NK","BC"),interaction_csv=interaction_csv)For each sample in the SeuratObject provided, this function will independently assign inferred doublet information to each cell. This allows for removing strong heterotypic doublet-based clusters downstream. A doublet results from a droplet that contained two cells, causing the ATAC-seq or scRNA data to be a mixture of the signal from each cell.
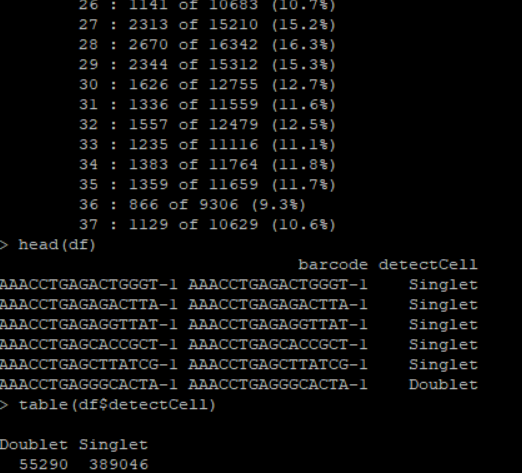
seurat<-calDoubletScores(seurat,sampleCol="Sample",threads=8) # caculate doublet score
scoreDF<-filterDoublets(seurat) # get doublet score data frameThis package can realize the processing of cytof data, the cytof data is sampled by sample, the default is 20,000 samples, and then processed into Seurat Object, can achieve umap, tsne, pca and other de-dimensional, cytof visualization, and can be used with the impute weight method;In addition, cytof data can be interacted with cellranger to generate close files for easy cell classification by researchers
sample_csv<-system.file("extdata", "cytofSample.csv", package = "yipCat")
config_csv<-system.file("extdata", "cytofConfig.csv", package = "yipCat")
seurat<-Cytof2Seurat(sample_csv=sample_csv,config_csv=config_csv,N=20000,path2barcode10X="3M-february-2018.txt")after run bellow command,will get data matrix in 10X format(eg,filtered_feature_bc_matrix) then run shell script to convert matrix into relative gzip 10X matrix
bash inst/extdata/cytofTo10x.sh filtered_feature_bc_matrixand,can run cellrnager reanalysis via(only support cellranger-3.1.0)
generateCloupe(...)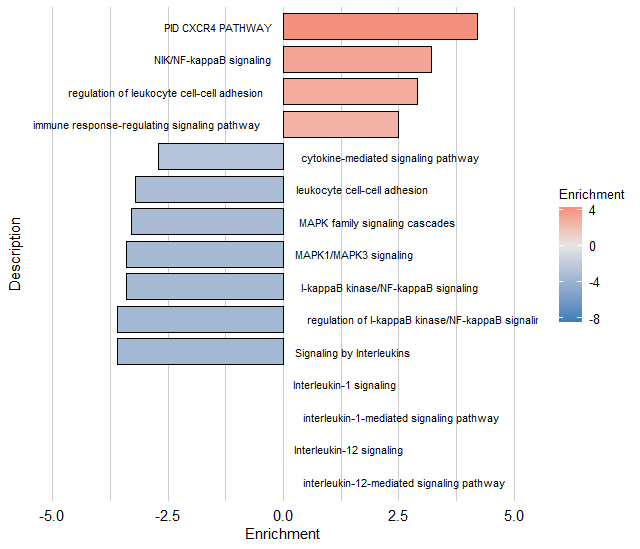 For additional usage, check out the package's function description
For additional usage, check out the package's function description
If this does not fix your problem, please report an issue on Github with the Bug Report form.