Kuramoto model for community detection
Method explanation
Let
The dynamic function is $$ \frac{\partial \theta_i}{\partial t} = \frac{1}{N}\sum_{j:(i,j) \in E} \sin(\theta_j - \theta_i) $$
Local order during simulation step
Solve the ODE until
Then we use
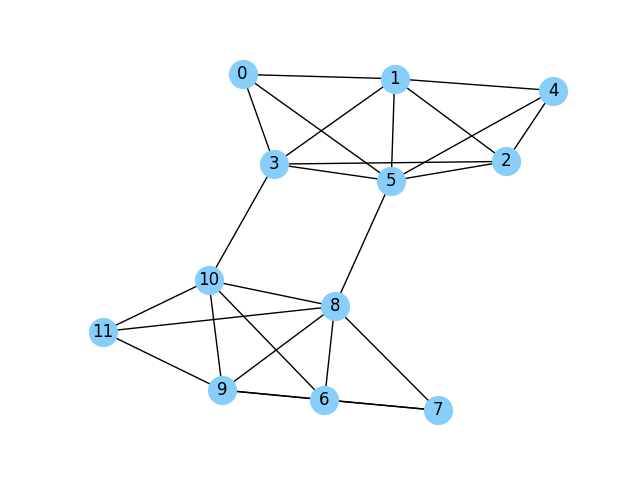
Example
For the above graph with 12 nodes. After
[4.85074828 4.84967319 4.84943349 4.86083052 4.846165 4.85887088
4.91879579 4.92206347 4.90935898 4.91855667 4.90739992 4.91748298]Pros and Cons
Pros: This method is bio-inspired, and shows potential in plausible explanation and theoretical justification.
Cons:
-
Slow as the number of nodes grows large,
since we need to solve an ODE system with
$N$ variables. - Low accuracy compared with other methods. We test this method on SBM(100,2,16,4), which shows that it has low accuracy compared with SDP-based method.
Reference
Implementation of syncnet in pyclustering.