The human microbiome consists of trillions of microorganisms cohabitating a variety of body niches. Microbiota can modulate the host physiology through microbiota-derived molecule and metabolite interactions. Microbiome-based biomarkers have great potential for numerous disease states but current approaches have yielded limited success.
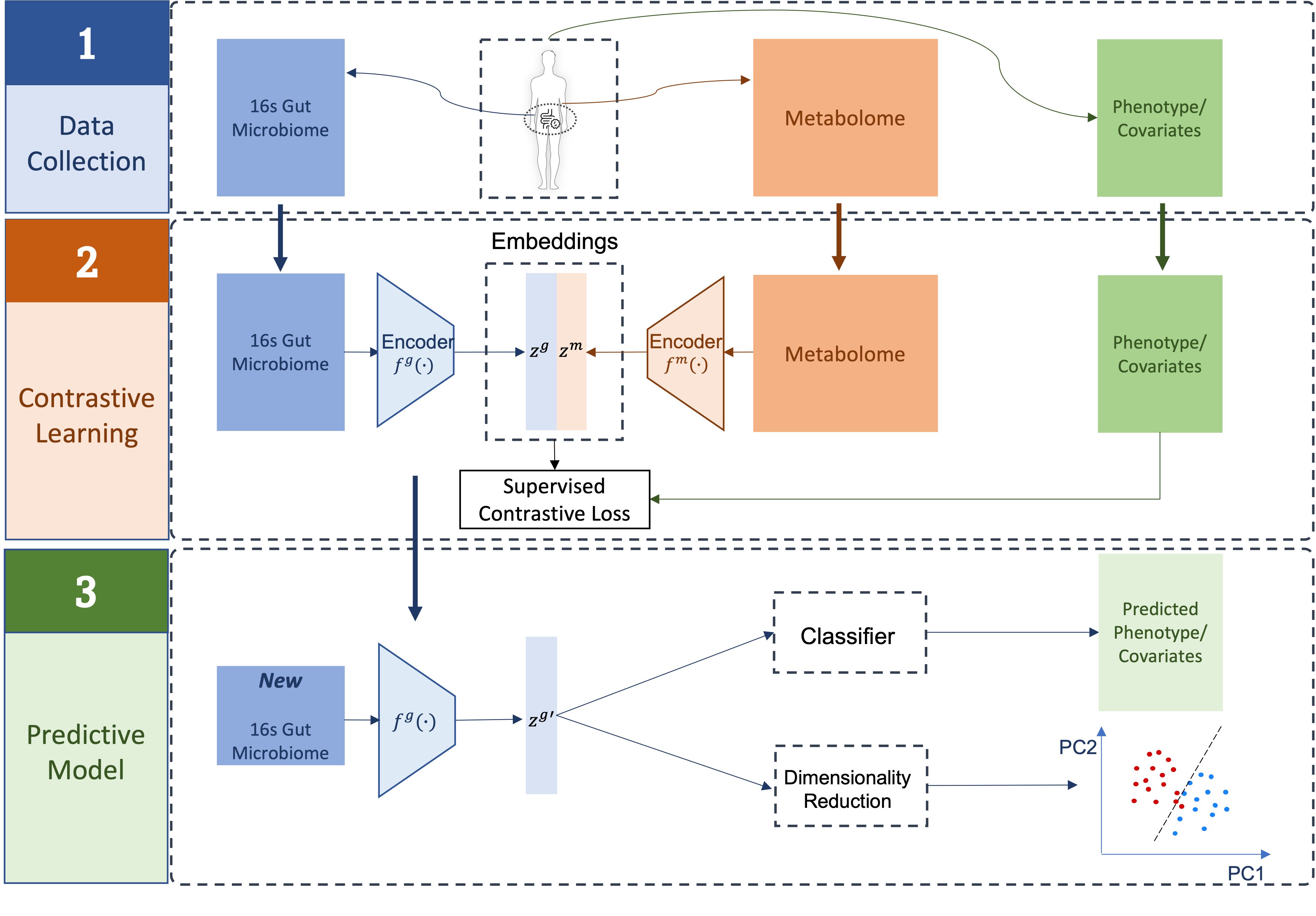
Here, we propose a novel integrative modeling framework, Microbiome-based Supervised Contrastive Learning Framework (MB-SupCon) to improve microbiome-based prediction models. By integrating microbiome and metabolomics data under a supervised contrastive learning scheme, MB-SupCon trained two encoder networks to maximize the similarity between microbiome embedding and metabolomics embedding. Compared to the original microbiome data, the microbiome embedding can lead to improved prediction performances.
Fig.1 MB-SupCon: Microbiome-based Supervised Contrastive Learning Framework
Folders
"data" folder includes:
-
Raw gut 16s microbiome data and metabolome data;
-
Clinical covariates data of all subjects;
-
Output index labels for training, validation and testing datasets;
-
Output PC1 and PC2 scores (by PCA) used for lower-dimensional scatter plots.
"feature_data" folder includes feature embeddings by MB-SupCon in representation space for all covariates.
"figures" folder includes all loss curves and generated lower-dimensional scatter plots.
"models" folder includes trained models.
"results" folder includes some history data during training.
"other methods" folder includes the codes, plots and outputs of the other methods for comparison with MB-SupCon.
Codes
1a - train MB-SupCon_categorical covariates.ipynb: Jupyter Notebook used for training MB-SupCon models for all covaraites and generating corresponding feature embeddings in the representation domain;
2a - prediction from embeddings of MB-SupCon.ipynb: Jupyter Notebook used for prediction of each covariates by logistic regression with elastic net regularization;
3a - lower-dim plotting by PCA on embeddings of MB-SupCon.ipynb: Jupyter Notebook used for performing PCA on embeddings and plotting on the lower-dimensional space (PC2 vs. PC1; colored by different covariate labels);
MB_SupCon_utils.py, plotting_utils.py, pred_utils.py, utils_eval.py: utility functions;
supervised_loss.py: a function used for calculating supervised contrastive loss.
Some system information - System: Linux; Release: 3.10.0-957.el7.x86_64.
GPU: Tesla V100-PCIE-32GB.
Python version: 3.8.5.
Some main packages used in this study:
pytorch: 1.7.1 (Build: py3.8_cuda11.0.221_cudnn8.0.5_0);
numpy: 1.19.2;
pandas: 1.2.1;
scikit-learn: 0.23.2;
matplotlib: 3.3.2;
seaborn: 0.11.2;
plotly: 4.14.3.
Sen Yang:
senyang@smu.edu | sen.yang@utsouthwestern.edu
Department of Statistical Science
Southern Methodist University
Dallas, TX 75275
Xiaowei Zhan:
xiaowei.zhan@utsouthwestern.edu
Quantitative Biomedical Research Center, Department of Population and Data Sciences
Center for Genetics of Host Defense
University of Texas Southwestern Medical Center
Dallas, TX 75390