CNNs, transfer learning, VGG16 pre-trained model, Keras, TensorFlow backened, TF serving for production/ deployment
Transfer learning using VGG16 and convolutional neural networks (CNNs) to classify images of freshwater snails and parasites. This automated detection approach is expected to make infectious disease control more efficiently in the field. This is also the FIRST deep-learning-powered image classifer for parasite worms of Schistosomiasis. The classifier of snails (4 species) has 99% accuracy; parasite classifer has 92% accuracy (11 species).
The script is in Python, building CNNs and transfer learning (with VGG16), using Keras package with Tensorflow backened. Augmenting in training data is also built in the code (to overcome the unbalanced datasets).
Main script: cnn_transfer_learning_2.py --> the scripts here can be used for different image classification tasks.
Deployment script: tf_serving_index.py --> built with TensorFlow serving (make sure you have already produced bottleneck_features_train.npy and bottleneck_features_validation.npy in the directory)
python tf_serving_index.py
Note that the image dataset is the property of De Leo Lab of Hopkins Marine Station at Stanford University. The images will be released with the publication in few months.
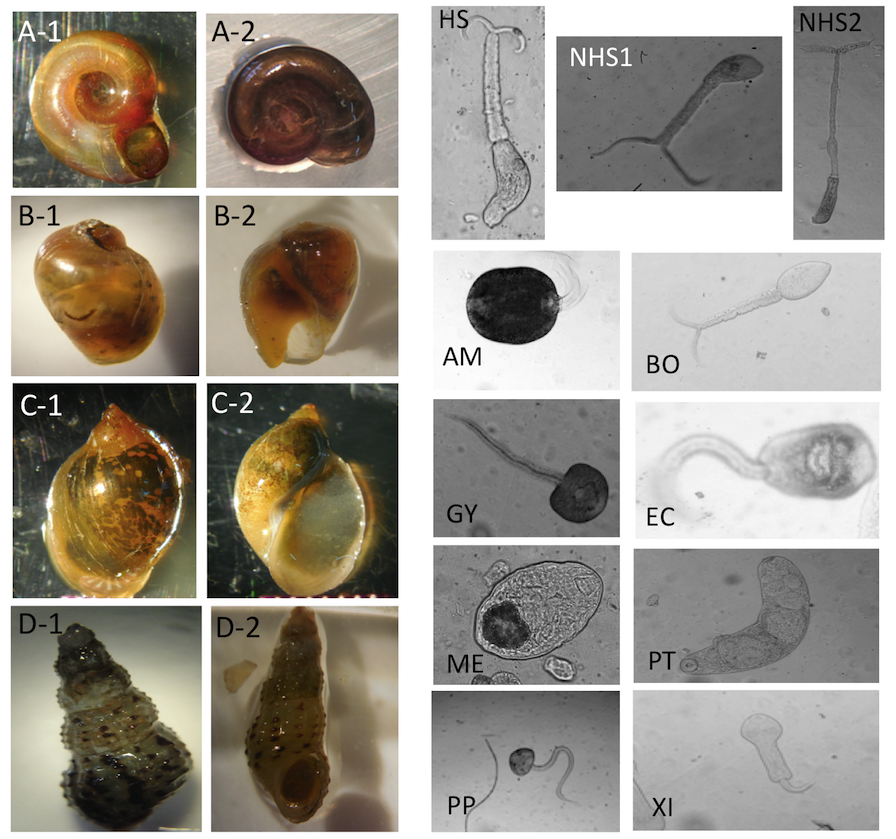
Image examples for snail and parasite classes. For snail classes, A-1 and A-2: Biomphalaria. B-1 and B-2: Bulinus. C-1 and C-2: Lymnaea. D-1 and D-2: Melanoides. For parasite classes, HS: Human-schisto, NHS1: Nonhuman- schisto forktail type I, NHS2: Nonhuman- schisto forktail type II, AM: Amphistome, BO: Bovis, EC: Echino, GY: Gymno, ME: Metacerc, PP: Parapleurolophocercous, PT: Parthenitae, XI-Xiphidiocercariae.
Links: