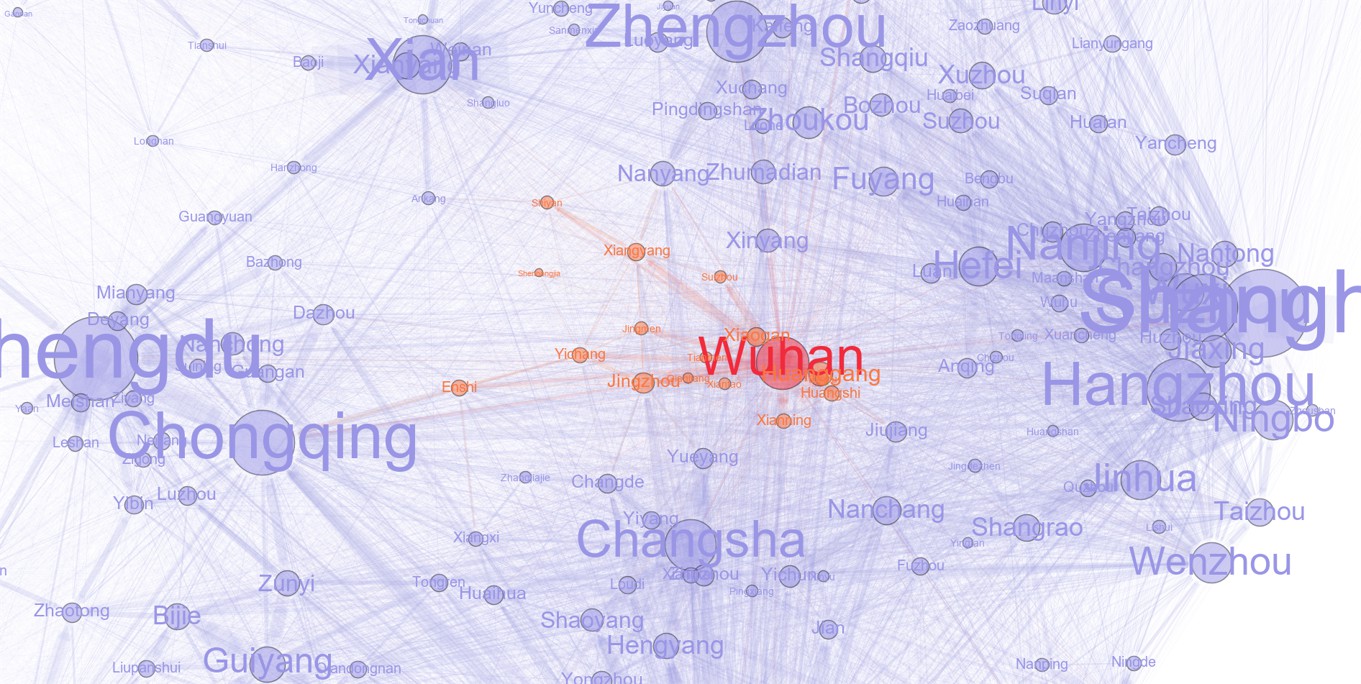
Figure1-Plot.ipynb: plot Figure 1 in Python
Figure2-Spatial analysis by R.ipynb: spatial analysis of the spread of COVID-19 in R
Figure2-Spatial analysis and plot.ipynb: spatial analysis of the spread of COVID-19 and plot Figure 2 in Python
Figure3-Outbreak analysis by R.ipynb: outbreak analysis of the spread of COVID-19 in R
Figure3-Plot.ipynb: plot Figure 3 in Python
Figure4-Plot.ipynb: plot Figure 4 in Python
To execute the tutorial, make sure you have Python 3, R/R-Studio and Jupyter notebook installed.
To run R script in Jupyter, IRkernel is required. This package is available on CRAN and you can install it in R/R-Studio Console by:
install.packages('IRkernel')
IRkernel::installspec() # register the kernel in the current R installation
os
numpy
pandas
itertools
seaborn
matplotlib
statsmodels
collections
readr
MASS
dplyr
tidyr
texreg
Should you have any further quires about the code and data, please contact at: yflyzhang_at_gmail.com