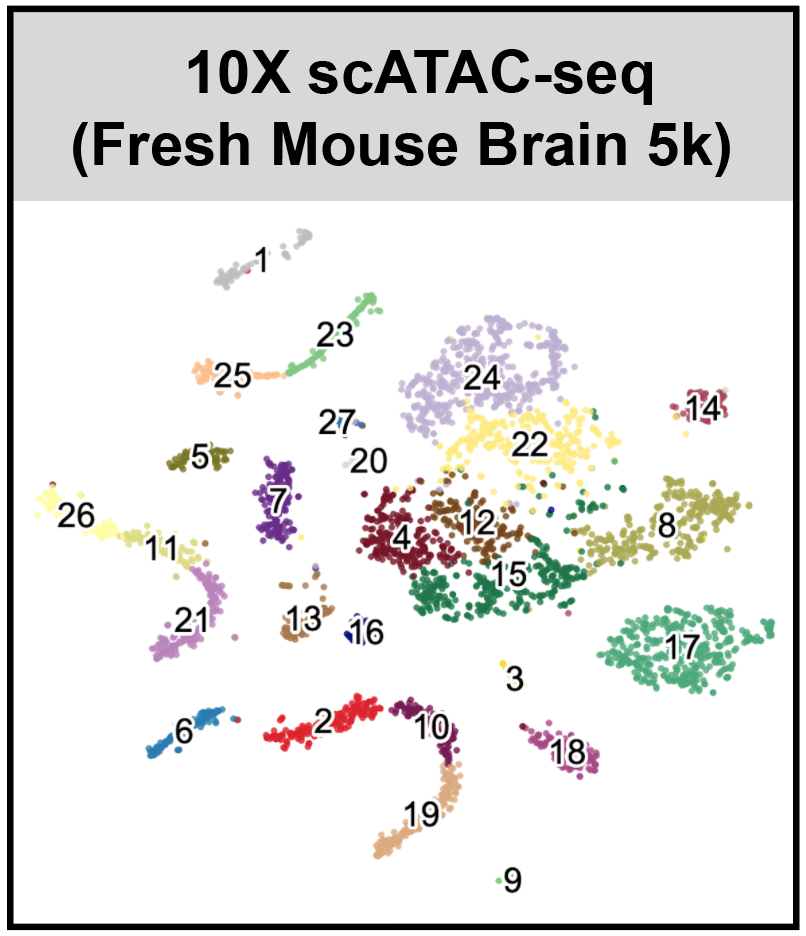
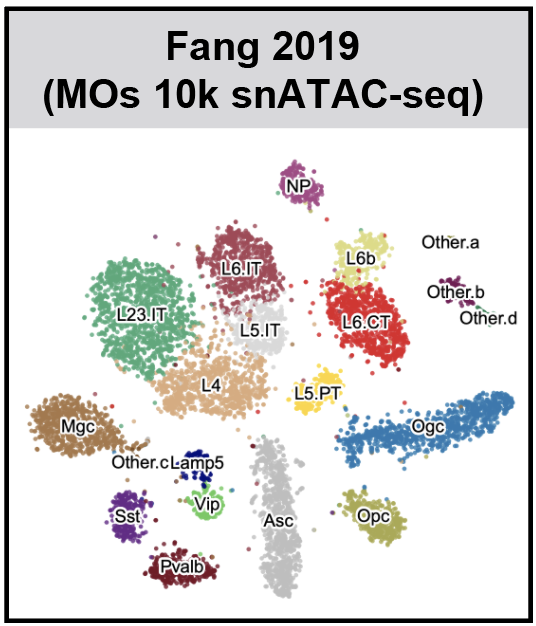
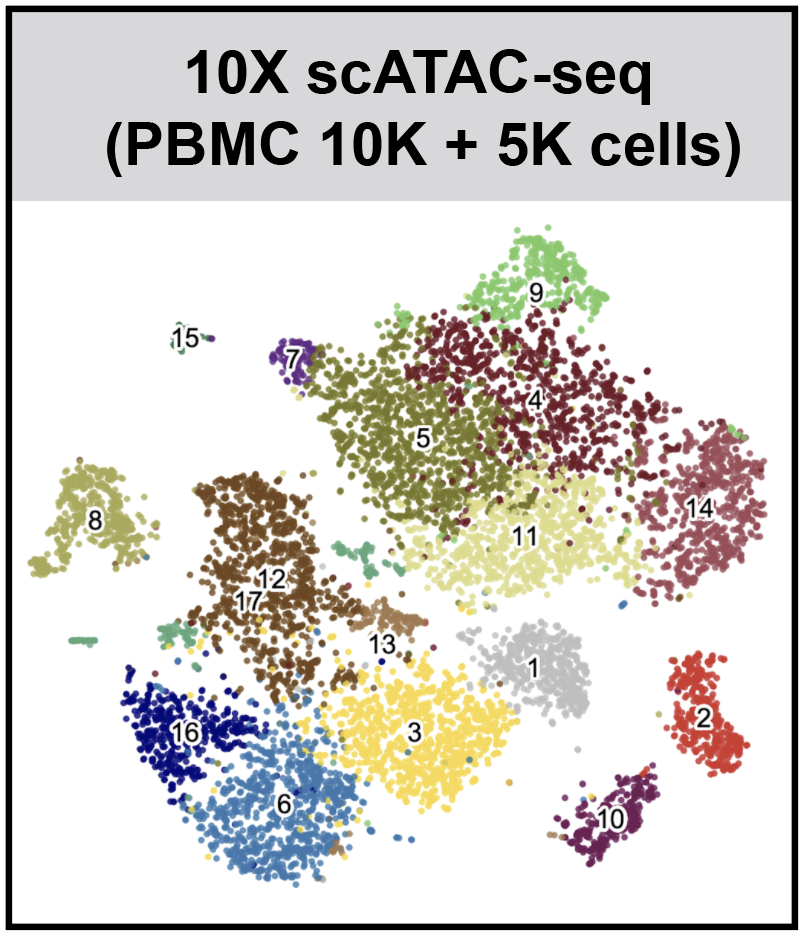
SnapATAC (Single Nucleus Analysis Pipeline for ATAC-seq) is a fast and accurate method for analyzing single cell ATAC-seq datasets. SnapATAC 1) overcomes the limitation of reliance on population-level peak annotation, 2) improves the clustering accuracy by integrating "off-peak" reads, 3) controls for the major bias using a regression-based normalization method and 4) substantially outperforms current methods in scalability.
- How to run SnapATAC on 10X dataset?
- I already ran CellRanger, can I use its output for SnapATAC?
- How can I analyze combine multiple samples together?
- How to group reads from any subset of cells?
- What is a snap file anyway?
- How to create a snap file from fastq file?
- Linux/Unix
- Python (>= 2.7) (SnapTools)
- R (>= 3.4.0) (SnapATAC)
SnapATAC has two components: Snaptools and SnapATAC.
- SnapTools - a python module for pre-processing and working with snap file.
- SnapATAC - a R package for the clustering, annotation, motif discovery and downstream analysis.
Install snaptools from PyPI. See how to install snaptools on FAQs.
$ pip install snaptoolsInstall SnapATAC R pakcage (development version).
$ R
> library(devtools)
> install_github("r3fang/SnapATAC")