Tools for working with volumes at DTU G-bar.
Log on a real node using X11 forwarding:
linuxsh -X
Get the code from this repository. For example, from the command line:
wget https://github.com/vedranaa/vis3d/archive/refs/heads/main.zip
unzip main.zip
(You can place the directory with the code wherever it suits you, and you can also rename it. Also, there is a subfolder NOT_USED, which you may delete.)
Navigate to the project folder (i.e. the folder containing init.sh and setup.py):
cd <PROJECT FOLDER>
Run configuration script which will load modules, create a virtual environment, and activate it:
. init.sh
Run editable install (from the folder containing setup.py):
pip install -e .
Test the setup by running:
vis3d
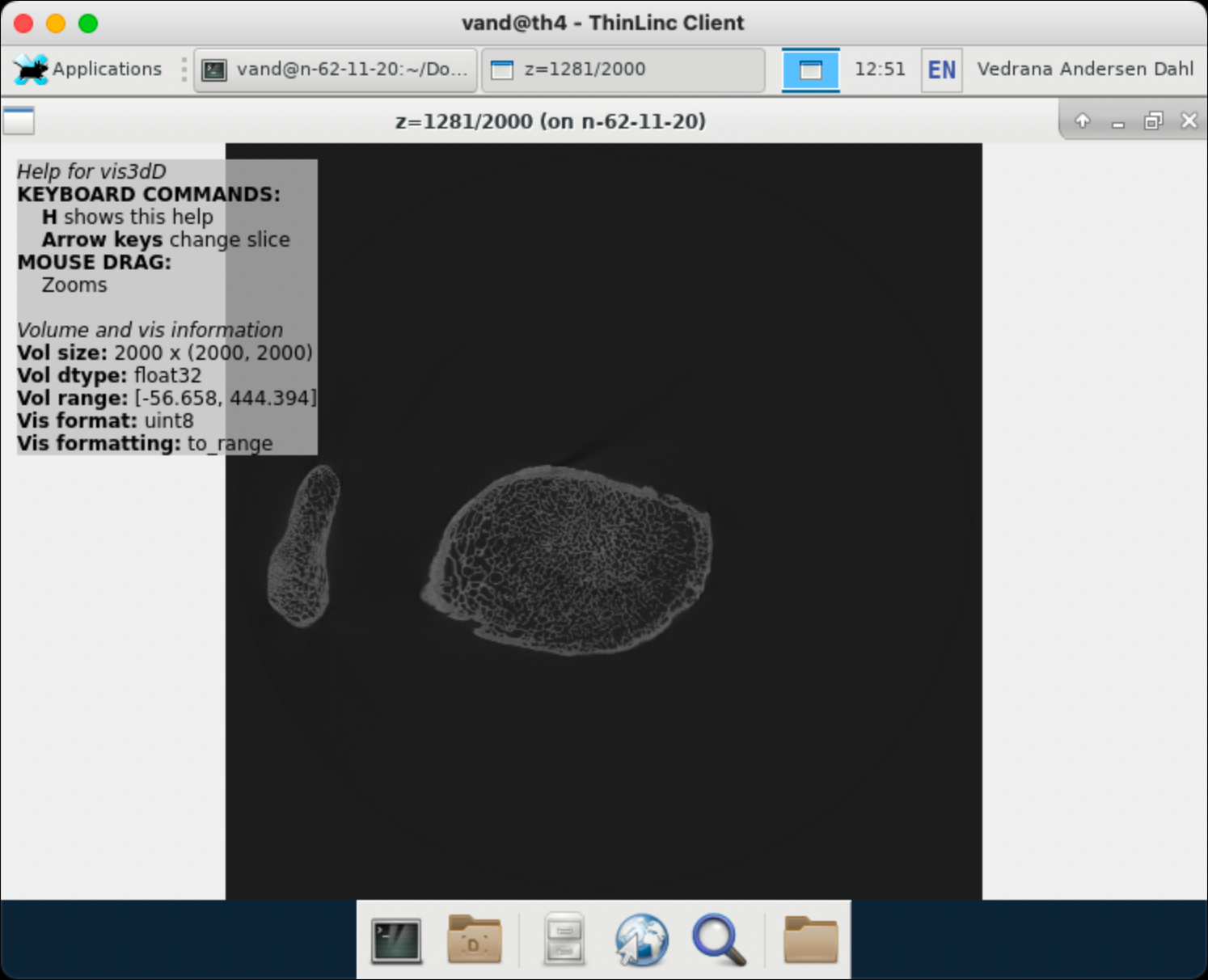
This should open a file navigation window. Navigate to folder links_gbar and open for example bone_reconstruction.txt. This should result in another window being opened, showing a central slice from the volume. The window is interactive, hold key H for help.
Use the commands below to log on a node, navigate to the project folder, and run init.sh:
linuxsh -X
cd <PROJECT FOLDER>
. init.sh
Use vis3d from the terminal with either:
vis3d
(which should open a file navigation window allowing you to open a file/folder, for example, one of the files from links_gbar) or by specifying a file/folder:
vis3d <PATH TO FILE/FOLDER>
This should open an interactive window, hold key H for help.
- folder containing images
- .tif file with stacked images
- URL to .tif file
- .vgi and corresponding .vol file
- .txm file
- .txt file containing a URL or file/folder path
Save a volume as downscaled tif from the command line using
tiffify <SOURCE> <DESTINATION> --factor <FACTOR>
For example:
tiffify somewhere/something.vgi here/this.tif --factor 4
- When the slicer points to a non-existent file/folder, it errors saying something strange. TODO: Before trying to open the volume using any slicer, check that all needed files exist, and if not, give an informative error message.