SharpTNI is a problem which takes a timed phylogeny with leaf labeling and host entry-exit times as input and finds counts and uniformly samples the number of minimum transmission number host labelings. It also counts and samples the solution space of transmission networks with minimum transmission number and a fixed co-tranmsission number.
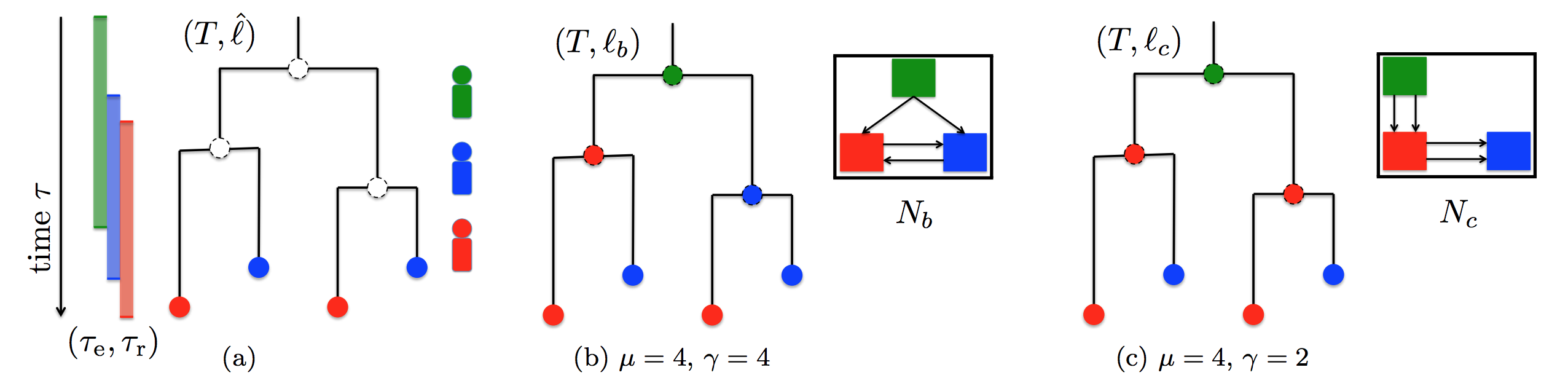 The evolutionary history of the pathogenic strains in an outbreak is
described by a timed phylogeny T, assigning a time-stamp to
every vertex.
In addition, each leaf is labeled by the host where
the corresponding strain was observed (indicated by colors).
Epidemiological data further constrain the entrance and removal time of each host.
In the TNI problem, we seek a host labeling with minimum
transmission number and subsequently smallest co-transmission number.
(b) Host labeling with minimum transmission but not
the smallest co-transmission number, resulting in a complex
transmission network.
(c) Host labeling with minimum transmission and
smallest co-transmission number, resulting in a
parsimonious transmission network.
The evolutionary history of the pathogenic strains in an outbreak is
described by a timed phylogeny T, assigning a time-stamp to
every vertex.
In addition, each leaf is labeled by the host where
the corresponding strain was observed (indicated by colors).
Epidemiological data further constrain the entrance and removal time of each host.
In the TNI problem, we seek a host labeling with minimum
transmission number and subsequently smallest co-transmission number.
(b) Host labeling with minimum transmission but not
the smallest co-transmission number, resulting in a complex
transmission network.
(c) Host labeling with minimum transmission and
smallest co-transmission number, resulting in a
parsimonious transmission network.
SharpTNI solver is written in C++11 and requires a modern C++ compiler (GCC >= 4.8.1, or Clang). In addition it has the following dependencies
Graphviz is required to visualize the resulting DOT files, but is not required for compilation.
To compile execute the following commands from the root of the repository
$ mkdir build
$ cd build
$ cmake ..
$ make
In case CMake fails to detect LEMON, run the following command with adjusted paths:
$ cmake -DLIBLEMON_ROOT=~/lemon
The compilation results in the following files in the `build' directory
| EXECUTABLE | DESCRIPTION |
|---|---|
sankoff |
count/enumerate the minimum transmission number host labelings |
sample_sankoff |
uniformly sample minimum transmission number host labelings |
gamma |
optimum clique partitioning for a given host labeling |
dimacs |
SAT formulation for SharpTNI problem |
The SharpTNI input is text based. There are two input files, host file and ptree file. Each line of the host file has exactly 3 entries separated by ' '. The format of each line of the host file is '<host name> <entry time> <removal time>' in each line. The number of lines in the host file is the number of sampled hosts. A ptree file gives the timed phylogeny with the leaf labeling. Each line of ptree file has exactly 4 entries separated by ' '. The format for each line of the ptree file is '<node name> <child1 name> <child2 name> <host label>'. The number of lines in the ptree file is the number of nodes in the timed phylogeny. The nodes of the tree in the file must be in post-order (all nodes must be preceded by their children). For a leaf the must be '0'.
Usage:
./sankoff [--help|-h|-help] [-b] [-c] [-e] [-l int] [-r int] [-t] [-u
int]
<host> / <transmission_tree> <ptree> <output_ptree>
Where:
--help|-h|-help
Print a short help message
-b
is the tree non binary (default: false)
-c
Find consensus Sankoff solution (deafault: false)
-e
Enumerate all the solutions (default: false)
-l int
Enumeration solution number limit (default: intMax)
-r int
Root label (default: 0)
-t
Transmission tree instead of host file
-u int
Number of unsampled hosts (default: 0)
An example execution:
$ ./sankoff ../data/sample/sample_host.out ../data/sample/sample_ptree.out ../data/sample/sample_enum.out -u 1 -e -l 5
Usage:
./sample_sankoff [--help|-h|-help] [-b] [-l int] [-r int] [-u int]
<host>
<ptree> <output_prefix>
Where:
--help|-h|-help
Print a short help message
-b
is the tree non binary (default: false)
-l int
Number of samples (default: 11000)
-r int
Root label (default: 0)
-u int
Number of unsampled hosts (default: 0)
An example execution:
$ ./sample_sankoff -l 1 -u 1 ../data/sample/sample_host.out ../data/sample/sample_ptree.out ../data/sample/sample_
Usage:
./gamma [--help|-h|-help] [-b] [-u int] <host> <ptree_sol>
Where:
--help|-h|-help
Print a short help message
-b
is the tree non binary (default: false)
-u int
Number of unsampled hosts (default: 0)
An example execution:
$ ./gamma -u 1 ../data/sample/sample_host.out ../data/sample/sample_idx0_count1.out 2> ../data/sample/example.dot
$ dot -Tpng ../data/sample/example.dot -o ../data/sample/example.png
Usage:
./dimacs [--help|-h|-help] [-k int] [-r int] [-t] [-u int]
<host> / <transmission_tree> <ptree> <output_dimacs_file>
<output_varlist_file>
Where:
--help|-h|-help
Print a short help message
-k int
number of co-infection events (default: m-1)
-r int
Root label (default: 0)
-t
Transmission tree instead of host file
-u int
Number of unsampled hosts (default: 0)
An example execution:
$ ./dimacs ../data/sample/sample_host.out ../data/sample/sample_ptree.out ../data/sample/sample_dimacs.cnf ../data/sample/sample_varlist.txt -u 1 -k 4