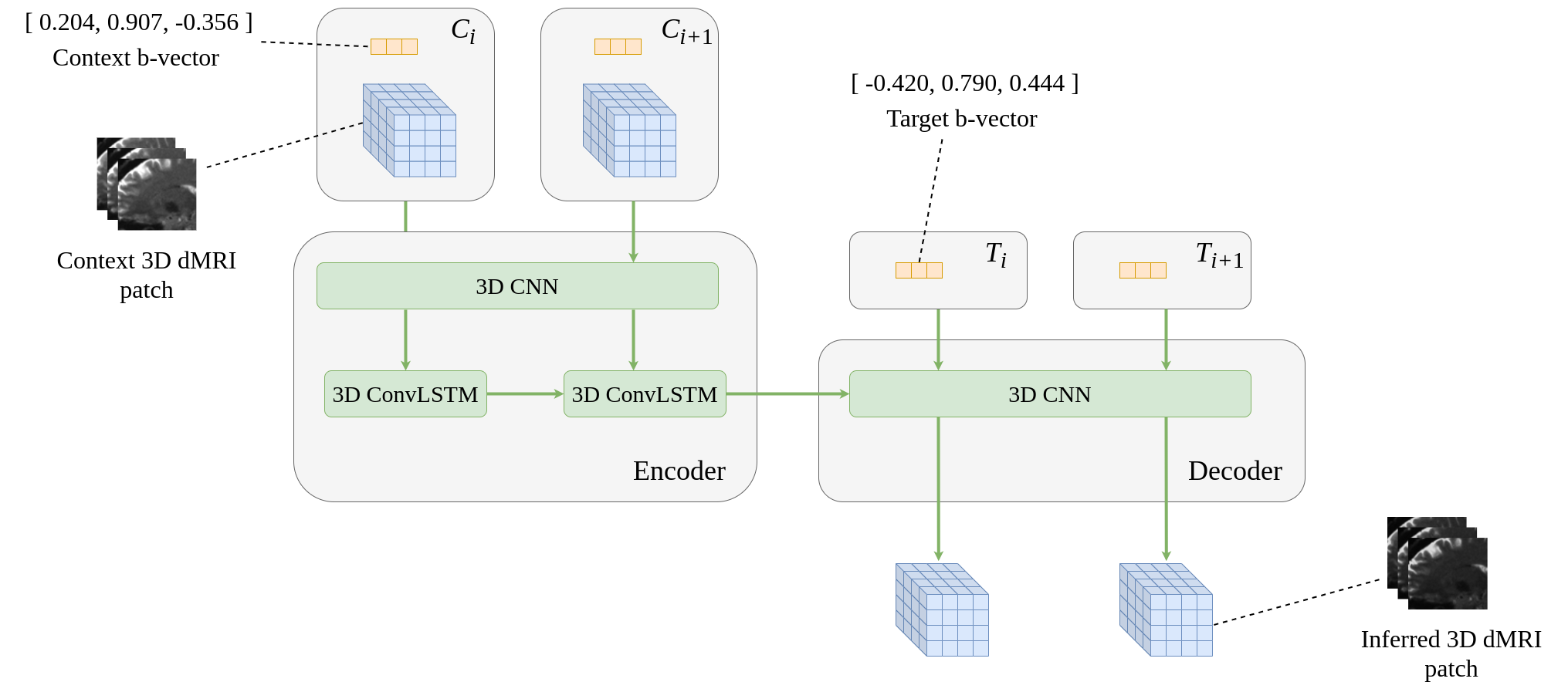
This project enhances the angular resolution of dMRI data through the use of a Recurrent CNN.
dMRI-RCNN can be installed by first downloading a release, then install via pip:
pip install dMRI-RCNN-{version}.tar.gzdMRI-RCNN uses TensorFlow as the deep learning architecture.
Listed below are the requirements for this package.
tensorflow>=2.6.0numpyeinopsnibabel
Once installed, use run_dmri_rcnn.py to perform inference of new dMRI volumes. Below lists the data requirements to use the script, and the commandline arguments available for inference.
To run this script, dMRI data is required in the following format:
- Context dMRI file. The dMRI data used as context within the model to infer other volumes
- File format:
NIfTI - Single-shell: containing only one b-value.
- Dimensions:
(i, j, k, q_in).(i, j, k)are the spatial dimensions of the dataq_innumber of samples within the q-space dimension. This can either be6,10, or30and will affect which of the trained models is used.
- File format:
- Context b-vector file. The corresponding b-vectors for the context dMRI file.
- File format: text file, whitespace delimited.
3rows corresponding to thex, y, zco-ordinates of q-spaceq_incolumns corresponding to the q-space directions sampled.q_inmust either be6,10, or30.
- Target b-vector file. The corresponding b-vectors for the inferred dMRI data.
- File format: text file, whitespace delimited.
3rows corresponding to thex, y, zco-ordinates of q-spaceq_outcolumns corresponding to the q-space directions sampled.
- Brain mask file. Binary brain mask file for dMRI data.
- File format:
NIfTI - Dimensions:
(i, j, k). Same spatial dimensions as used in the dMRI data.
- File format:
The script will create the following data:
- Inferred dMRI file. dMRI volumes inferred from the model as defined by the target b-vectors.
- File format:
NIfTI - Dimensions:
(i, j, k, q_out).q_outnumber of samples within the q-space dimension. This can any number, though using higher numbers will require more GPU memory if using.
- File format:
Bring up the following help message via run_dmri_rcnn.py -h:
usage: `run_dmri_rcnn.py` [-h] -dmri_in DMRI_IN -bvec_in BVEC_IN -bvec_out BVEC_OUT -mask MASK -dmri_out DMRI_OUT -s {1000,2000,3000} [-m {1,3}] [-c] [-b BATCH_SIZE]
optional arguments:
-h, --help show this help message and exit
-dmri_in DMRI_IN Context dMRI NIfTI volume. Must be single-shell and contain q_in 3D volumes
-bvec_in BVEC_IN Context b-vector text file. Whitespace delimited with 3 rows and q_in columns
-bvec_out BVEC_OUT Target b-vector text file. Whitespace delimited with 3 rows and q_out columns
-mask MASK Brain mask NIfTI volume. Must have space spatial dimensions as dmri_in.
-dmri_out DMRI_OUT Inferred dMRI NIfTI volume. This will contain q_out inferred volumes.
-s {1000,2000,3000}, --shell {1000,2000,3000}
Shell to perform inference with. Must be same shell as context/target dMRI and b-vectors
-m {1,3}, --model-dim {1,3}
Model dimensionality, choose either 1 or 3.
-c, --combined Use combined shell model. Currently only applicable with 3D model and 10 q_in.
-b BATCH_SIZE, --batch-size BATCH_SIZE
Batch size to run model inference with.
The following example performs b = 1000 inference with the 3D dMRI RCNN.
$ run_dmri_rcnn.py -dmri_in context_dmri.nii.gz -bvec_in context_bvecs -bvec_out target_bvecs -mask brain_mask.nii.gz -dmri_out inferred_dmri.nii.gz -s 1000 -m 3
This example would take ~2 minutes to infer 80 volumes on an NVIDIA RTX 3080.
Future Additions & Improvements:
- Training Pipeline.
- Addition of the training pipeline to allow finetuning & further user experimentation within the framework.
- Docker support.