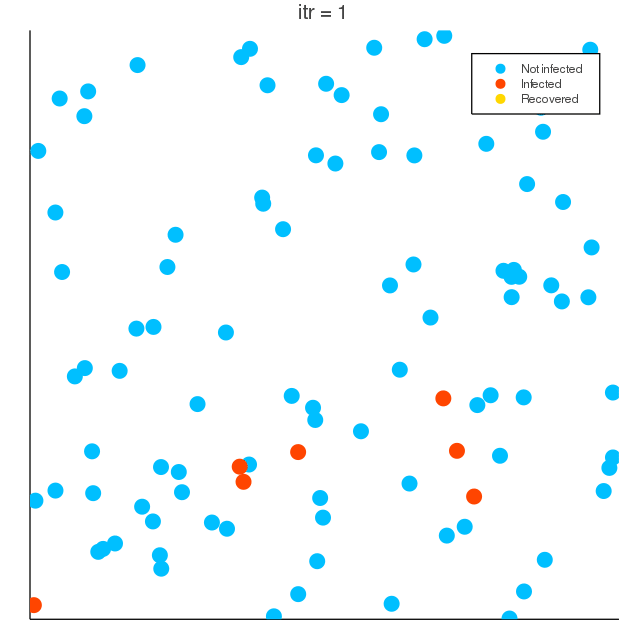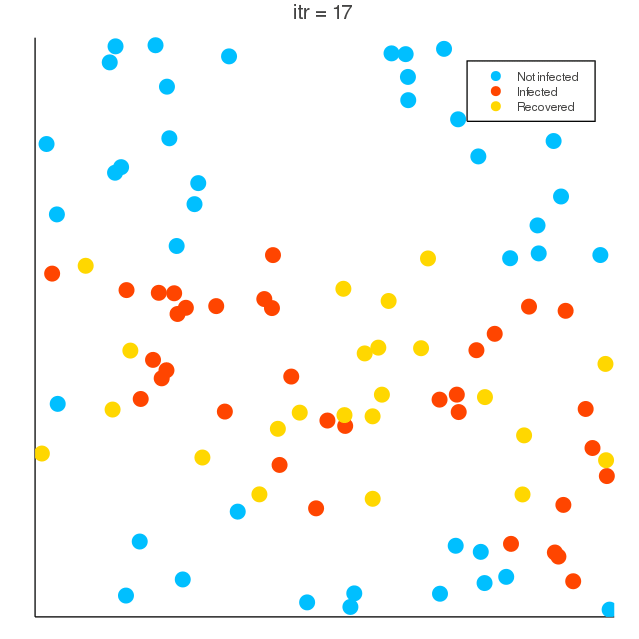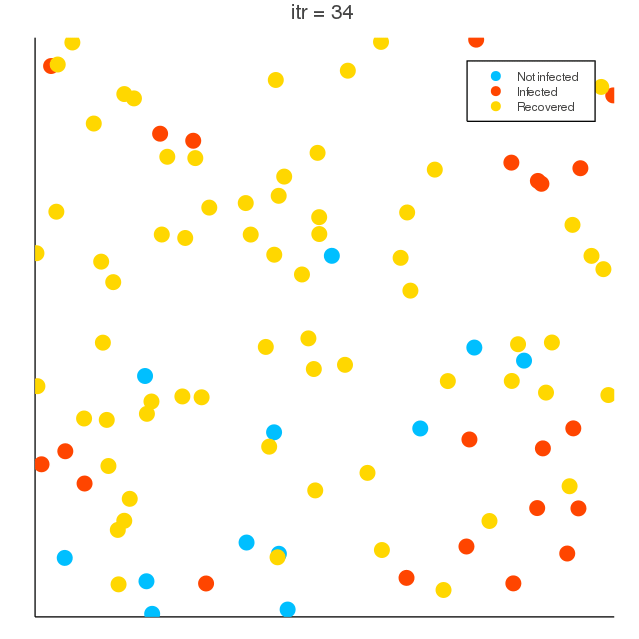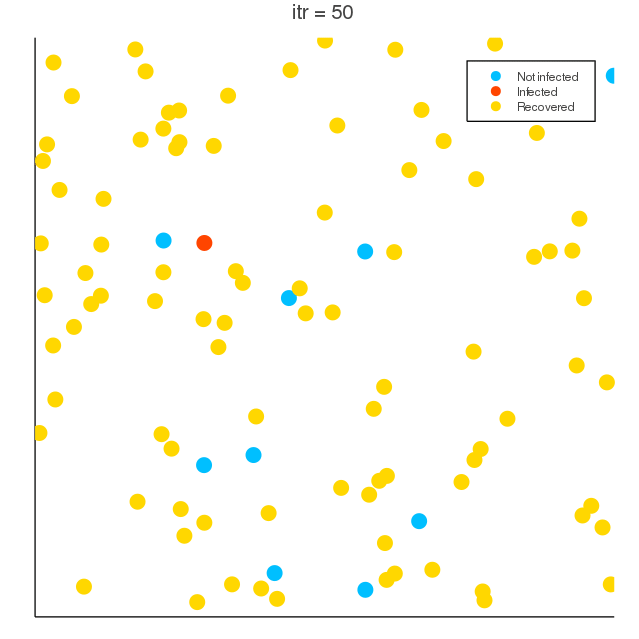
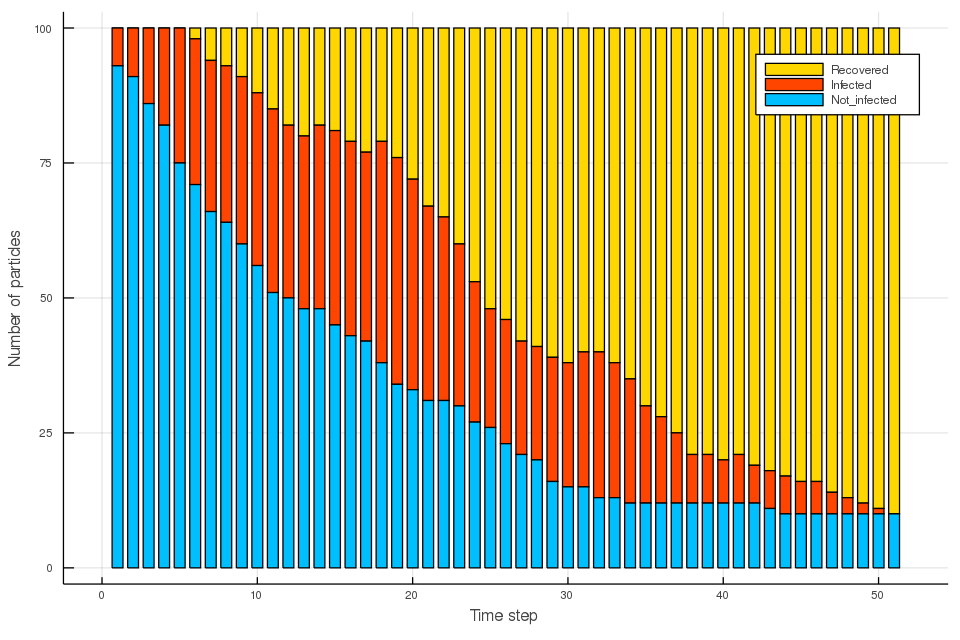
COVID-19 spreading simulation, Julia translation of https://github.com/MuAuan/collective_particles with several changes and new functionalities.
num_particlesparticles are simulated overmax_iterationtemporal iteration.- Computational domain is
[0, x_range]×[0, y_range]. - Initial particle velocity
(vel_x, vel_y)obeys Gaussian distribution, with averagevel_meanand standard deviationvel_σ. ratio_infection_initpercentage of particles are initially infected by the virus.- Particles in the nearby (below relative distance
radius_infection) of the infected particle can be infected by the chance rateinfection_chanceper time step. - Infected particles (deterministically) recover after
recovery_timesteps and never infected again. - During temporal development, particle velocity alters by up to
vel_flc.
flag_multiple_infectionallows recovered particles to re-infect the virus.- In the case of multiple infection,
infection_chanceis modified by number of past infection.
- In the case of multiple infection,
flag_infected_isolationforces infected particles to stop their movement during infection.
- The code is tested on Julia version 1.3.1.
./fig/subdirectory is needed for gif video.- Module dependencies are ProgressMeter, Distributions, Printf and Plots with GR backend.