An R package to estimate auto- and cross-correlations from time-series
data sampled with any arbitrary sampling scheme. Developed with Nick Jacobson and Kejin Wu
The package takes a time-series dataset with measurement timing information as the input. Relying on a simple data-stacking approach and Generalized Additive Mixed Models (GAMMs), it allows researchers to:
- Estimate auto- and cross-correlations at different time-intervals requested by the user.
- Visualize how lagged correlations vary and evolve as a function of the time-interval between measurements
- Construct confidence intervals (CIs) to quantify uncertainty around estimated lagged correlations.
This repository contains an R package used by Ryan, Wu & Jacobson (in
prep) Exploratory Continuous-Time Modeling (expct): Extracting Dynamic
Features from Irregularly Spaced Time Series.
The current version of this package can be installed directly from github using
devtools::install_github("ryanoisin/expct")
library(expct)The package takes as input a time-series dataset or long-format longitudinal data. The dataset should have the following columns
ida column denoting the id number of each participant in the dataset. Note that for single-subject time series, this should simply be a column with a single number repeated, e.g.data$id = 1timea column containing timing information for each observation. The specific format required is “time elapsed since the first observation” in a unit of the users choice (hours, minutes, etc.). For example, if the first observation of the dataset is taken at 3pm, and the second observation at 5pm, then the first two entries of the time column should readtime[1:2] = c(0,2), encoding time in the unit of hours.- The remaining columns should contains the variables that the user wants to compute auto and/or cross-correlations for.
load("data/simdata.rda")
head(simdata)## id time Y1 Y2
## [1,] 1 0.000000 1.91866673 0.7673046
## [2,] 1 3.338756 -0.76230415 0.6131818
## [3,] 1 5.878493 -0.58990520 0.4041070
## [4,] 1 7.950279 1.72802101 -0.3894506
## [5,] 1 10.060890 1.34938143 1.4604215
## [6,] 1 14.870422 0.01682572 3.1393569
The main function of this package is expct. The key input options are:
dataset: the dataset in the format described aboveTpred: A vector which indicates the time-intervals at which the user wants to estimate the auto- and cross- correlations.output_type: Determines the method used to construct credible or confidence intervals. If output_type == “CI”, then default GAM confidence intervals are supplied, which can be interpreted as “point-wise” CIs for each of the values inTpred. If output_type ==“SCI”, then simultaneous CIs are supplied. These can be interpreted as “function-wide” CIs, see here for an introduction. If output_type ==“LLCI”, CIs are computed by approximating analytic auto- and cross-correlation standard errors from the time-series literature. The default value is “CI”. We recommend using either “CI” or “SCI”, with the latter being more conservative
Other input options allow users to request bootstrap estimation, request
pre-estimation detrending, and control estimation of the lagged
correlations by passing arguments to mcgv::gam().
The output of this functions is a list which contains below elements:
est: Matrix containing point estimates of lagged correlationshighCI: The upper 95 percent CI of the point estimation.lowCI: The lower 95 percent CI of the point estimation.- The ``stacked’’ dataset used in estimation of lagged effects
attributes: All attributes used to estimate GAMMs.
Perform analyses with expct:
# library(mgcv)
Tpred = seq(1,30,1)
out <- expct(
dataset = simdata,
Time = "time", # name of the column in dataset with timing informaiton
outcome = c("Y1","Y2"), # optional: which variables to compute lagged corrleations for
ID = "id", # name of id column
Tpred = Tpred,
plot_show = F, # plot output
method = "bam" # option to be passed to mgcv::gam()
)Plot output, for instance using
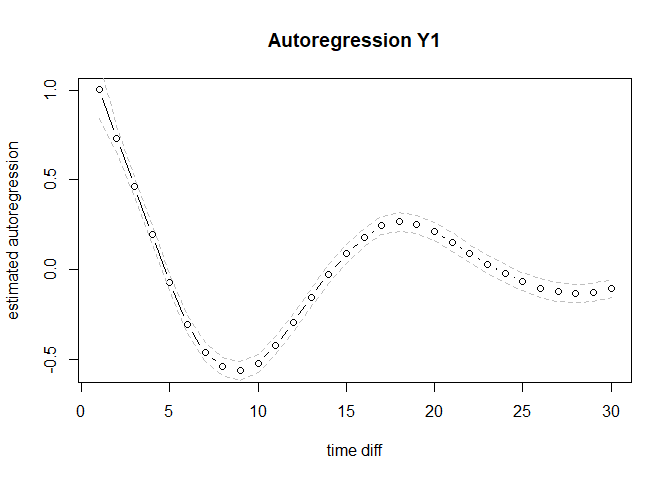
plot(out$est$Y1toY1, type = "b", ylab = "estimated autoregression", xlab = "time diff", main = "Autoregression Y1")
lines(out$highCI$Y1toY1, lty = 2, col = "gray")
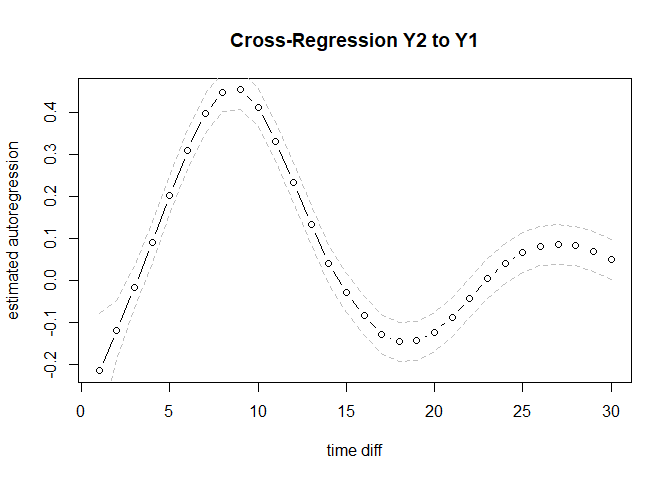
lines(out$lowCI$Y1toY1, lty = 2, col = "gray")plot(out$est$Y2toY1, type = "b", ylab = "estimated autoregression", xlab = "time diff", main = "Cross-Regression Y2 to Y1")
lines(out$highCI$Y2toY1, lty = 2, col = "gray")
lines(out$lowCI$Y2toY1, lty = 2, col = "gray")For more details please contact o.ryan@uu.nl