This repository is forked from huawei-noah/vega which contains the implementation of NAGO.
- The configuration file for running NAGO on CIFAR10
nago.ymlis under./examples/nas/nago/. - The function class for defining the search space of NAGO (i.e. given the generator hyperparameters, we draw a sample architecture from the defined generator and convert it into a trainable PyTorch network model) is
nago.pyunder./zeus/networks/pytorch/customs/nago.py. - The key utility functions (
logical_graph.py,layer.pyandops.py) required innago.pyare under./zeus/networks/pytorch/customs/utils/.
In this code package VEGA, we (a) use BOHB as the search strategy over the generator hyperparameters, (b) draw a trainable architecture sample from the generator via nago.py and (c) use the default trainer to train and evaluate the architecture sample from the architecture generator. If you want to use your own search strategies and/or your own trainer, you just need to take the relevant code files for NAGO and replace component (a) and (c) with your own implementation
The search space of NAGO described above can be specified in the configuration file nago.yml as follows.
search_space:
type: SearchSpace
hyperparameters:
- key: network.custom.G1_nodes
type: INT
range: [3, 10]
- key: network.custom.G1_K
type: INT
range: [2, 5]
- key: network.custom.G1_P
type: FLOAT
range: [0.1, 1.0]
- key: network.custom.G2_nodes
type: INT
range: [3, 10]
- key: network.custom.G2_P
type: FLOAT
range: [0.2, 1.0]
- key: network.custom.G3_nodes
type: INT
range: [3, 10]
- key: network.custom.G3_K
type: INT
range: [2, 5]
- key: network.custom.G3_P
type: FLOAT
range: [0.1, 1.0]Note despite we are using the NAS pipeline, we define the search space following HPO pipeline format as we use BOHB, which is a HPO method, to perform the search.
Our NAGO search space is amenable to any Bayesian optimisation search strategies. In this code package, we use BOHB to perform the optimisation and the configuration of BOHB needs to be specified in nago.yml. The example below defines a BOHB run with eta=2 and t=50 search iterations; each iteration evaluate 7 configurations. The minimum and maxmimum training epochs used for evaluating a recommended configuration is 30 and 120 respectively.
search_algorithm:
type: BohbHpo
policy:
total_epochs: -1
repeat_times: 50
num_samples: 350
max_epochs: 120
min_epochs: 30
eta: 2- Install and set-up vega following the instruction. Note
MMDectionpackage is not needed for NAGO. - Define the NAGO configuration file
nago.ymlas suggested above and put dataset at thedata_pathspecified innago.yml - Run the command
cd ./examples & python3 run_pipeline.py ./nas/nago/nago.yml
The following two files are generated in the specified output directory (the default directory is ./example/tasks/task_id/output/nas/):
- The
output.csvfile contains the best architecture generator hyperparameters found by BOHB - The
reports.csvfile contains all the architecture generator hyperparameters queried by BOHB at different epoch.
Note the exact validation accuracy of the architectures found during the search phase may be slightly lower than that reported in the paper because in VEGA, the batch size and learning rate needs to be prespecified and fixed for the network trainer but in our original code base, we will adjust the batch size to maximise the use of the GPU memory and scale the learning rate accordingly.
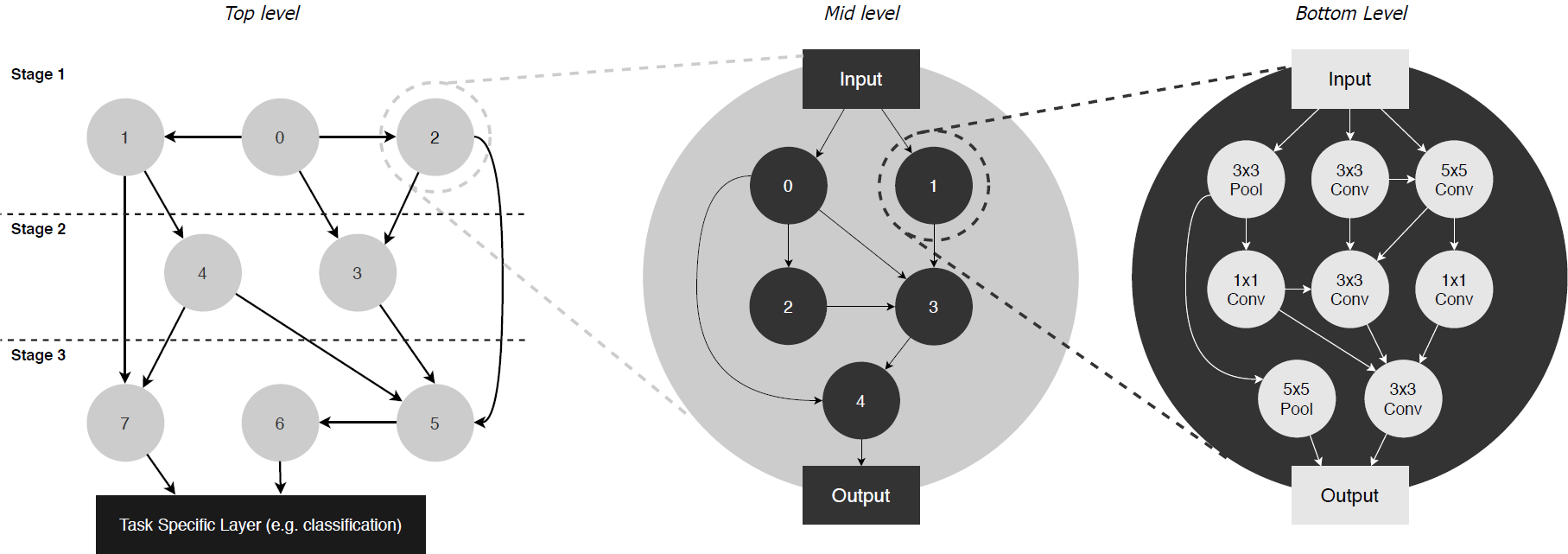
Neural Architecture Generator Optimization (NAGO) casts NAS as a problem of finding the optimal network generator and proposes a new, hierarchical and graph-based search space capable of representing an extremely large variety of network types, yet only requiring few continuous hyper-parameters. This greatly reduces the dimensionality of the problem, enabling the effective use of Bayesian optimisation as a search strategy. At the same time, we expand the range of valid architectures, motivating a multi-objective learning approach.
Our network search space is modelled as a hierarchical graph with three levels. At the top-level, we have a graph of cells. Each cell is itself represented by a mid-level graph. Similarly, each node in a cell is a graph of basic operations (conv3x3, conv5x5, etc.). This results in 3 sets of graph hyperparameters:
X_{top}, X_{mid}, X_{bottom}, each of which independently defines the graph generation model in each level.
Following Xie et al. (2019) we use the Watts-Strogatz (WS) model as the random graph generator for the top and bottom levels, with hyperparameters X_{top}=[N_{t}, K_{t}, P_{t}] and X_{bottom}=[N_{b}, K_{b}, P_{b}]; and use the Erdos-Renyi (ER) graph generator for the middle level, with hyperparameters X_{mid}=[N_{m}, P_{m}], to allow for the single-node case.
By varying the graph generator hyperparameters and thus the connectivity properties at each level, we can produce a extremely diverse range of architectures. For instance, if the top-level graph has 20 nodes arranged in a feed-forward configuration and the mid-level graph has a single node, then we obtain networks similar to those sampled from the DARTS search space. While if we fix the top-level graph to 3 nodes, the middle level to 1 and the bottom-level graph to 32, we can reproduce the search space from (Xie et al., 2019).
Our proposed hierarchical graph-based search space allows us to represent a wide variety of neural architectures with a small number of continuous hyperparameters, making NAS amenable to a wide range of powerful BO methods. In this code package, we use the multi-fidelity BOHB approach, which uses partial evaluations with smaller-than-full budget in order to exclude bad configurations early in the search process, thus saving resources to evaluate more promising configurations and speeding up optimisation. Given the same time constraint, BOHB evaluates many more configurations than conventional BO which evaluates all configurations with full budget. Please refer to hpo.md for more details on BOHB algorithm.
Please cite our paper if you would like to use the code
@article{ru2020neural,
title={Neural Architecture Generator Optimization},
author={Ru, Binxin and Esperanca, Pedro and Carlucci, Fabio},
journal={arXiv preprint arXiv:2004.01395},
year={2020}
}
For a Chinese version of the document on NAGO, please refer to nago.md. For more details on the VEGA package, please refer to VEGA_README.md.