Message passing based decoders for LDPC codes with Python/NumPy. Includes implementations of
- min-sum (MSA) and sum-product (SPA) algorithms using sparse matrices (scipy.sparse)
- maximum-likedlood (ML) and linear-programming (LP) decoders (only for short length codes like Hamming(7,4)) based on Using Linear Programming to Decode Binary Linear Codes
- ADMM decoder based on Decomposition Methods for Large Scale LP Decoding
for binary erasure (BEC), binary symmetric (BSC) and binary-AWGN (biawgn) channels.
Following Python/package versions (or higher) are required.
Python version 3.5.2numpy version 1.12.0scipy version 0.18.1
Checkout assests branch to see all pre-compupted results. These include different codes, simulation results and plots.
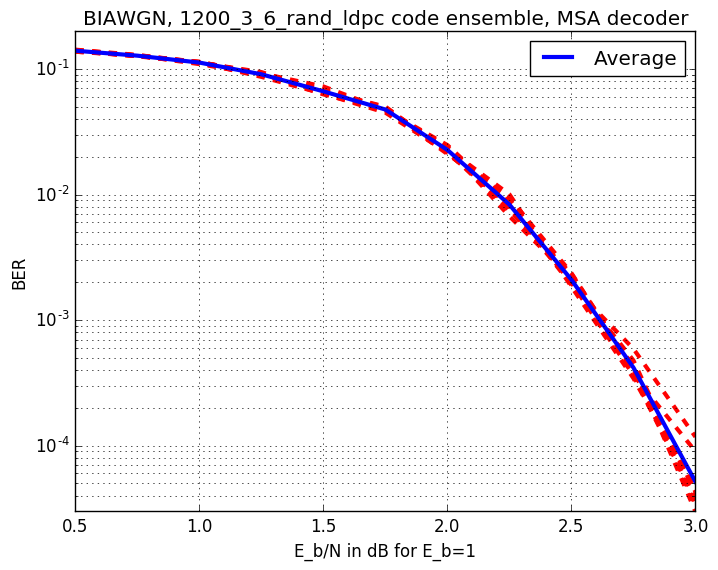
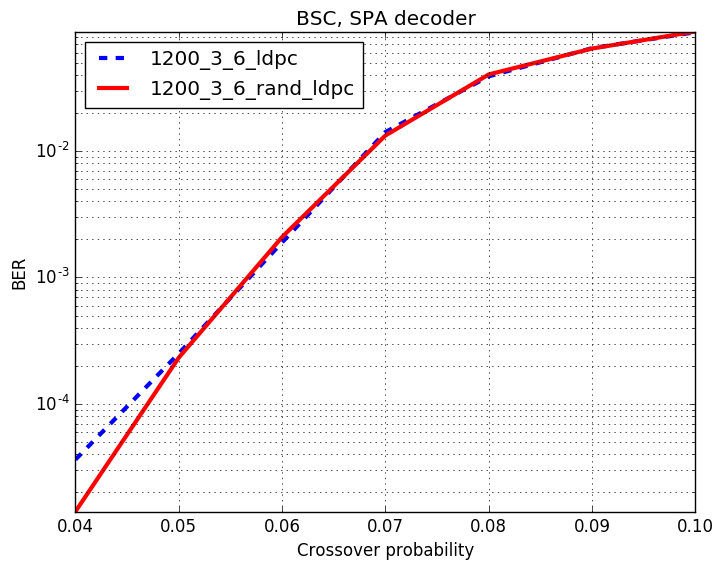
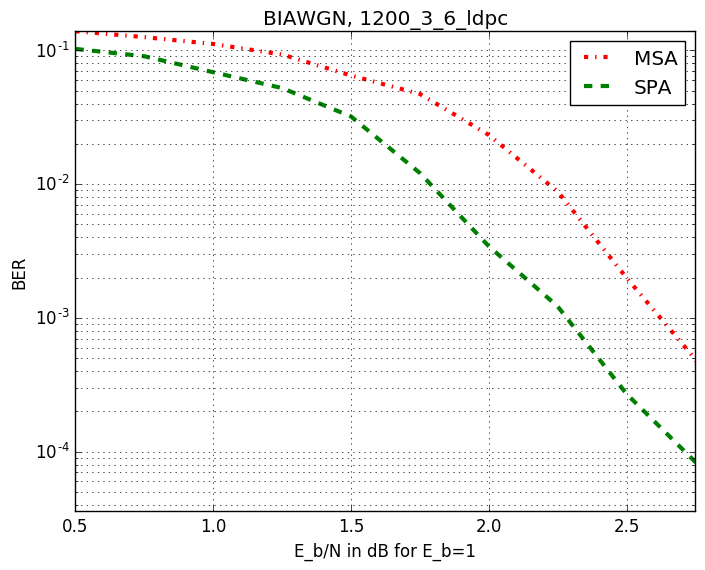
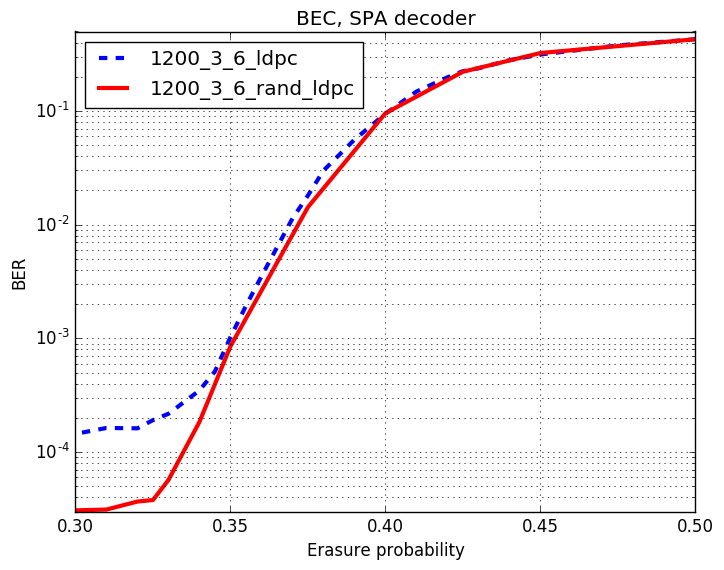
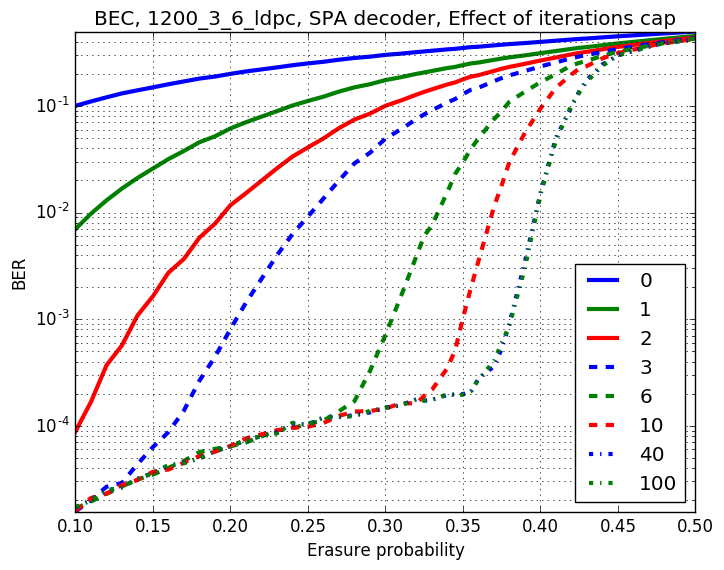
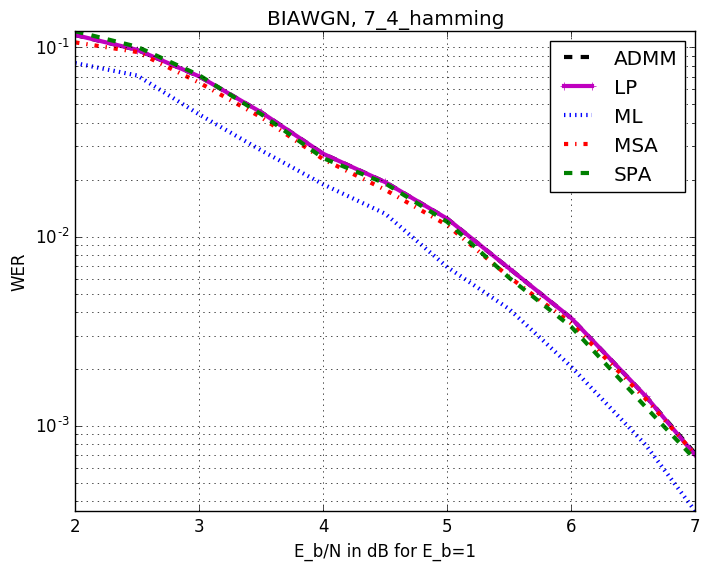
-
Make directories
codesanddatain the root directory. -
Execute
python src/codes.py 10 1200 3 6to generate 10 random samples fromLDPC(1200,3,6)ensemble. -
Run simulations using
run_sims.sh {PARA} {CASE} {ARGS}command. For example,./run_sims.sh SEQL HMGexecutes all Hamming code related simulations sequentially../run_sims.sh SEQL REG_ENS --data-dir=./data --consoleexecutes some regular LDPC related simulations sequentially while printing logs onto console../run_sims.sh PARA REG_ENS --data-dir=./dataexecutes the same in parallel.- Use the latter only on a dedicated server as it will take large amount of CPU.
- See
run_sims.shfor other choices of{CASE}.
-
If running the simulations on Niagara cluster, need to setup environment first by executing
setup_env.sh. -
Execute
./run_sims.sh PARA HMG --data-dir=$SCRATCHto test if simulations run properly. -
All simulations can be submitted for later excution on Niagara by using command
sbatch niagara/submit_job.sh.
- Make directory
plotsin the root directory. - Execute
./plot_results.py HMGto view and save Hamming code related plots in PNG format. - Execute
./plot_results.py REG_ENS --ext=pdfto view and save regular ensemble related plots in PDF format. - Execute
./plot_results.py HMG REG_ENS --ext=png --data-dir=./data --plots-dir=./plots --silent --error=berto silently save both Hamming code related and regular ensemble related bit-error-rate plots. - If running on the Niagara cluster execute
./plot_results.py HMG REG_ENS --data-dir=$SCRATCH --plots-dir=$SCRATCH --silent --aggto use the proper back-end formatplotlib.
python src/main.py bec 1200_3_6_rand_ldpc_1 SPA --codeword=1 --console --params .5 .475 .45 .425 .4 .375 .35 .325 .3python src/main.py biawgn 7_4_hamming SPA --codeword=1 --params .01 .05 .1 .5 1 2 4 6 --console- See
run_sims.shandsimulations.pyfor more.
python src/stats.py bec 1200_3_6_rand_ldpc SPA
python src/graph.py bec 1200_3_6_ldpc SPA single --error berpython src/graph.py bsc 7_4_hamming SPA ML comp_dec --error wer- See
plot_results.pyfor more.
First of the following plots the density evolution while the second generates the optimal irregular distribution a given check node distribution (rho).
python src/ldpc.py pltpython src/ldpc.py irg --count=10 --len=1200 --rho=5 --rate=.5
Execute the following to reproduce the Figure 50.4 (histogram for LT code with length 10000) in Information Theory, Inference, and Learning Algorithms by David J.C. MacKay. Replace in the first command with 0.01, 0.03, 0.1.
python -u src/luby.py 10000 12000 <c> .5 250 --pool=4python src/luby_graph.py .01 .03 .1