Fast multi-dimensional histogram with convenient interface for C++11 and Python
This C++11 library provides an easy-to-use powerful n-dimensional histogram class for your statistics needs. It is very customisable through policy classes, but the default policies were carefully designed so that most users won't need to customize anything. The library has a convenient uniform interface, is memory efficient, and very fast. If the default policies are used, bin counts cannot overflow or loose precision.
The histogram class comes in two implementations with a common interface. The static variant uses compile-time information to provide maximum performance, at the cost of potentially larger executables and reduced runtime flexibility. The dynamic variant makes the opposite trade-off. Python bindings for the latter are included, implemented with boost.python.
The histogram supports value semantics. Move operations and trips over the language boundary from C++ to Python are cheap. Histogram instances can be streamed from/to files and pickled in Python. Numpy is supported to speed up operations in Python: histograms can be filled with Numpy arrays at high speed (faster than numpy's own histogram functions) and are convertible into Numpy arrays without copying data.
My goal is to submit this project to Boost, that's why it uses the Boost directory structure and namespace. The code is released under the Boost Software License.
Check out the full documentation. Highlights are given below.
- N-dimensional histogram
- Simple and convenient interface
- Value semantics with efficient move operations
- Support for various binning schemes (user-extensible)
- Optional underflow/overflow bins for each dimension
- Counts cannot overflow or loose precision (+)
- Support for weighted input
- Statistical variance can be queried for each bin
- High performance (see benchmarks)
- Efficient use of memory (dynamically grows as needed)
- Serialization support using
boost.serialization - Language support: C++11, Python 2.x and 3.x
- Numpy support
(+) In the standard configuration and if you don't use weighted input.
The library can be build with b2 within the boost directory structure, but if you are not a boost developer, use cmake instead.
git clone https://github.com/HDembinski/histogram.git
mkdir build && cd build
cmake ../histogram/build
make # or 'make install'To run the tests, do make test.
For the full version of the following examples with explanations, see Tutorial.
Example 1: Fill a 1d-histogram in C++
#include <boost/histogram/histogram.hpp> // proposed for inclusion in Boost
#include <iostream>
#include <cmath>
int main(int, char**) {
namespace bh = boost::histogram;
// create 1d-histogram with 10 equidistant bins from -1.0 to 2.0,
// with axis of histogram labeled as "x"
auto h = bh::make_static_histogram(bh::regular_axis<>(10, -1.0, 2.0, "x"));
// fill histogram with data
h.fill(-1.5); // put in underflow bin
h.fill(-1.0); // included in first bin, bin interval is semi-open
h.fill(-0.5);
h.fill(1.1);
h.fill(0.3);
h.fill(1.7);
h.fill(2.0); // put in overflow bin, bin interval is semi-open
h.fill(20.0); // put in overflow bin
h.wfill(5.0, 0.1); // fill with a weighted entry, weight is 5.0
// iterate over bins, loop includes under- and overflow bin
for (const auto& bin : h.axis<0>()) {
std::cout << "bin " << bin.idx
<< " x in [" << bin.left << ", " << bin.right << "): "
<< h.value(bin.idx) << " +/- " << std::sqrt(h.variance(bin.idx))
<< std::endl;
}
/* program output:
bin -1 x in [-inf, -1): 1 +/- 1
bin 0 x in [-1, -0.7): 1 +/- 1
bin 1 x in [-0.7, -0.4): 1 +/- 1
bin 2 x in [-0.4, -0.1): 0 +/- 0
bin 3 x in [-0.1, 0.2): 5 +/- 5
bin 4 x in [0.2, 0.5): 1 +/- 1
bin 5 x in [0.5, 0.8): 0 +/- 0
bin 6 x in [0.8, 1.1): 0 +/- 0
bin 7 x in [1.1, 1.4): 1 +/- 1
bin 8 x in [1.4, 1.7): 0 +/- 0
bin 9 x in [1.7, 2): 1 +/- 1
bin 10 x in [2, inf): 2 +/- 1.41421
*/
}Example 2: Fill a 2d-histogram in Python with data in Numpy arrays
import histogram as bh
import numpy as np
# create 2d-histogram over polar coordinates, with
# 10 equidistant bins in radius from 0 to 5 and
# 4 equidistant bins in polar angle
h = bh.histogram(bh.regular_axis(10, 0.0, 5.0, "radius",
uoflow=False),
bh.circular_axis(4, 0.0, 2*np.pi, "phi"))
# generate some numpy arrays with data to fill into histogram,
# in this case normal distributed random numbers in x and y,
# converted into polar coordinates
x = np.random.randn(1000) # generate x
y = np.random.randn(1000) # generate y
rphi = np.empty((1000, 2))
rphi[:, 0] = (x ** 2 + y ** 2) ** 0.5 # compute radius
rphi[:, 1] = np.arctan2(y, x) # compute phi
# fill histogram with numpy array
h.fill(rphi)
# access histogram counts (no copy)
count_matrix = np.asarray(h)
print count_matrix
# program output:
#
# [[37 26 33 37]
# [60 69 76 62]
# [48 80 80 77]
# [38 49 45 49]
# [22 24 20 23]
# [ 7 9 9 8]
# [ 3 2 3 3]
# [ 0 0 0 0]
# [ 0 1 0 0]
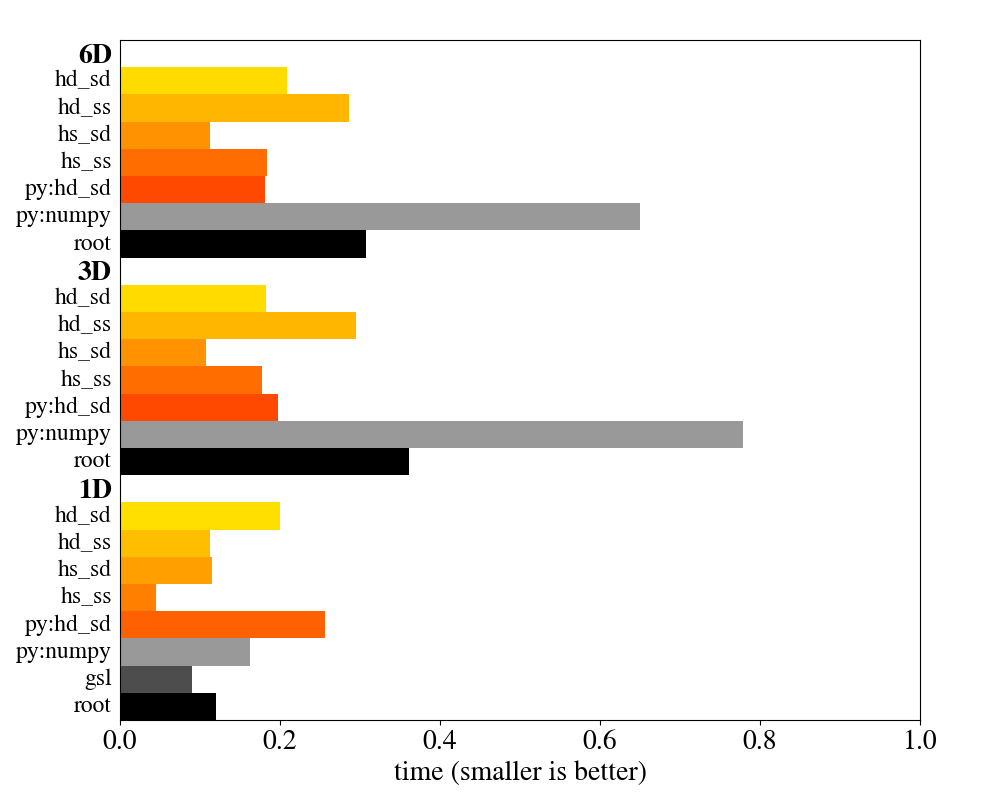
# [ 0 0 0 0]]Thanks to modern meta-programming and intelligent memory management, this library is not only more flexible and convenient to use, but also faster than the competition. In the plot below, its speed is compared to classes from the ROOT framework and to Numpy. The orange to red items are different compile-time configurations of the histogram in this library. More details on the benchmark are given in the documentation
There is a lack of a widely-used free histogram class. While it is easy to write an 1-dimensional histogram, writing an n-dimensional histogram poses more of a challenge. If you add serialization and Python/Numpy support onto the wish-list, the air becomes thin. The main competitor is the ROOT framework. This histogram class is designed to be more convenient to use, and as fast or faster than the equivalent ROOT histograms. It comes without heavy baggage, instead it has a clean and modern C++ design which follows the advice given in popular C++ books, like those of Meyers and Sutter and Alexandrescu.
Read more about the rationale of the design choices in the documentation
The histogram is feature-complete. More than 500 individual tests make sure that the implementation works as expected. Comprehensive documentation is available. User feedback is appreciated!