This is the repository of the SAN paper, found here:
http://ecai2020.eu/papers/1721_paper.pdf (please, cite if you are using it!). Note that the full code with datasets to reproduce the paper can be found here: https://gitlab.com/skblaz/attentionrank (the code is in benchmark-ready form). The purpose of this repository is to provide all functionality in a user-friendly way. Disclaimer: this code was not extensively benchmarked and can contain bugs. If you find one, please open an issue.
python setup.py install
or
pip install git+https://github.com/https://github.com/SkBlaz/san
A simple usecase is given next:
from scipy import sparse
import numpy as np
from sklearn.datasets import load_breast_cancer
import san
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.feature_selection import chi2,f_classif,mutual_info_classif
from sklearn.ensemble import RandomForestClassifier
sns.set_style("whitegrid")
dataobj = load_breast_cancer()
X = dataobj['data']
Y = dataobj['target']
names = dataobj['feature_names']
# let's overfit, just for demo purposes
clf = san.SAN(num_epochs = 32, num_heads = 2, batch_size = 8, dropout = 0.2, hidden_layer_size = 32)
X = sparse.csr_matrix(X)
clf.fit(X, Y)
preds = clf.predict(X)
global_attention_weights = clf.get_mean_attention_weights()
local_attention_matrix = clf.get_instance_attention(X)
mutual_information = mutual_info_classif(X,Y)
rf = RandomForestClassifier().fit(X,Y).feature_importances_
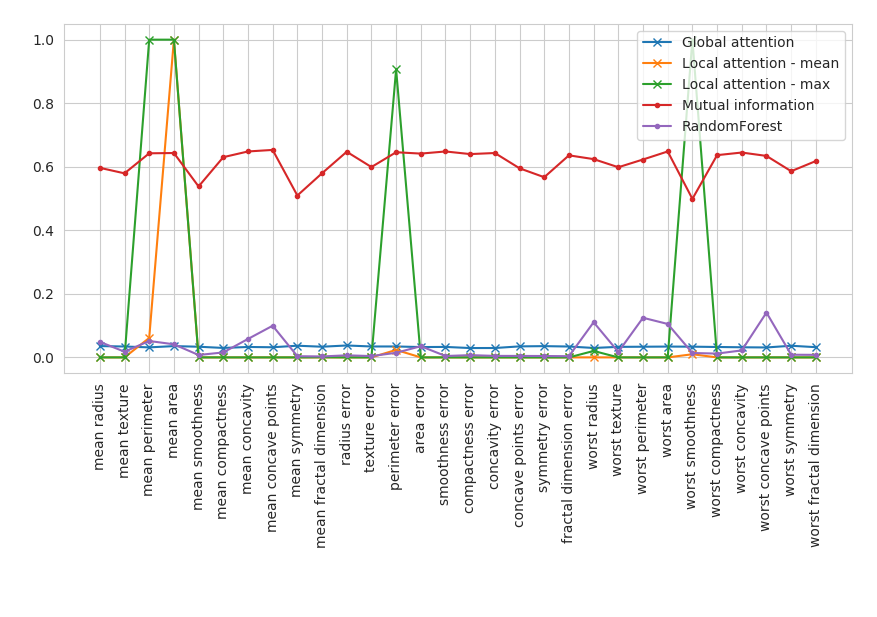
plt.plot(names, global_attention_weights, label = "Global attention", marker = "x")
plt.plot(names, np.mean(local_attention_matrix, axis = 0), label = "Local attention - mean", marker = "x")
plt.plot(names, np.max(local_attention_matrix, axis = 0), label = "Local attention - max", marker = "x")
plt.plot(names, mutual_information, label = "Mutual information", marker = ".")
plt.plot(names, rf, label = "RandomForest", marker = ".")
plt.legend(loc = 1)
plt.xticks(rotation = 90)
plt.tight_layout()
plt.show()
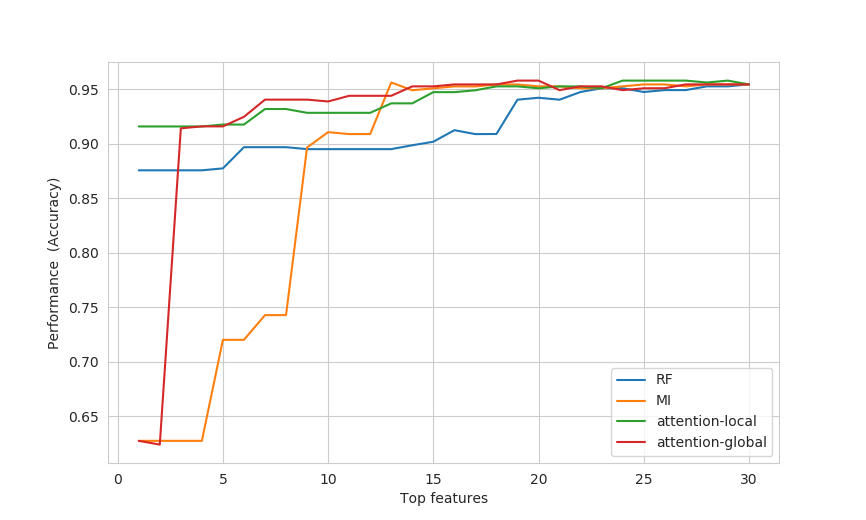
Example mock evaluation is shown below (examples/example_benchmark.py):