TBtools is a Toolset written by CJ-Chen for wet-lab biologists.It aims at facilitating some common and elaborate but simple molecular analysis tasks such as sequences extract, gene set functional enrichment It is easy to use and can run in all Operating system including Mac, Linux, Windows, which has Java run time environment higher than 1.6 . However, TBtools may mainly focus on data analysis procedure which make it difficult to plot fancy figures as well as being publishable. To cover the shortage, we implement an interactive plot websever using shiny programing env, and named TBploter
Click here to redirect to TRploter website.
Screen shoot of TBploter
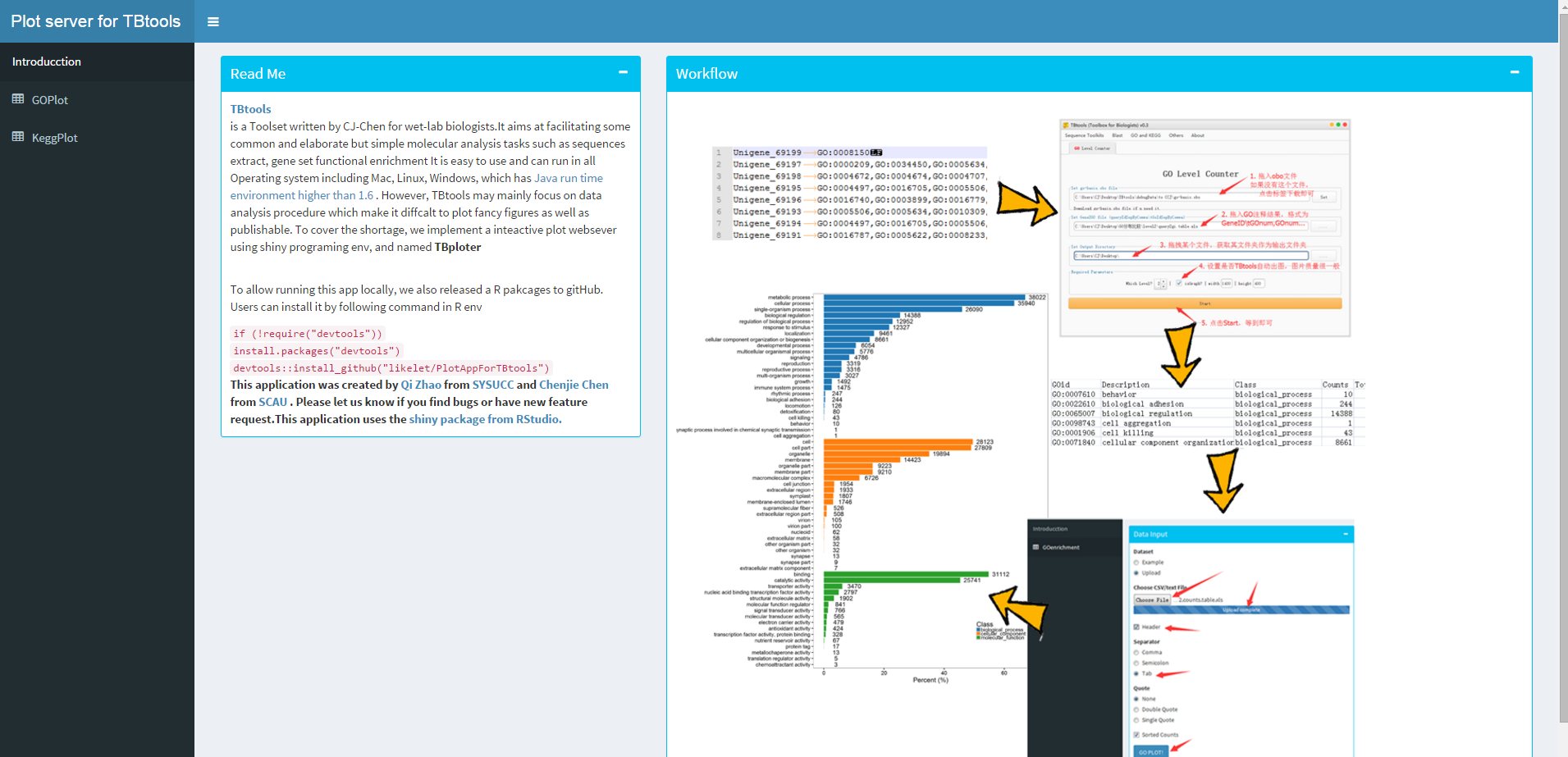
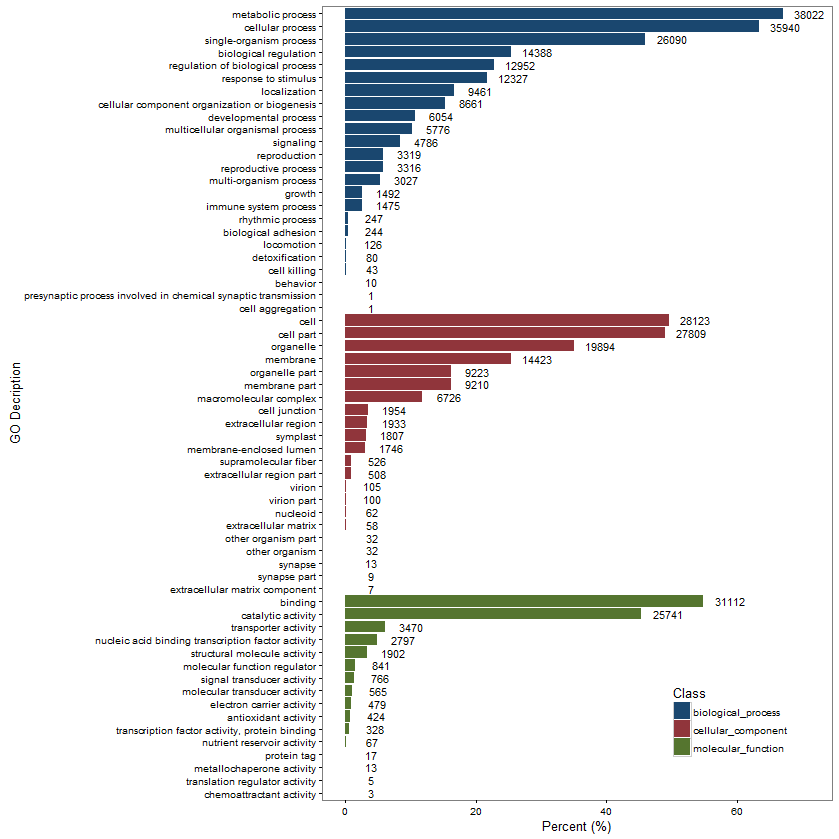
GOenriment ploter
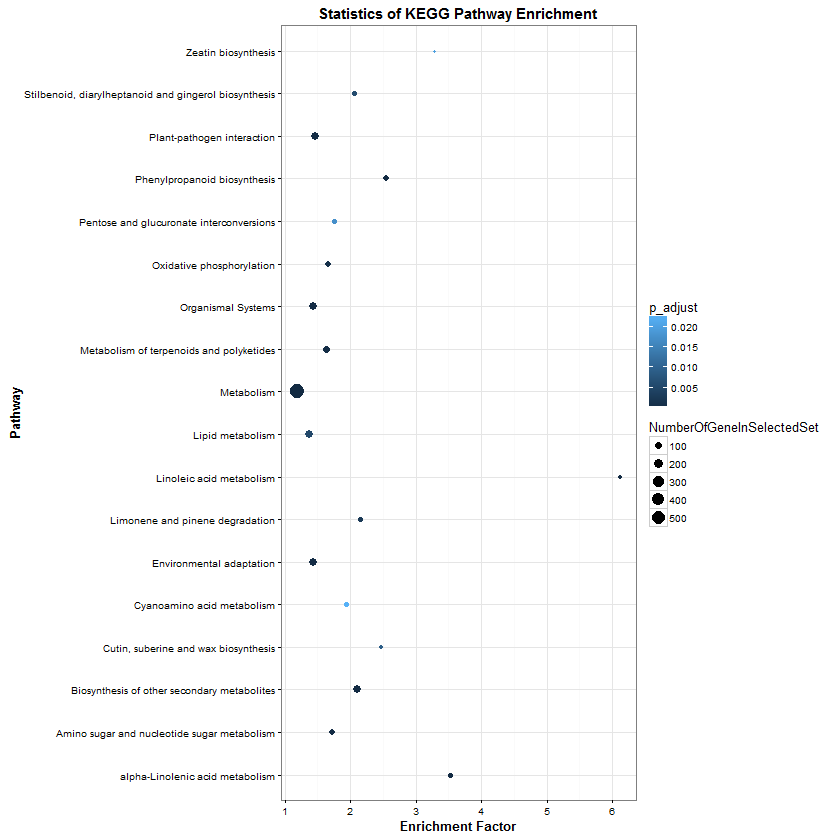
 KEGGenrichment ploter
KEGGenrichment ploter
To check the dependencies installed correctly, this command can help users to check the status of each installation
library("Packages for check")Code for install dependencies R packages
cDep <- c("ggplot2","shiny","shinyBS","shinydashboard","ggthemes","DT")
###INSTALLED PACKAGES
#get installed list
inst <- packageStatus()$inst
#check and install DEPENDENCIES from CRAN
for(i in 1:length(cDep)){
tag = which(inst$Package == cDep[i])
if(length(tag)){
remove.packages(cDep[i])
}
install.packages(cDep[i])
}
To install the latest development build directly from GitHub, run this:
if (!require("devtools"))
install.packages("devtools")
devtools::install_github("likelet/PlotAppForTBtools")Qi Zhao, zhaoqi3@mail2.sysu.edu.cn
Qi Zhao, zhaoqi3@mail2.sysu.edu.cn
Chenjie Chen,120509419@qq.com
Qi Zhao
Please feel free contact us.
MIT license
during preparation