GERNERMED - An Open German Medical NER Model
Note: The project repository will be updated upon acceptance.
About
The is the project repository for GERNERMED, a named entity recognition (NER) model in the context of German medical natural language processing (NLP).
In particular, GERNERMED is the first open neural NER model for medical entities designed for German data.
The following entities are supported:
- Drug
- Strength
- Route
- Form
- Dosage
- Frequency
- Duration
The evaluation scores on the test set are as follows:
| NER Tag | Precision | Recall | F1-Score |
|---|---|---|---|
| Drug | 67.33 | 66.17 | 66.74 |
| Strength | 92.34 | 90.99 | 91.66 |
| Route | 89.93 | 90.14 | 90.04 |
| Form | 91.94 | 89.24 | 90.57 |
| Dosage | 87.83 | 87.57 | 87.70 |
| Frequency | 79.14 | 76.92 | 78.01 |
| Duration | 67.86 | 52.78 | 59.37 |
| total | 82.31 | 80.79 | 81.54 |
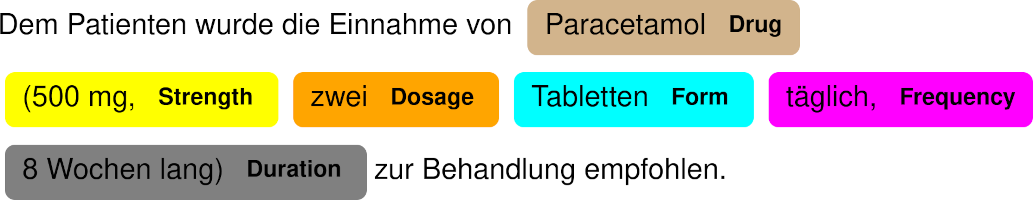
NER Demonstration
Setup and Usage
For a quick test, run python3 example.py to run the NER model pipeline.
The package NER pipeline is available in /data/de_GERNERMED-1.0.0.tar.gz and can be installed by:
# From local fs
python3 -m pip install ./data/de_GERNERMED-1.0.0.tar.gz
# From GitHub
python3 -m pip install https://github.com/frankkramer-lab/GERNERMED/blob/main/data/de_GERNERMED-1.0.0.tar.gz?raw=trueThe pipeline can be used in Python:
import spacy
nlp = spacy.load("de_GERNERMED")
doc = nlp("Dem Patienten wurde die Einnahme von Paracetamol (500 mg, zwei Tabletten täglich, 8 Wochen lang) zur Behandlung empfohlen.")
# Show entities
print(doc.ents)Reproducibility
The evaluation scores on the testset can be obtained by ./run_eval.sh.
The custom German dataset is stored in /data/GERNERMED_dataset.json.
Citation
Please cite the following ArXiv paper from http://arxiv.org/abs/2109.12104
@misc{frei2021gernermed,
title={GERNERMED -- An Open German Medical NER Model},
author={Johann Frei and Frank Kramer},
year={2021},
eprint={2109.12104},
archivePrefix={arXiv},
primaryClass={cs.CL}
}