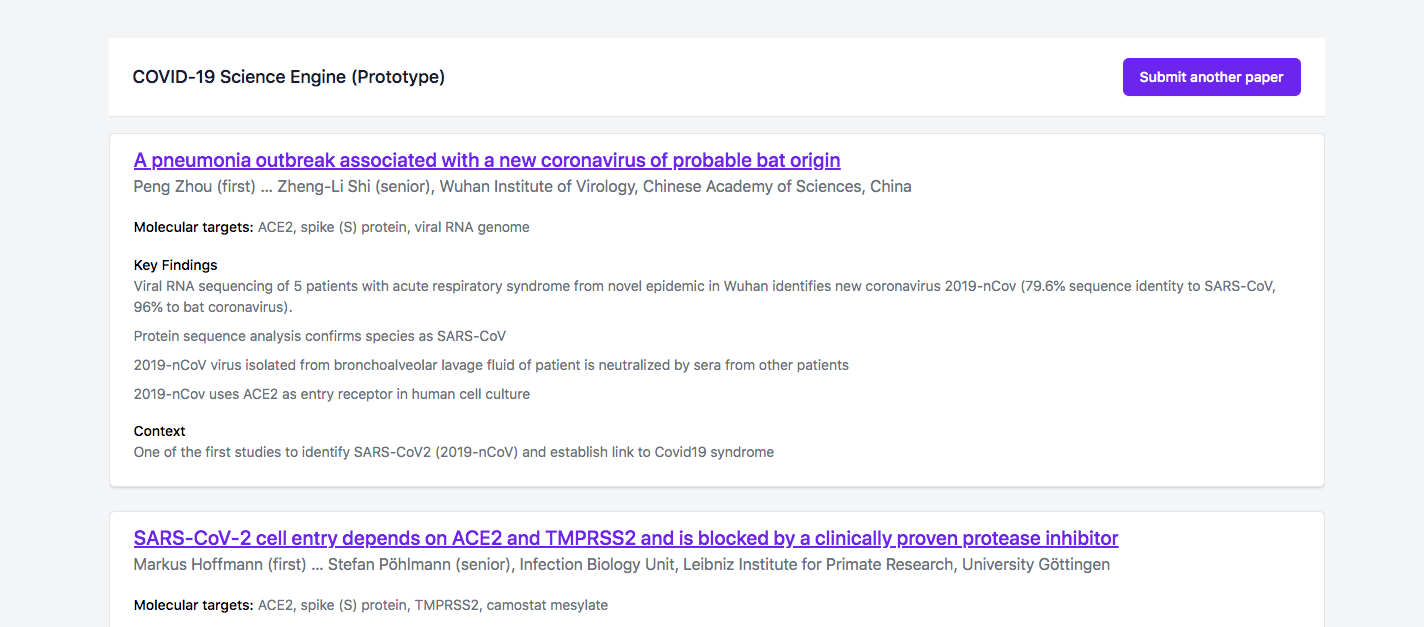
A tool for researchers to quickly understand and contextualize Covid19 research papers.
Biomedical researchers have to painstakingly search for relevant information in thousands of papers and in each paper in a lot of irrelevant text.
We want to use a combination of human and machine intelligence to accelerate this process drastically. We are building an NLP algorithm that is able to extract meaningful information and contextualize it. In the meantime, we are allowing information to be manually submitted so we can build a useful database of papers.
Summaries and metadata can be manually submitted (this will be automated later)
The prototype is available at https://covid-science-engine-prototype.herokuapp.com/ (currently password protected)
The stack is:
- Ruby on Rails 6.0
- React (with TypeScript)
- Tailwind CSS
- Postgres
- Heroku
Install and start postgresql:
- On macOS, you can use
pg_ctl -D /usr/local/var/postgres start - (To stop postgres use
pg_ctl -D /usr/local/var/postgres stop)
Install dependencies:
bundle install
yarn install
Setup the database and seed data:
rails db:setup
You can use your editor of choice but please make sure to reformat your code with Prettier according to the .prettierrc file.
The following environment variables can be set:
| Environment variable | Description |
|---|---|
AUTH_USERNAME |
Username for basic authentication |
AUTH_PASSWORD |
Password for basic authentication |
rails server
Then go to http://localhost:3000 to view the app.
Currently, we have some basic authentication in front of the website. We will be replacing this with something better soon.
In development, the username and password is foo / bar.
rails spec
We are looking for new contributors to help us build this project.
👉 Please read our Contributing guide to get started. 👈
You can also join our Discord server to get in touch.
This code uses the MIT license.