FUSegNet and x-FUSegNet are implemented on top of qubvel's implementation.
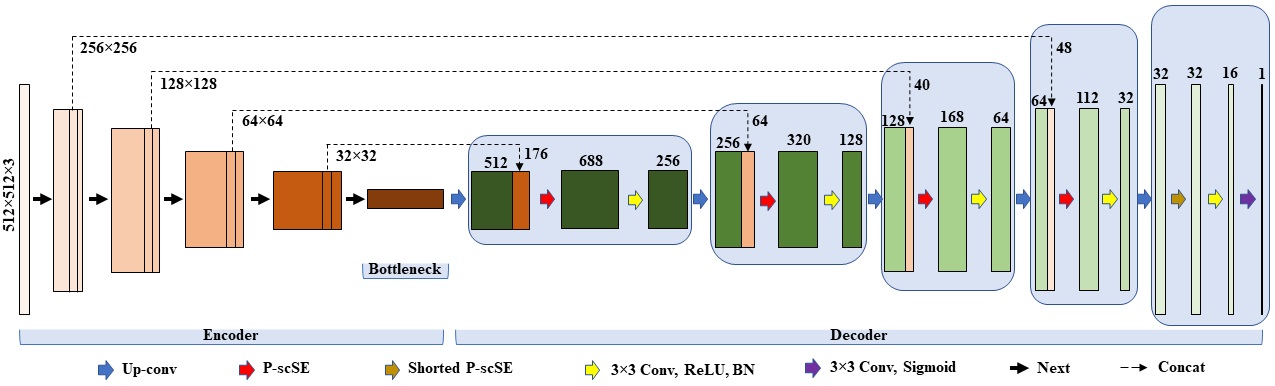
FUSegNet is a novel model for foot ulcer segmentation in diabetes patients. The model introduces the parallel scSE (P-scSE) module, combining additive and max-out scSE, fused in the middle of each decoder stage. FUSegNet achieves a data-based dice score of 92.70% on a chronic wound dataset, outperforming other state-of-the-art models. In the MICCAI 2021 FUSeg Challenge, the submitted x-FUSegNet model achieves a top score of 89.23%, leading the leaderboard.
Preprint link.
Our saved (trained) models can be downloaded from the following links-
- utils
|--category.py: Lists AZH Chronic wound test imgaes into 10 categories. Categories are created based on %GT area in images. Categorized test image names are stored in a json file called categorized_oldDfu.json
|--eval.py: Performs data-based evaluation.
|--eval_categorically.py: Performs data-based evaluation for each category.
|--eval_boxplot.py: Performs image-based evaluation for each category that is required for boxplot. The final output is an excel file with multiple sheets. Each sheet stores results for a perticular category.
|--boxplot.py: Creates a boxplot. It utilizes the excel file generated byeval_boxplot.py.
|--contour.py: Draws contours around the wound region.
|--runtime_patch.py: Creates patch during runtime. fusegnet_all.py: It's an end-to-end file contains codes for dataloader, training and testing using the FUSegNet model.fusegnet_train.py: It is to train a dataset using the FUSegNet model.fusegnet_test.py: It is to perform inference using the FUSegNet model.xfusegnet_all.py: It's an end-to-end file contains codes for dataloader, training and testing using the xFUSegNet model.xfusegnet_train.py: It is to train a dataset using the xFUSegNet model.xfusegnet_test.py: It is to perform inference using the xFUSegNet model.FUSegNet_feature_visualization.ipynb: Demonstrates intermediate features.
pip install -r requirements.txt
Note: It is better to install torch and its associated packages manually as these are very sensitive to hardware and the OS.
The torch packages mentioned in the requirements.txt file are used for a 64-bit Ubuntu PC with an 8-core 3.4 GHz CPU and a single NVIDIA RTX 2080Ti GPU with a CUDA compilation version of 10.1.
- Proposed FUSegNet overview
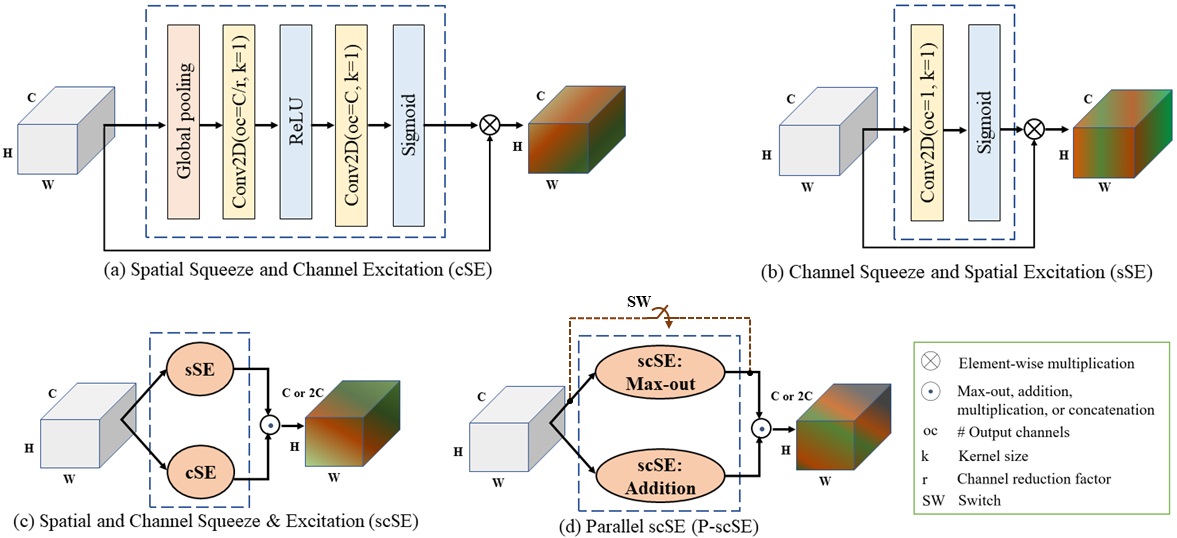
- Proposed Parallel scSE (P-scSE) module
The directory structure is shown below. Note that if checkpoints, plots, and predictions folders are not created beforehand, they will be generated automatically.
.
|-- fusegnet_all.py
|-- fusegnet_train.py
|-- fusegnet_test.py
|-- xfusegnet_all.py
|-- xfusegnet_train.py
|-- xfusegnet_test.py
|-- utils
|-- dataset
|-- train
|-- images
|-- (training and validation images are kept here)
|-- labels
|-- (training and validation labels are kept here)
|-- test
|-- images
|-- (test images are kept here)
|-- labels
|-- (test labels are kept here)
|-- checkpoints
|-- (models will be stored here)
|-- plots
|-- (loss curves will be stored here)
|-- predictions
|-- (model predictions will be store here)
fusegnet_all.py, fusegnet_train.py, xfusegnet_all.py, and xfusegnet_train.py have a section called Parameters where the user can set the model parameters. The following are the model parameters used to train FUSegNet and xFUSegNet.
BASE_MODEL = 'FuSegNet' # give any name for the model
ENCODER = 'efficientnet-b7' # encoder model
ENCODER_WEIGHTS = 'imagenet' # encoder weights
BATCH_SIZE = 2 # no. of batches
IMAGE_SIZE = 224 # height and width
n_classes = 1 # no. of classes excluding background
ACTIVATION = 'sigmoid' # output activation. sigmoid for binary and softmax for multi-class segmentation
DEVICE = torch.device("cuda" if torch.cuda.is_available() else "cpu") # sets gpu if available
LR = 0.0001 # learning rate
EPOCHS = 200 # no. of epochs
WEIGHT_DECAY = 1e-5 # for L2 penalty
SAVE_WEIGHTS_ONLY = True # if True, saves weights only
TO_CATEGORICAL = False # if True, converts to onehot
SAVE_BEST_MODEL = True # if True, saves the best model only
SAVE_LAST_MODEL = False # if True, saves the model after completing the training
PERIOD = None # periodically save checkpoints
RAW_PREDICTION = False # if true, then stores raw predictions (i.e. before applying threshold)
PATIENCE = 30 # no. of epoches waits before early stopping
EARLY_STOP = True # if True, enables early stoppingMode: end-to-end
- In this mode, training and inference codes are embedded in a single .py file. *
fusegnet_all.pyandxfusegnet_all.pyfiles are written in this mode.- Once the model parameters are set in the
Parametersection, the user can run (train, validation, and test) using the following commands -python fusegnet_all.pyorpython xfusegnet_all.py. fusegnet_all.pyandxfusegnet_all.pycan directly be run from any IDE (e.g. Spyder, PyCharm, Jupyter Notebook, etc.)
Mode: train only
- In this mode, only training code is embedded in the .py file.
fusegnet_train.pyandxfusegnet_train.pywork in this mode.- Once the model parameters are set in the
Parametersection, the user can train the model using the following commands -python fusegnet_train.pyorpython xfusegnet_train.py. fusegnet_train.pyandxfusegnet_train.pycan directly be run from any IDE (e.g. Spyder, PyCharm, Jupyter Notebook, etc.)
Mode: test (inference) only
- In this mode, only inference code is embedded in the .py file.
fusegnet_test.pyandxfusegnet_test.pywork in this mode.- The user can test the model using the following commands -
python fusegnet_test.pyorpython xfusegnet_test.py. fusegnet_train.pyandxfusegnet_train.pycan directly be run from any IDE (e.g. Spyder, PyCharm, Jupyter Notebook, etc.).- To test with our saved models, put the saved models in the
checkpointsdirectory and then perform either one of the above two steps.
Mode: feature visualization
- In this mode, intermediate feature maps are visualized.
FUSegNet_feature_visualization.ipynbdemonstrates the output feature maps of the parallel scSE (P-scSE) modules and each decoder stage.
Currently, our implementation supports the following SEs:
pscse: Parallel spatial and channel squeeze-and-excitationscse: Spatial and channel squeeze-and-excitationmaxout: max(cSE, sSE)additive: cSE + sSEconcat: concatenate(cSE, sSE)multiplication: cSE * sSEaverage: mean(stack(cSE, sSE))average-all: mean(stack(maxout, additive, concat, multiplication))
The user needs to pass the attention type to decoder_attention_type in fusegnet_all.py, fusegnet_train.py, xfusegnet_all.py, or xfusegnet_train.py. For instance,decoder_attention_type = 'pscse'
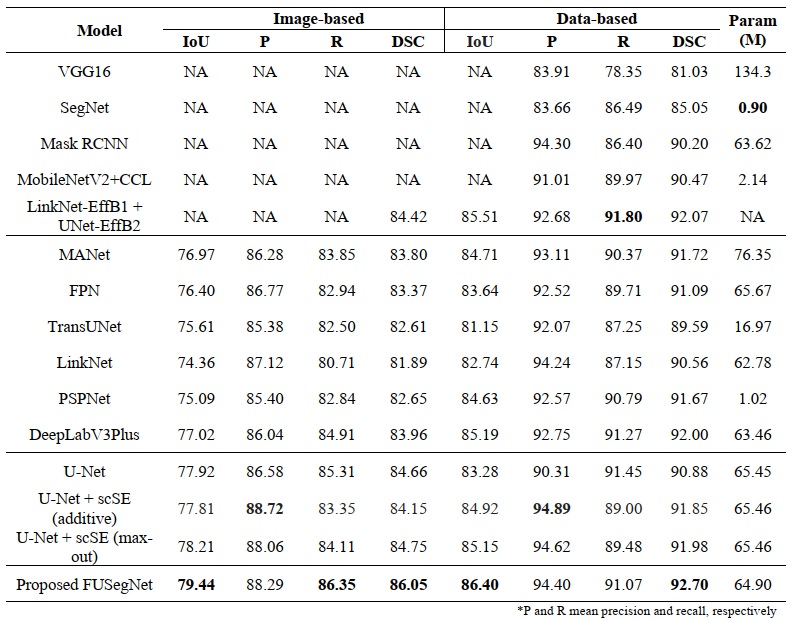
- Segmentation results on the Chronic Wound dataset
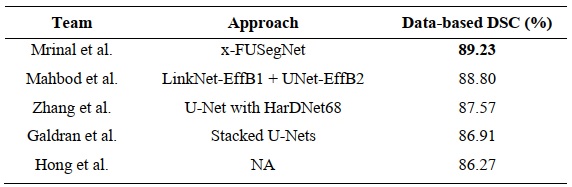
- Top five performers of the MICCAI FUSeg Challenge 2021
[1] Pavel Iakubovskii, "Segmentation Models Pytorch", GitHub repository, GitHub, 2019. URL: https://github.com/qubvel/segmentation_models.pytorch
[2] C. Wang et al., “Fully automatic wound segmentation with deep convolutional neural networks,” Sci. Rep., vol. 10, no. 1, 2020.
[3] MICCAI FUSeg Challenge 2021. URL: https://fusc.grand-challenge.org/