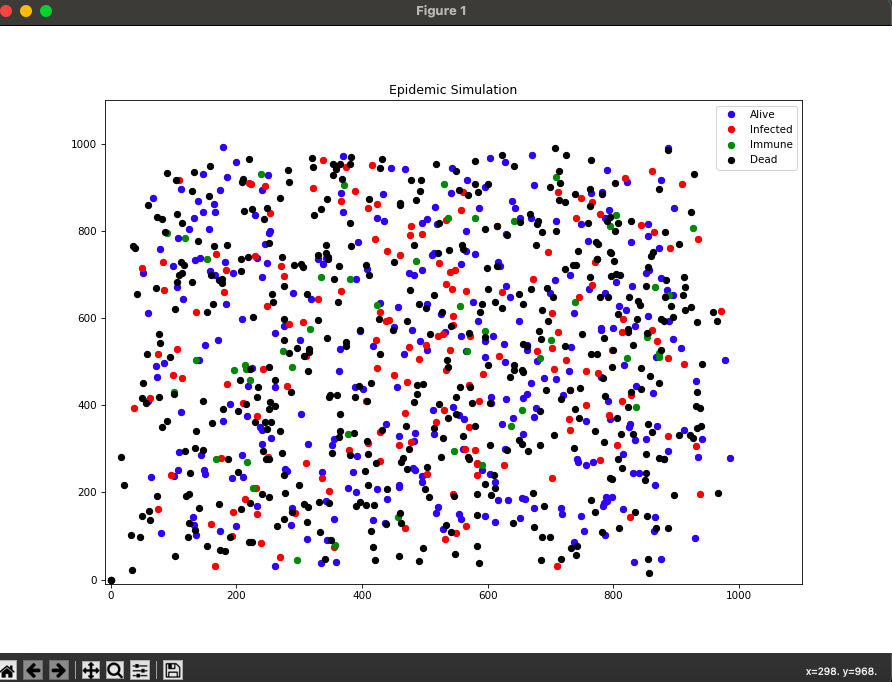
This is a small simulation of an epidemic inside a mobile population. It is implemented in C++ and is not to be used as representing real epidemics.
The Simulation spawns a number of People with common attributes:
struct Person
{
// Status
bool alive; // 0 if dead
bool infected; // 1 if true
bool immune; // 1 if true;
int days_infected = 0;
Person() : alive(true), infected(false), immune(false), days_infected(0), xpos(rand()%1000), ypos(rand()%1000){};
// Position
double xpos = rand()%1000;
double ypos = rand()%1000;
};
Everything happens inside a 1000x1000 square plane.
Each Person gets a random starting point, from which they move in a random walk (implemented with std::mt19937). If they get too close to the "walls", they are moved to a random point inside the plane.
If a person gets too close to an infected person, they get infected.
For each timestep, there is a chance SPONT_INF of spontaneous infection. If the person is infected for more than 14 timesteps (days), they get immune; if they are infected, they can also die with a chance of LETALITY.
The simulation is started from the command line by invoking
./epidemic <Number of Timesteps> <Size of initial population>
e.g.: ./epidemic 100 1000.
Epidemic depends on lava/matplotlib-cpp for building. See the instructions and installation requirements there.
If you do not whish to plot the output, you can export the data in ParaView-compatible .csv.NUM_STEP files (where NUM_STEP is a running index).
Uncomment the #define CSVOUTPUT and comment out the #define ANIMATION guards in the src/Simulation.cpp file to enable CSV-only output.
Build with CMake inside a build directory.
python
matplotlib
git clone https://github.com/mjjrt/Epidemic
cd Epidemic
mkdir build && cd build
cmake ..
cmake --build .
./epidemic 100 1000
TBD.
- The
matplotlibcpp.hdependency seems to have a bug in theplt::legend()template function. Therefore, it is not possible to fix the legend to one place without restructuring the image. That's why the plot looks so off-center.
- Implement infection susceptibility
- Faster Check Loops
- Better Plotting/Rendering
- Documentation