PCA for every single pair of populations in the 1000 Genomes Populations
30-second explanation on how these were made
- I used the the .vcf file from the affy_6 chip affy_6
- Converted to plink format
- Removed SNPs with maf < 0.01
- Created one set of plink files for all pairs of populations
- LD based pruning using plink with arguments 50 5 0.8
- Ran PCA using plink
- Used homemade python script to generate the pdfs
- The big pdf, with all the populations, was made using pdfjam
- More questions? Simply go look at the code
Examples
Low quality png render of a huge pdf with all pcas on it (I recommend you download it!)

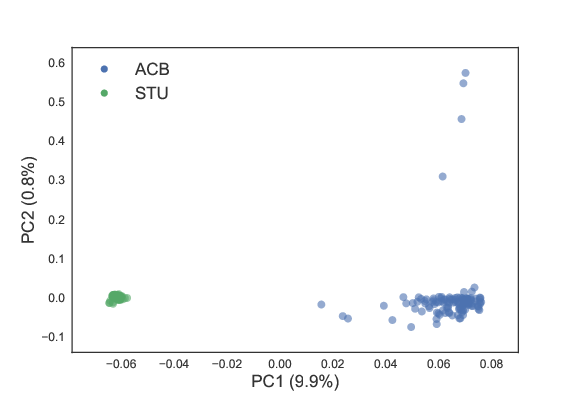
Example of pca for a pair of populations. All of the pairs can be found in the figures folder.