How to use this repo
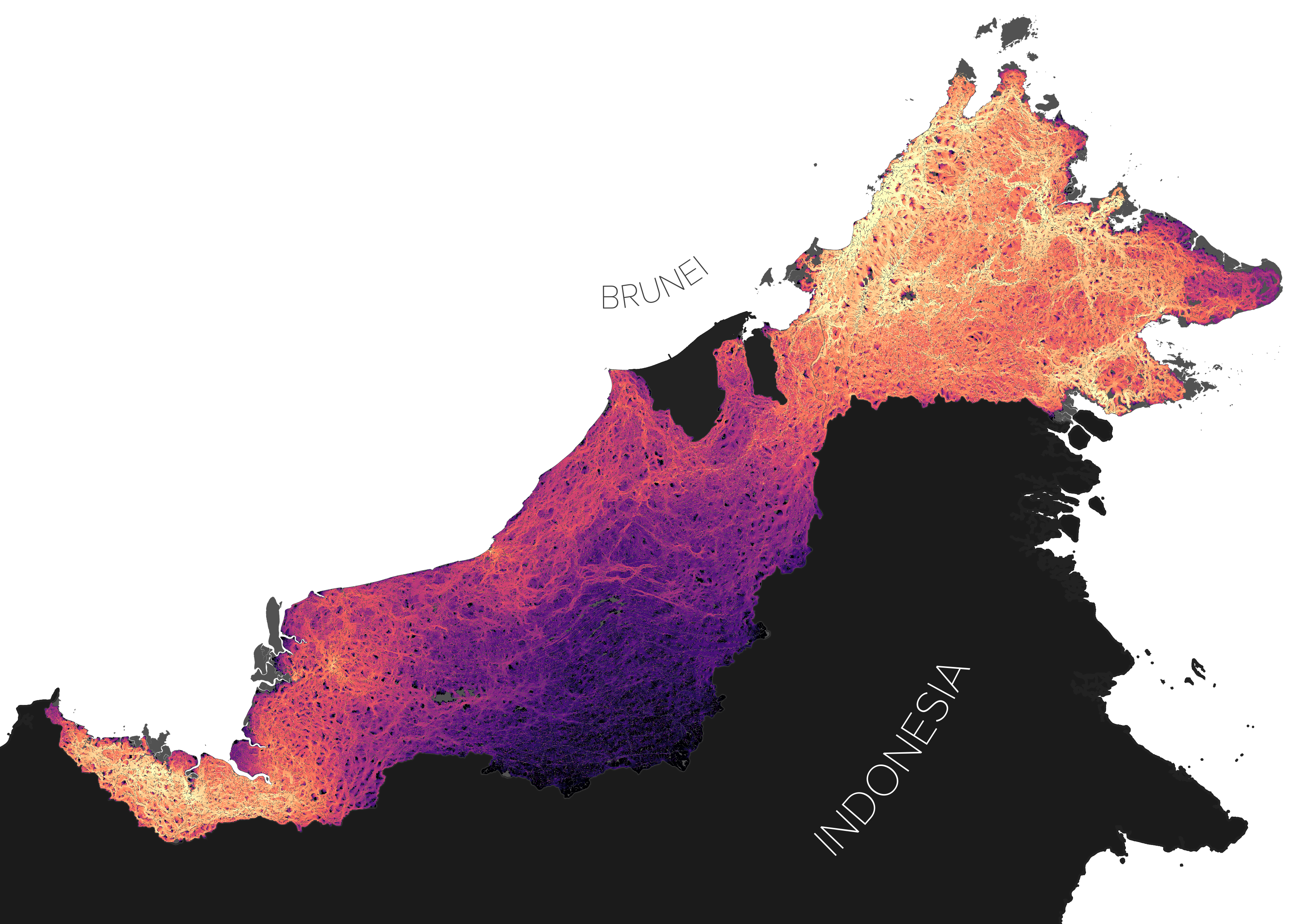
This repo contains all code from Deith and Brodie, 2020 - "Predicting defaunation – accurately mapping bushmeat hunting pressure over large areas". The code was used to create the following map of Malaysian Borneo's accessibility to wild meat hunters:
Script files are organized into folders based on their function.
- Creation of a resistance surface
- ResistanceMapGeneration.py - using geospatial data layers with the same extent and resolution, performs calculations based on a. landcover, b. slope (from a digital elevation model), c. roadways, and d. rivers and stream order (from a digital elevation model).
- Circuit-theory based simulations across the resistance surface (using GFLOW software by Leonard et al.)
- GRASS_IsochroneSampler.py - using settlement location geospatial data layer (as a .shp file), create GFLOW-specific source and sink node files (in .txt and .tsv format) to use as input to GFLOW simulations. This Python script must be run within an interactive GRASS session's terminal.
- CircuitTheoreticMapCreation_GFLOW.sh - modification of script from GFLOW to loop iteratively over the source-sink pairs created with
GRASS_IsochroneSampler.py. Output maps are created and modified according to local population density (inserted into the file names of the.txtand.tsvfiles created in theGRASS_IsochroneSampler.py). - GFLOWOutput_Summation.py - this script iterates through output files created with
CircuitTheoreticMapCreation_GFLOW.sh, modifies the outputs of these files based on population density at each source village, and sums these into a final cumulative .tif map of circuit-theoretic accessibility for all source-sink pairings. This may need to be modified if the ouputs ofCircuitTheoreticMapCreation_GFLOW.share modified within the shell script.
- N-mixture model creation and assessment
- NMixtureModel_Workflow.R - an R file which contains all steps of the N-mixture model creation process (broken down into individual files in the subfolder
NMixModel_Steps):- main.R - identical to the workflow script, this will run all stages of analysis by sourcing the following scripts:
- functions.R - helper functions that are used later in the
- load.R - load the raw data for covariates in the N-mixture models
- clean.R - cleans input data and prepares it in the correct format for N-mixture model fitting
- modelfitting.R - given a list of covariates, automatically creates formulae for N-mixture model and fits these models (designed to run on a cluster).