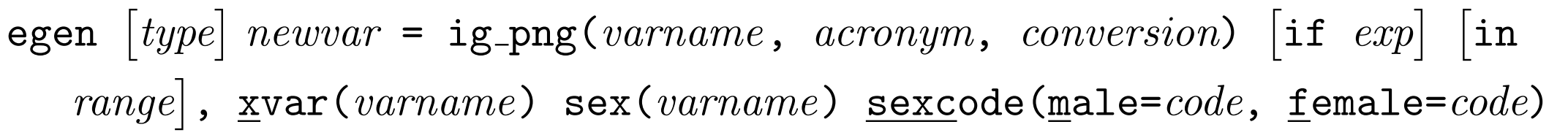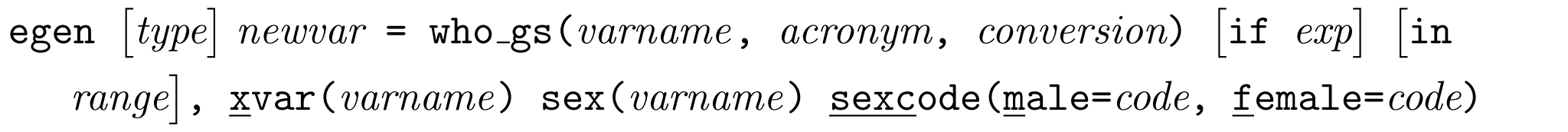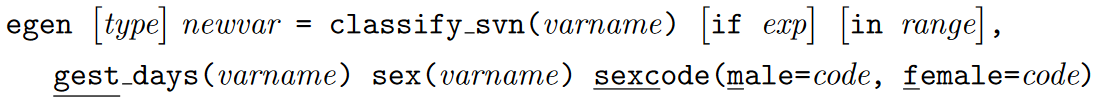gigs: Newborn and infant growth assessment in Stata
About
Produced as part of the Guidance for International Growth Standards (GIGS)
project, gigs provides a single, simple interface for working with the WHO
Child Growth Standards and outputs from the INTERGROWTH-21st project.
You will find functions for converting between anthropometric measures (e.g.
weight or length) to z-scores and centiles, and the inverse. Also included are
functions for classifying newborn and infant growth according to
literature-based cut-offs.
Installation
The gigs package is available for Stata version 16 and over. You can install
the latest stable release of gigs from GitHub using the
github module for Stata:
. github install lshtm-gigs/gigs-stataAlternatively, you can download a stable release of your choice from GitHub using the
net install command from Stata. Simply go to the stable release of gigs that
you want to download from the
releases page on GitHub, and
download the zipped archive. Unzip this downloaded archive. Within this unzipped
folder will be another folder, inside which will be the .ado/.dta files needed
for gigs to work. Put the path to the folder containing the .ado/.dta files
in the from() option of net install, and Stata will install the necessary
files.
. net install gigs, from("directory/of/unzipped/folder/with/ado/files")
Available standards
-
ig_nbs- INTERGROWTH-21st standards for newborn sizeComponent standards
Acronym Description Unit gest_days()rangewfgaWeight-or-gestational age kg 168 to 300 days lfgaLength-for-gestational age cm 168 to 300 days hcfgaHead circumference-for-gestational age cm 168 to 300 days wlrfgaWeight-to-length ratio-for-gestational age kg/cm 168 to 300 days ffmfgaFat-free mass-for-gestational age kg 266 to 294 days bfpfgaBody fat percentage-for-gestational age % 266 to 294 days fmfgaFat mass-for-gestational age kg 266 to 294 days -
ig_png- INTERGROWTH-21st standards for postnatal growth in preterm infantsComponent standards
Acronym Description Unit xvar()rangewfaweight-for-age kg 27 to <64 exact weeks lfalength-for-age cm 27 to <64 exact weeks hcfahead circumference-for-age cm 27 to <64 exact weeks wflweight-for-length kg 35 to 65 cm -
who_gs- WHO Child Growth Standards for term infantsComponent standards
Acronym Description Unit xvar()rangewfaweight-for-age kg 0 to 1856 days bfaBMI-for-age kg/m2 0 to 1856 days lhfalength/height-for-age cm 0 to 1856 days hcfahead circumference-for-age cm 0 to 1856 days wflweight-for-length kg 45 to 110 cm wfhweight-for-height kg 65 to 120 cm acfaarm circumference-for-age cm 91 to 1856 days ssfasubscapular skinfold-for-age mm 91 to 1856 days tsfatriceps skinfold-for-age mm 91 to 1856 days
Conversion functions
Each conversion function has similar syntax. The main function call determines
the set of standards in use, the acronym parameter specifies which component
standard is being used, and the conversion parameter specifies the type of
conversion you wish to perform. This conversion parameter can take one of four
values: "v2z" (value-to-z-score), "v2c" (value-to-centile), "z2v"
(z-score-to-value), "c2v" (centile-to-value). The sex() and sexcode()
options are used to give the function sex data - as the growth standards are
sex-specific, the standards cannot be applied correctly without this information.
INTERGROWTH-21st Newborn Size standards, including very preterm
This function can be used to convert between measurements and z-scores/centiles in each of the INTERGROWTH-21st Newborn Size Standards.
INTERGROWTH-21st Postnatal Growth standards
This function can be used to convert between measurements and z-scores/centiles in each of the INTERGROWTH-21st Postnatal Growth of Preterm Infants Standards.
WHO Child Growth Standards
This function can be used to convert between measurements and z-scores/centiles in each of the WHO Child Growth Standards.
Classification functions
These functions are used to classify infant growth according to published cut-offs. These publications are discussed in the attached paper.
Size for gestational age
This function outputs a variable with the following values and labels. Severely
SGA infants are only labelled if the severe option is specified:
| Value | Meaning | Centile range |
|---|---|---|
| -2 | Severely small for gestational age | <3rd |
| -1 | Small for gestational age (SGA) | <10th |
| 0 | Appropriate for gestational age (AGA) | 10th to 90th |
| 1 | Large for gestational age (LGA) | >90th |
Small vulnerable newborns
This function outputs a variable with the following values and labels:
| Value | Meaning | Term Status | Centile range |
|---|---|---|---|
| -4 | Preterm SGA | Preterm | <10th |
| -3 | Preterm AGA | Preterm | 10th to 90th |
| -2 | Preterm LGA | Preterm | >90th |
| -1 | Term SGA | Term | <10th |
| 0 | Term AGA | Term | 10th to 90th |
| 1 | Term LGA | Term | >90th |
Stunting
The function outputs a variable with the following values and labels. Outlier
observations are only labelled if the outliers option is specified:
| Value | Meaning | Z-score range |
|---|---|---|
| -2 | Severe stunting | -5 to -3 |
| -1 | Stunting | -3 to -2 |
| 0 | Normal | -2 to 5 |
| -10 | Implausible | <-5 or >5 |
Wasting
The function outputs a variable with the following values and labels. Outlier
observations are only labelled if the outliers option is specified:
| Value | Meaning | Z-score range |
|---|---|---|
| -2 | Severe wasting | -5 to -3 |
| -1 | Wasting | -3 to -2 |
| 0 | Normal | -2 to 2 |
| 1 | Overweight | 2 to 5 |
| -10 | Implausible | <-5 or >5 |
Weight-for-age
The function outputs a variable with the following values and labels. Outlier
observations are only labelled if the outliers option is specified:
| Value | Meaning | Z-score range |
|---|---|---|
| -2 | Severely underweight | -6 to -3 |
| -1 | Underweight | -3 to -2 |
| 0 | Normal weight | -2 to 2 |
| 1 | Overweight | 2 to 5 |
| -10 | Implausible | <-6 or >5 |
Examples
This section illustrates a possible use case using life6mo.dta, an extract of
data from the Low birthweight Infant Feeding Exploration (LIFE) Study. It
contains weight measurements for term and preterm infants from birth
(visitweek == 0) to around six months of age (visitweek == 26).
. use life6mo
. local 37weeks 7 * 37
. list in f/10, noobs abbreviate(10) sep(10)
___________________________________________________________
| infantid gestage pma sex visitweek meaninfwgt |
|-----------------------------------------------------------|
| 101-1002-1 132 133 2 0 2300 |
| 101-1002-1 132 139 2 1 2270 |
| 101-1002-1 132 146 2 2 2465 |
| 101-1002-1 132 162 2 4 2700 |
| 101-1002-1 132 174 2 6 3160 |
| 101-1002-1 132 203 2 10 4004 |
| 101-1002-1 132 230 2 14 4893.333 |
| 101-1002-1 132 258 2 18 5690 |
| 101-1002-1 132 317 2 26 6950 |
| 101-1003-1 237 238 1 0 1900 |
|-----------------------------------------------------------|Conversion
We can use the conversion functions listed above to generate weight-for-age z-scores (WAZs) in the different study populations (i.e. term vs preterm) and measurement timings (i.e. z-scores for newborns with INTERGROWTH-21st Newborn Size Standards, WHO/INTERGROWTH Postnatal standards after birth).
. egen double waz_nbs = ig_nbs(meaninfwgt/1000, "wfga", "v2z") ///
> if agedays == 0, ///
> gest_days(gestage) sex(sex) sexcode(m=1, f=2)
(8,228 missing values generated)
. egen double waz_who = who_gs(meaninfwgt/1000, "wfa", "v2z") ///
> if agedays > 0 & gestage >= `37weeks', ///
> xvar(agedays) sex(sex) sexcode(m=1, f=2)
(4,073 missing values generated)
. gen pma_weeks = pma / 7
. gen pma_weeks_floored = floor(pma / 7)
. egen double waz_png = ig_png(meaninfwgt/1000, "wfa", "v2z") ///
> if gestage < `37weeks' & agedays > 0, ///
> xvar(pma_weeks_floored) sex(sex) sexcode(m=1, f=2)
(4,659 missing values generated)
. drop pma_weeks pma_weeks_floored
. gen age_corrected = pma - `40weeks´
. egen waz_whocorr = who_gs(meaninfwgt/1000, "wfa", "v2z") ///
> if gestage < `37weeks´ & visitweek == 26, ///
> xvar(age_corrected) sex(sex) sexcode(m=1, f=2)
(7,996 missing values generated)
. drop age_correctedWe can then combine these WAZs into one overall waz variable:
. gen double waz = waz_who if gestage > `37weeks'
(4,749 missing values generated)
. replace waz = waz_png if gestage < `37weeks'
(3,757 real changes made)
. replace waz = waz_nbs if agedays == 0
(188 real changes made)
. list visitweek gestage pma waz_* waz in f/10, noobs sep(10)
+------------------------------------------------------------------------+
| visitw~k gestage pma waz_nbs waz_who waz_png waz |
|------------------------------------------------------------------------|
| 0 132 133 . . . . |
| 1 132 139 . . . . |
| 2 132 146 . . . . |
| 4 132 162 . . . . |
| 6 132 174 . . . . |
| 10 132 203 . . 7.8594946 7.8594946 |
| 14 132 230 . . 7.2548282 7.2548282 |
| 18 132 258 . . 6.0985216 6.0985216 |
| 26 132 317 . . 3.8344571 3.8344571 |
| 0 237 238 . . -.43606208 -.43606208 |
+------------------------------------------------------------------------+This waz variable can then be used to determine whether infants are
underweight at different age points, or to track the growth trajectory of
individual children.
Classification
This dataset contains information on weight at birth, so could be used to
calculate size-for-gestational age classifications. We can reload the data, then
remove any observations which were not made at birth. We then use the
classify_sga() command to give us our classifications:
. use life6mo, clear
. keep if (gestage - pma) == 0
(7,303 observations deleted)
. egen sga = classify_sga(meaninfwgt/1000), ///
> gest_days(gestage) sex(sex) sexcode(m=1, f=2)
(2 missing values generated)Known issues and bug reporting
We kindly request that users note any bugs, issues, or feature requests on the GitHub issues page.
Authors
S. R. Parker
Maternal, Adolescent, Reproductive, and Child Health Centre
London School of Hygiene and Tropical Medicine
Dr E. O. Ohuma
Maternal, Adolescent, Reproductive, and Child Health Centre
London School of Hygiene and Tropical Medicine