GM-AE
The repository of GM-AE is an implementation of the paper 'Parameterized Gompertz-guided Morphological AutoEncoder for Predicting Pulmonary Nodule Growth', accepted by TMI. Nodules used for GM-AE is listed in 'data/original coordinates/dataset_part_2301.csv'
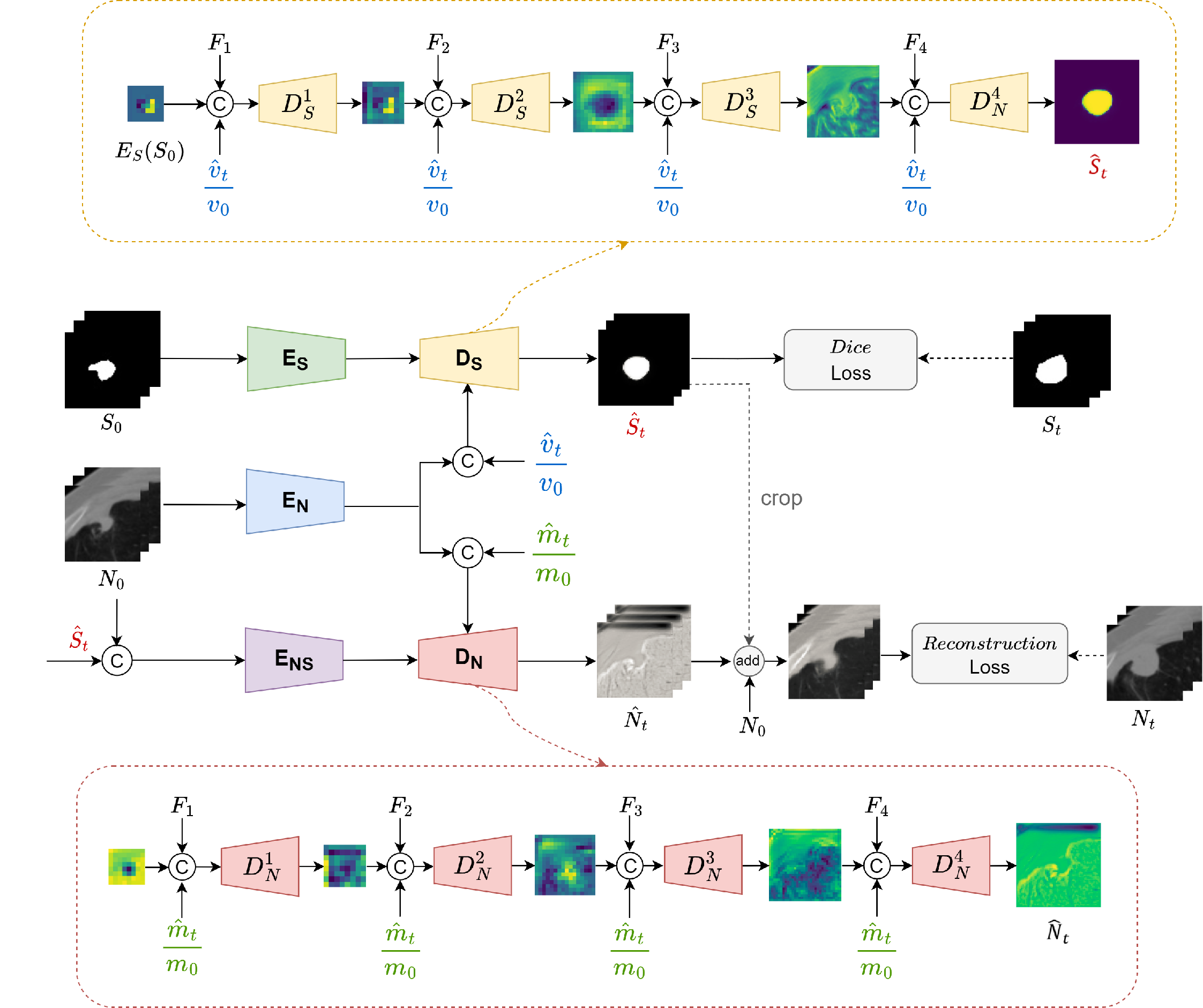
Architecture of morphological autoencoder equipped with shape-aware (DS) and texture-aware (DN ) decoders.
ST-Mixer
Introduction
This repository is an implementation of the paper 'Siamese Encoder-based Spatial-Temporal Mixer for Growth Trend Prediction of Lung Nodules on CT Scans', accepted by MICCAI2022, and is built upon DeiT, thank them very much!
Schematic of our proposed method, including a siamese encoder, a spatial-temporal mixer (STM) and a two-layer H-loss.
Usage
Training:
Run
python main_ddp.py --resume --gpu 1,2 --batch-size 16 --epochs 60 --distributed
Inference:
Run
python inference.py
Datasets
The pipeline of organizing the temporal CT dataset.
The pipeline of organizing the temporal CT dataset (NLSTt), including CT scan registration, ROI pairing, and class annotation. The letter d denotes the diameter of lung nodules, and c indicates their corresponding texture types, i.e., solid, part-solid (PS), ground-glass nodule (GGN).
The paired nodules from NLST is listed in './data/registration coordinates/data.csv' (NLSTt), and please download raw CT scans from NLST
Statistics of benchmark splits of the NLSTt dataset and in-house dataset.
| Types | Train set | Validation set | Test set | In-house set |
|---|---|---|---|---|
| GGN | 2,683 | 564 | 658 | 129 |
| Solid | 4,121 | 900 | 892 | 352 |
| PS | 148 | 26 | 37 | 6 |
| Total | 6,952 | 1,490 | 1,587 | 487 |
Note
Unfortunately, we saved and used coordinates of nouldes after the regisration in the Paper, which is inconvenient to sharing coordinates. So we will release the original coordinates step by step in 'data/original coordinates'.
Results
AUC (in %) of different mixers and encoders on the test and in-house sets.
| Encoder | Mixer | Test Set AUC@H1 | Test Set AUC@H2 | In-house set AUC@H1 | In-house set AUC@H2 |
|---|---|---|---|---|---|
| CNN | Concat | 80.8 | 75.3 | 67.2 | 67.2 |
| CNN | LSTM | 81.8 | 75.0 | 64.0 | 71.0 |
| CNN | STM (Ours) | 83.0 | 76.3 | 73.5 | 71.6 |
| ViT | Concat | 82.6 | 75.2 | 64.2 | 64.1 |
| ViT | LSTM | 82.6 | 76.3 | 67.1 | 74.7 |
| ViT | STM (Ours) | 83.6 | 77.5 | 72.8 | 78.5 |
ACC of nodule types of the test set on different methods.
| Method | ACC-GGN | ACC-Solid | ACC-PS |
|---|---|---|---|
| CNN+STM | 90.9 | 88.2 | 56.8 |
| ViT+STM | 92.4 | 91.6 | 59.5 |
ACC of nodule types of in-house set on different methods.
| Method | ACC-GGN | ACC-Solid | ACC-PS |
|---|---|---|---|
| CNN+STM | 87.6 | 91.2 | 58.1 |
| ViT+STM | 93.8 | 90.6 | 60.5 |
| Clinician A | 85.3 | 93.2 | 60.5 |
| Clinician B | 86.0 | 94.0 | 62.8 |
Examples of predicting the growth trend by our model.
Examples of predicting the growth trend by our model and clinicians A and B. The first row is the ground-truth of the evolution classes, and the predicted results in red color are the incorrect predictions. The symbols →, ↑, and ↓ denote the classesstability, dilatation and shrinkage, respectively.
License
This repository is released under the Apache 2.0 license as found in the LICENSE file.
Citation
Please cite ST-Mixer in your publications if it helps your research.
@article{fang2022siamese,
title={Siamese Encoder-based Spatial-Temporal Mixer for Growth Trend Prediction of Lung Nodules on CT Scans},
author={Fang, Jiansheng and Wang, Jingwen and Li, Anwei and Yan, Yuguang and Hou, Yonghe and Song, Chao and Liu, Hongbo and Liu, Jiang},
journal={arXiv preprint arXiv:2206.03049},
year={2022}
}