R package for subsampling genomic data based on epidemiological time series data.
You can install the development version of subsamplerr from GitHub with:
devtools::install_github("leke-lyu/subsamplerr")Count the number of genome samples by Epi-Week and location, and Integrate daily count of case data into weekly count:
library(subsamplerr)
texasSeq <- texasSeqMeta %>% metaTableToMatrix(., "location", "date") %>% exactDateToEpiweek(.)
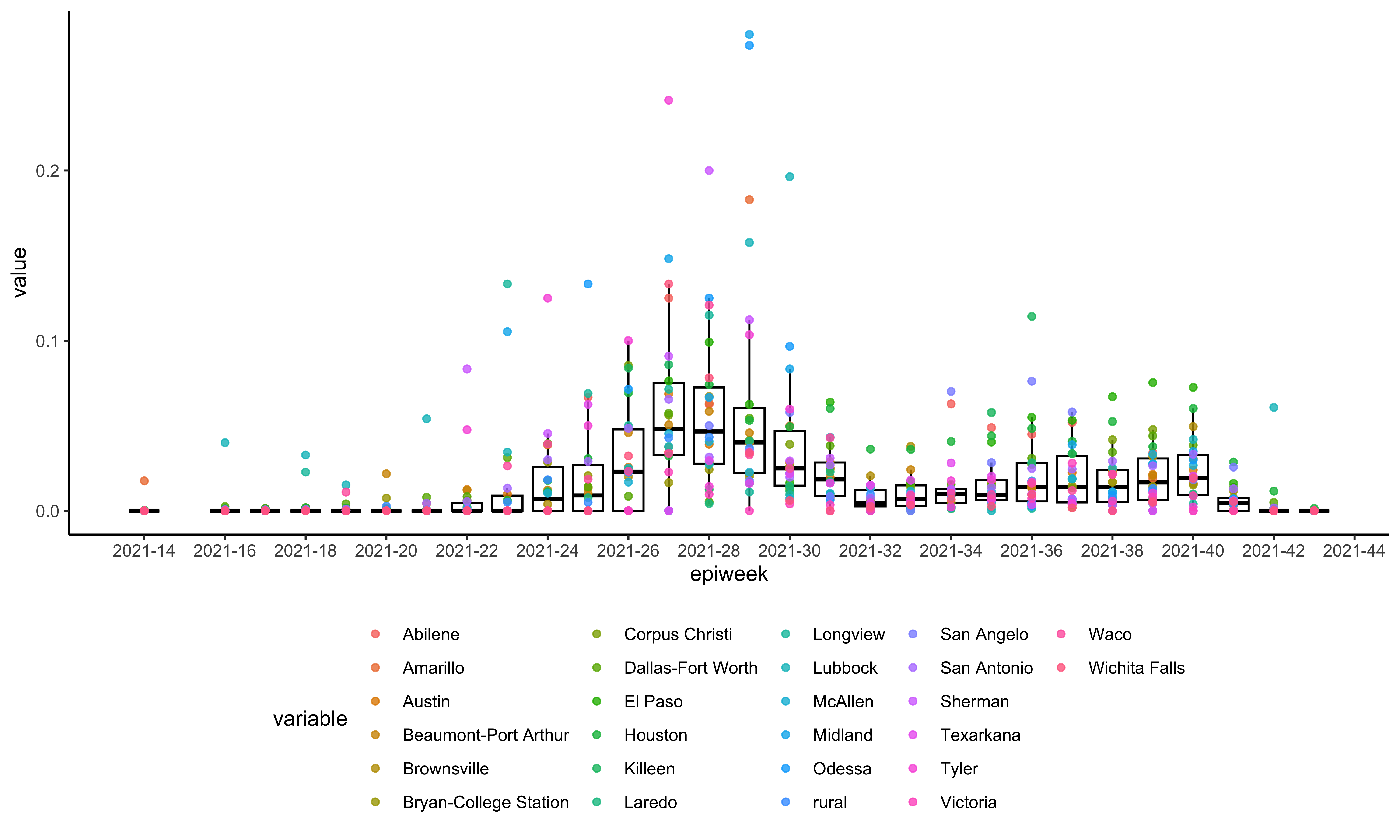
texasCase %<>% exactDateToEpiweek(.)Inspect the sampling heterogeneity of the Texas dataset:
plotSequencingPercentage(texasSeq, texasCase)Generate sampled dataset with baseline equals 0.006
texasSample <- expectedSampleMatrix(0.006, texasSeq, texasCase)
id <- proportionalSampling(texasSample, texasSeqMeta)
#> [1] "Given the basline equals 0.006, 5899 genomes are sampled."
#> .
#> Dallas-Fort Worth Houston San Antonio
#> 1835 1523 593
#> rural Austin McAllen
#> 516 465 113
#> Corpus Christi Beaumont-Port Arthur Killeen
#> 102 84 78
#> Brownsville Bryan-College Station El Paso
#> 66 60 57
#> Lubbock Waco Tyler
#> 48 48 43
#> Amarillo Laredo Midland
#> 38 32 29
#> Wichita Falls Sherman Odessa
#> 26 25 24
#> Longview Abilene Victoria
#> 23 22 21
#> Texarkana San Angelo
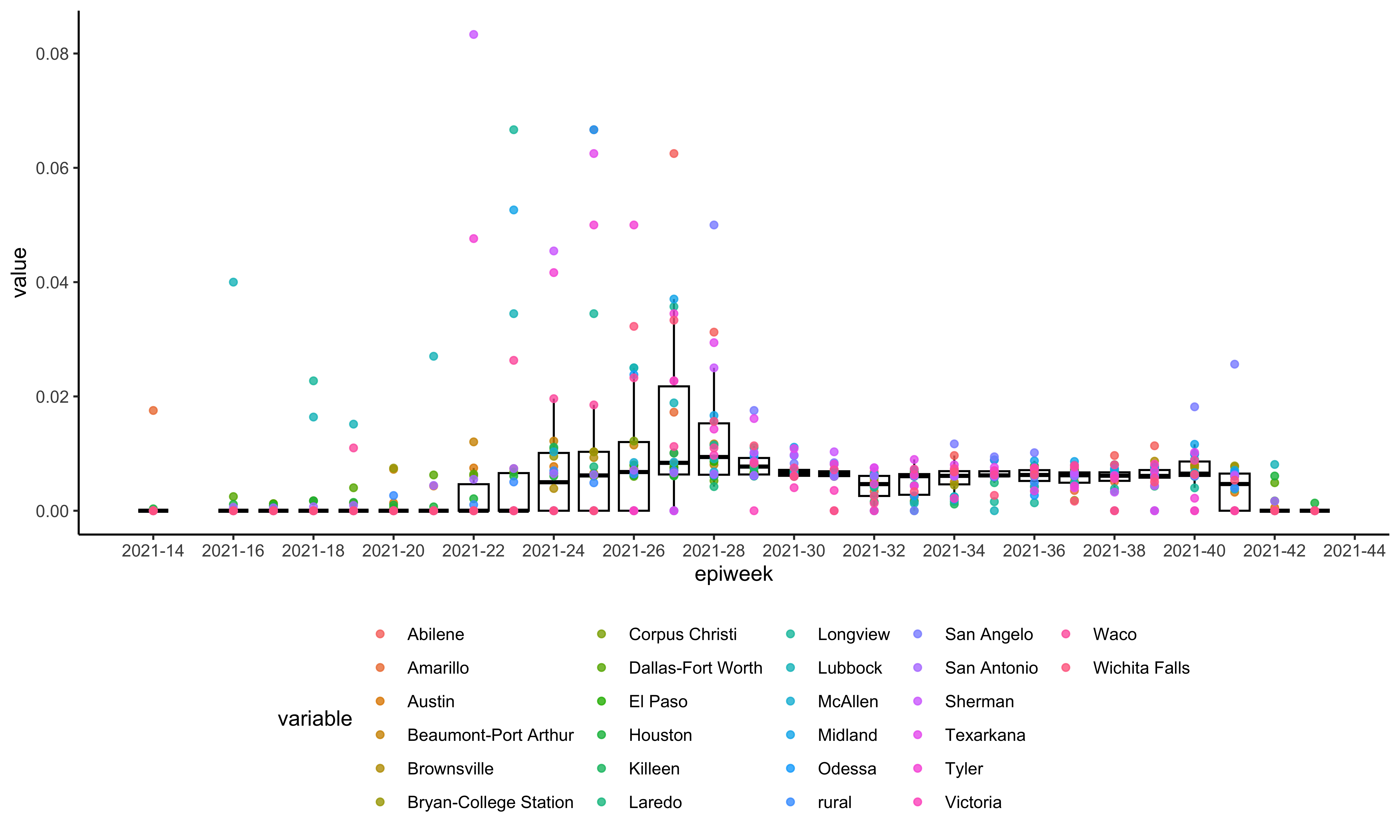
#> 15 13Inspect the sampling heterogeneity of the sampled dataset:
plotSequencingPercentage(texasSample, texasCase)