PySDM is a package for simulating the dynamics of population of particles. It is intended to serve as a building block for simulation systems modelling fluid flows involving a dispersed phase, with PySDM being responsible for representation of the dispersed phase. Currently, the development is focused on atmospheric cloud physics applications, in particular on modelling the dynamics of particles immersed in moist air using the particle-based (a.k.a. super-droplet) approach to represent aerosol/cloud/rain microphysics. The package features a Pythonic high-performance implementation of the Super-Droplet Method (SDM) Monte-Carlo algorithm for representing collisional growth (Shima et al. 2009), hence the name.
There is a growing set of example Jupyter notebooks exemplifying how to perform various types of calculations and simulations using PySDM. Most of the example notebooks reproduce results and plot from literature, see below for a list of examples and links to the notebooks (which can be either executed or viewed "in the cloud").
There are also a growing set of tutorials, also in the form of Jupyter notebooks. These tutorials are intended for teaching purposes and include short explanations of cloud microphysical concepts paired with widgets for running interactive simulations using PySDM. Each tutorial also comes with a set of questions at the end that can be used as homework problems. Like the examples, these tutorials can be executed or viewed "in the cloud" making it an especially easy way for students to get started.
PySDM has two alternative parallel number-crunching backends
available: multi-threaded CPU backend based on Numba
and GPU-resident backend built on top of ThrustRTC.
The Numba backend (aliased CPU) features multi-threaded parallelism for
multi-core CPUs, it uses the just-in-time compilation technique based on the LLVM infrastructure.
The ThrustRTC backend (aliased GPU) offers GPU-resident operation of PySDM
leveraging the SIMT
parallelisation model.
Using the GPU backend requires nVidia hardware and CUDA driver.
For an overview of PySDM features (and the preferred way to cite PySDM in papers), please refer to our JOSS papers:
- Bartman et al. 2022 (PySDM v1).
- de Jong, Singer et al. 2023 (PySDM v2).
PySDM includes an extension of the SDM scheme to represent collisional breakup described in de Jong, Mackay et al. 2023.
For a list of talks and other materials on PySDM, see the project wiki.
A pdoc-generated documentation of PySDM public API is maintained at: https://open-atmos.github.io/PySDM
PySDM dependencies are: Numpy, Numba, SciPy, Pint, chempy, pyevtk, ThrustRTC and CURandRTC.
To install PySDM using pip, use: pip install PySDM
(or pip install git+https://github.com/open-atmos/PySDM.git to get updates
beyond the latest release).
Conda users may use pip as well, see the Installing non-conda packages section in the conda docs. Dependencies of PySDM are available at the following conda channels:
- numba: numba
- conda-forge: pyevtk, pint and
- fyplus: ThrustRTC, CURandRTC
- bjodah: chempy
- nvidia: cudatoolkit
For development purposes, we suggest cloning the repository and installing it using pip -e.
Test-time dependencies can be installed with pip -e .[tests].
PySDM examples constitute the PySDM-examples package.
The examples have additional dependencies listed in PySDM_examples package setup.py file.
Running the example Jupyter notebooks requires the PySDM_examples package to be installed.
The suggested install and launch steps are:
git clone https://github.com/open-atmos/PySDM.git
pip install -e PySDM
pip install -e PySDM/examples
jupyter-notebook PySDM/examples/PySDM_examples
Alternatively, one can also install the examples package from pypi.org by
using pip install PySDM-examples (note that this does not apply to notebooks itself,
only the supporting .py files).
mindmap
root((PySDM))
Builder
Formulae
Particulator
((attributes))
(physics)
DryVolume: ExtensiveAttribute
Kappa: DerivedAttribute
...
(chemistry)
Acidity
...
(...)
((backends))
CPU
GPU
((dynamics))
AqueousChemistry
Collision
Condensation
...
((environments))
Box
Parcel
Kinematic2D
...
((initialisation))
(spectra)
Lognormal
Exponential
...
(sampling)
(spectral_sampling)
ConstantMultiplicity
UniformRandom
Logarithmic
...
(...)
(...)
((physics))
(hygroscopicity)
KappaKoehler
...
(condensation_coordinate)
Volume
VolumeLogarithm
(...)
((products))
(size_spectral)
EffectiveRadius
WaterMixingRatio
...
(ambient_thermodynamics)
AmbientRelativeHumidity
...
(...)
In order to depict the PySDM API with a practical example, the following
listings provide sample code roughly reproducing the
Figure 2 from Shima et al. 2009 paper
using PySDM from Python, Julia and Matlab.
It is a Coalescence-only set-up in which the initial particle size
spectrum is Exponential and is deterministically sampled to match
the condition of each super-droplet having equal initial multiplicity:
Julia (click to expand)
using Pkg
Pkg.add("PyCall")
Pkg.add("Plots")
Pkg.add("PlotlyJS")
using PyCall
si = pyimport("PySDM.physics").si
ConstantMultiplicity = pyimport("PySDM.initialisation.sampling.spectral_sampling").ConstantMultiplicity
Exponential = pyimport("PySDM.initialisation.spectra").Exponential
n_sd = 2^15
initial_spectrum = Exponential(norm_factor=8.39e12, scale=1.19e5 * si.um^3)
attributes = Dict()
attributes["volume"], attributes["multiplicity"] = ConstantMultiplicity(spectrum=initial_spectrum).sample(n_sd)Matlab (click to expand)
si = py.importlib.import_module('PySDM.physics').si;
ConstantMultiplicity = py.importlib.import_module('PySDM.initialisation.sampling.spectral_sampling').ConstantMultiplicity;
Exponential = py.importlib.import_module('PySDM.initialisation.spectra').Exponential;
n_sd = 2^15;
initial_spectrum = Exponential(pyargs(...
'norm_factor', 8.39e12, ...
'scale', 1.19e5 * si.um ^ 3 ...
));
tmp = ConstantMultiplicity(initial_spectrum).sample(int32(n_sd));
attributes = py.dict(pyargs('volume', tmp{1}, 'multiplicity', tmp{2}));Python (click to expand)
from PySDM.physics import si
from PySDM.initialisation.sampling.spectral_sampling import ConstantMultiplicity
from PySDM.initialisation.spectra.exponential import Exponential
n_sd = 2 ** 15
initial_spectrum = Exponential(norm_factor=8.39e12, scale=1.19e5 * si.um ** 3)
attributes = {}
attributes['volume'], attributes['multiplicity'] = ConstantMultiplicity(initial_spectrum).sample(n_sd)The key element of the PySDM interface is the Particulator
class instances of which are used to manage the system state and control the simulation.
Instantiation of the Particulator class is handled by the Builder
as exemplified below:
Julia (click to expand)
Builder = pyimport("PySDM").Builder
Box = pyimport("PySDM.environments").Box
Coalescence = pyimport("PySDM.dynamics").Coalescence
Golovin = pyimport("PySDM.dynamics.collisions.collision_kernels").Golovin
CPU = pyimport("PySDM.backends").CPU
ParticleVolumeVersusRadiusLogarithmSpectrum = pyimport("PySDM.products").ParticleVolumeVersusRadiusLogarithmSpectrum
radius_bins_edges = 10 .^ range(log10(10*si.um), log10(5e3*si.um), length=32)
env = Box(dt=1 * si.s, dv=1e6 * si.m^3)
builder = Builder(n_sd=n_sd, backend=CPU(), environment=env)
builder.add_dynamic(Coalescence(collision_kernel=Golovin(b=1.5e3 / si.s)))
products = [ParticleVolumeVersusRadiusLogarithmSpectrum(radius_bins_edges=radius_bins_edges, name="dv/dlnr")]
particulator = builder.build(attributes, products)Matlab (click to expand)
Builder = py.importlib.import_module('PySDM').Builder;
Box = py.importlib.import_module('PySDM.environments').Box;
Coalescence = py.importlib.import_module('PySDM.dynamics').Coalescence;
Golovin = py.importlib.import_module('PySDM.dynamics.collisions.collision_kernels').Golovin;
CPU = py.importlib.import_module('PySDM.backends').CPU;
ParticleVolumeVersusRadiusLogarithmSpectrum = py.importlib.import_module('PySDM.products').ParticleVolumeVersusRadiusLogarithmSpectrum;
radius_bins_edges = logspace(log10(10 * si.um), log10(5e3 * si.um), 32);
env = Box(pyargs('dt', 1 * si.s, 'dv', 1e6 * si.m ^ 3));
builder = Builder(pyargs('n_sd', int32(n_sd), 'backend', CPU(), 'environment', env));
builder.add_dynamic(Coalescence(pyargs('collision_kernel', Golovin(1.5e3 / si.s))));
products = py.list({ ParticleVolumeVersusRadiusLogarithmSpectrum(pyargs( ...
'radius_bins_edges', py.numpy.array(radius_bins_edges), ...
'name', 'dv/dlnr' ...
)) });
particulator = builder.build(attributes, products);Python (click to expand)
import numpy as np
from PySDM import Builder
from PySDM.environments import Box
from PySDM.dynamics import Coalescence
from PySDM.dynamics.collisions.collision_kernels import Golovin
from PySDM.backends import CPU
from PySDM.products import ParticleVolumeVersusRadiusLogarithmSpectrum
radius_bins_edges = np.logspace(np.log10(10 * si.um), np.log10(5e3 * si.um), num=32)
env = Box(dt=1 * si.s, dv=1e6 * si.m ** 3)
builder = Builder(n_sd=n_sd, backend=CPU(), environment=env)
builder.add_dynamic(Coalescence(collision_kernel=Golovin(b=1.5e3 / si.s)))
products = [ParticleVolumeVersusRadiusLogarithmSpectrum(radius_bins_edges=radius_bins_edges, name='dv/dlnr')]
particulator = builder.build(attributes, products)The backend argument may be set to CPU or GPU
what translates to choosing the multi-threaded backend or the
GPU-resident computation mode, respectively.
The employed Box environment corresponds to a zero-dimensional framework
(particle positions are not considered).
The vectors of particle multiplicities n and particle volumes v are
used to initialise super-droplet attributes.
The Coalescence
Monte-Carlo algorithm (Super Droplet Method) is registered as the only
dynamic in the system.
Finally, the build() method is used to obtain an instance
of Particulator which can then be used to control time-stepping and
access simulation state.
The run(nt) method advances the simulation by nt timesteps.
In the listing below, its usage is interleaved with plotting logic
which displays a histogram of particle mass distribution
at selected timesteps:
Julia (click to expand)
using Plots; plotlyjs()
for step = 0:1200:3600
particulator.run(step - particulator.n_steps)
plot!(
radius_bins_edges[1:end-1] / si.um,
particulator.formulae.particle_shape_and_density.volume_to_mass(
particulator.products["dv/dlnr"].get()[:]
)/ si.g,
linetype=:steppost,
xaxis=:log,
xlabel="particle radius [µm]",
ylabel="dm/dlnr [g/m^3/(unit dr/r)]",
label="t = $step s"
)
end
savefig("plot.svg")Matlab (click to expand)
for step = 0:1200:3600
particulator.run(int32(step - particulator.n_steps));
x = radius_bins_edges / si.um;
y = particulator.formulae.particle_shape_and_density.volume_to_mass( ...
particulator.products{"dv/dlnr"}.get() ...
) / si.g;
stairs(...
x(1:end-1), ...
double(py.array.array('d',py.numpy.nditer(y))), ...
'DisplayName', sprintf("t = %d s", step) ...
);
hold on
end
hold off
set(gca,'XScale','log');
xlabel('particle radius [µm]')
ylabel("dm/dlnr [g/m^3/(unit dr/r)]")
legend()Python (click to expand)
from matplotlib import pyplot
for step in [0, 1200, 2400, 3600]:
particulator.run(step - particulator.n_steps)
pyplot.step(
x=radius_bins_edges[:-1] / si.um,
y=particulator.formulae.particle_shape_and_density.volume_to_mass(
particulator.products['dv/dlnr'].get()[0]
) / si.g,
where='post', label=f"t = {step}s"
)
pyplot.xscale('log')
pyplot.xlabel('particle radius [µm]')
pyplot.ylabel("dm/dlnr [g/m$^3$/(unit dr/r)]")
pyplot.legend()
pyplot.savefig('readme.png')The resultant plot (generated with the Python code) looks as follows:
The component submodules used to create this simulation are visualized below:
graph
COAL[":Coalescence"] --->|passed as arg to| BUILDER_ADD_DYN(["Builder.add_dynamic()"])
BUILDER_INSTANCE["builder :Builder"] -...-|has a method| BUILDER_BUILD(["Builder.build()"])
ATTRIBUTES[attributes: dict] -->|passed as arg to| BUILDER_BUILD
N_SD["n_sd :int"] ---->|passed as arg to| BUILDER_INIT
BUILDER_INIT(["Builder.__init__()"]) --->|instantiates| BUILDER_INSTANCE
BUILDER_INSTANCE -..-|has a method| BUILDER_ADD_DYN(["Builder.add_dynamic()"])
ENV_INIT(["Box.__init__()"]) -->|instantiates| ENV
DT[dt :float] -->|passed as arg to| ENV_INIT
DV[dv :float] -->|passed as arg to| ENV_INIT
ENV[":Box"] -->|passed as arg to| BUILDER_INIT
B["b: float"] --->|passed as arg to| KERNEL_INIT(["Golovin.__init__()"])
KERNEL_INIT -->|instantiates| KERNEL
KERNEL[collision_kernel: Golovin] -->|passed as arg to| COAL_INIT(["Coalesncence.__init__()"])
COAL_INIT -->|instantiates| COAL
PRODUCTS[products: list] ----->|passed as arg to| BUILDER_BUILD
NORM_FACTOR[norm_factor: float]-->|passed as arg to| EXP_INIT
SCALE[scale: float]-->|passed as arg to| EXP_INIT
EXP_INIT(["Exponential.__init__()"]) -->|instantiates| IS
IS["initial_spectrum :Exponential"] -->|passed as arg to| CM_INIT
CM_INIT(["ConstantMultiplicity.__init__()"]) -->|instantiates| CM_INSTANCE
CM_INSTANCE[":ConstantMultiplicity"] -.-|has a method| SAMPLE
SAMPLE(["ConstantMultiplicity.sample()"]) -->|returns| n
SAMPLE -->|returns| volume
n -->|added as element of| ATTRIBUTES
PARTICULATOR_INSTANCE -.-|has a method| PARTICULATOR_RUN(["Particulator.run()"])
volume -->|added as element of| ATTRIBUTES
BUILDER_BUILD -->|returns| PARTICULATOR_INSTANCE["particulator :Particulator"]
PARTICULATOR_INSTANCE -.-|has a field| PARTICULATOR_PROD(["Particulator.products:dict"])
BACKEND_INSTANCE["backend :CPU"] ---->|passed as arg to| BUILDER_INIT
PRODUCTS -.-|accessible via| PARTICULATOR_PROD
NP_LOGSPACE(["np.logspace()"]) -->|returns| EDGES
EDGES[radius_bins_edges: np.ndarray] -->|passed as arg to| SPECTRUM_INIT
SPECTRUM_INIT["ParticleVolumeVersusRadiusLogarithmSpectrum.__init__()"] -->|instantiates| SPECTRUM
SPECTRUM[":ParticleVolumeVersusRadiusLogarithmSpectrum"] -->|added as element of| PRODUCTS
click COAL "https://open-atmos.github.io/PySDM/PySDM/dynamics/collisions/collision.html"
click BUILDER_INSTANCE "https://open-atmos.github.io/PySDM/PySDM/builder.html"
click BUILDER_INIT "https://open-atmos.github.io/PySDM/PySDM/builder.html"
click BUILDER_ADD_DYN "https://open-atmos.github.io/PySDM/PySDM/builder.html"
click ENV_INIT "https://open-atmos.github.io/PySDM/PySDM/environments/index.html"
click ENV "https://open-atmos.github.io/PySDM/PySDM/environments/index.html"
click KERNEL_INIT "https://open-atmos.github.io/PySDM/PySDM/dynamics/collisions/collision_kernels/index.html"
click KERNEL "https://open-atmos.github.io/PySDM/PySDM/dynamics/collisions/collision_kernels/index.html"
click EXP_INIT "https://open-atmos.github.io/PySDM/PySDM/initialisation/spectra/index.html"
click IS "https://open-atmos.github.io/PySDM/PySDM/initialisation/spectra/index.html"
click CM_INIT "https://open-atmos.github.io/PySDM/PySDM/initialisation/sampling/spectral_sampling.html"
click CM_INSTANCE "https://open-atmos.github.io/PySDM/PySDM/initialisation/sampling/spectral_sampling.html"
click SAMPLE "https://open-atmos.github.io/PySDM/PySDM/initialisation/sampling/spectral_sampling.html"
click PARTICULATOR_INSTANCE "https://open-atmos.github.io/PySDM/PySDM/particulator.html"
click BACKEND_INSTANCE "https://open-atmos.github.io/PySDM/PySDM/backends/numba.html"
click BUILDER_BUILD "https://open-atmos.github.io/PySDM/PySDM/builder.html"
click NP_LOGSPACE "https://numpy.org/doc/stable/reference/generated/numpy.logspace.html"
click SPECTRUM_INIT "https://open-atmos.github.io/PySDM/PySDM/products/size_spectral/particle_volume_versus_radius_logarithm_spectrum.html"
click SPECTRUM "https://open-atmos.github.io/PySDM/PySDM/products/size_spectral/particle_volume_versus_radius_logarithm_spectrum.html"
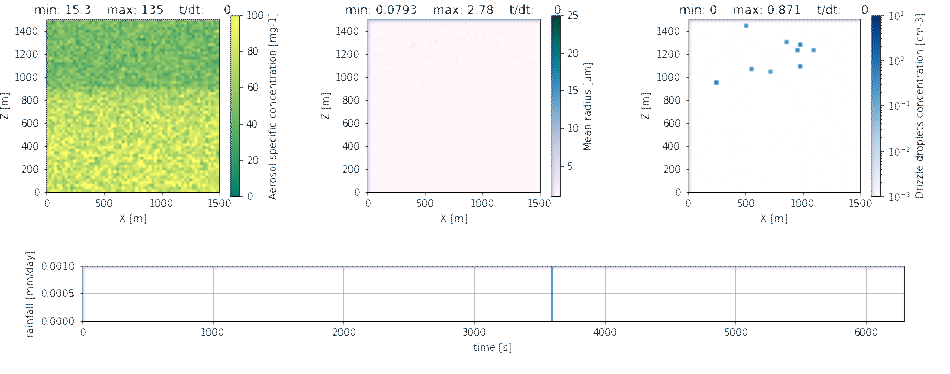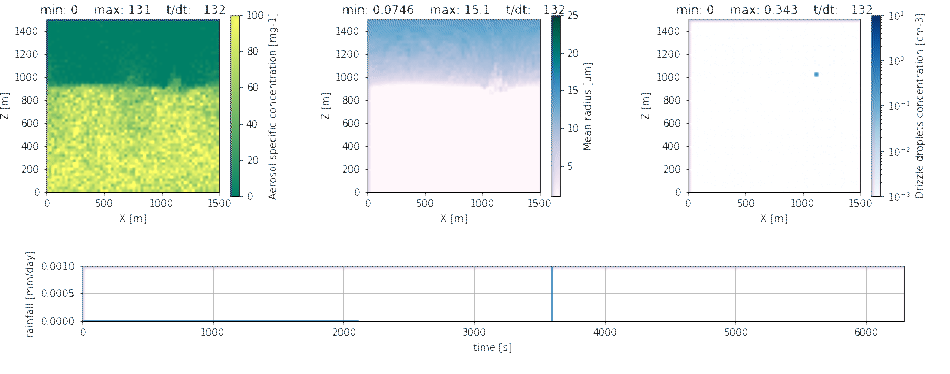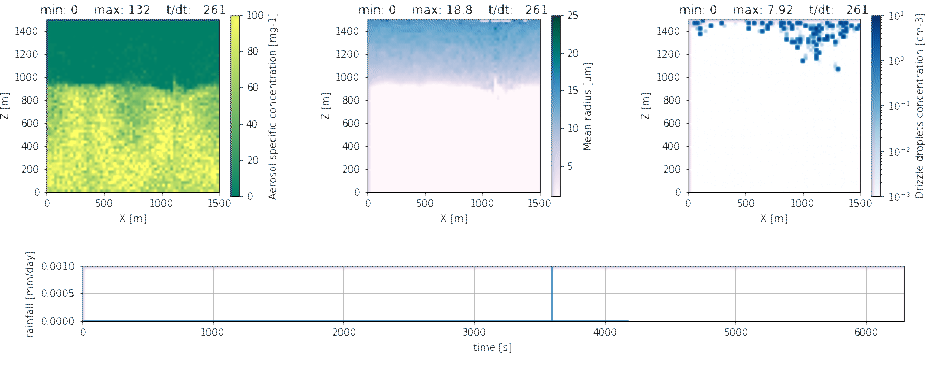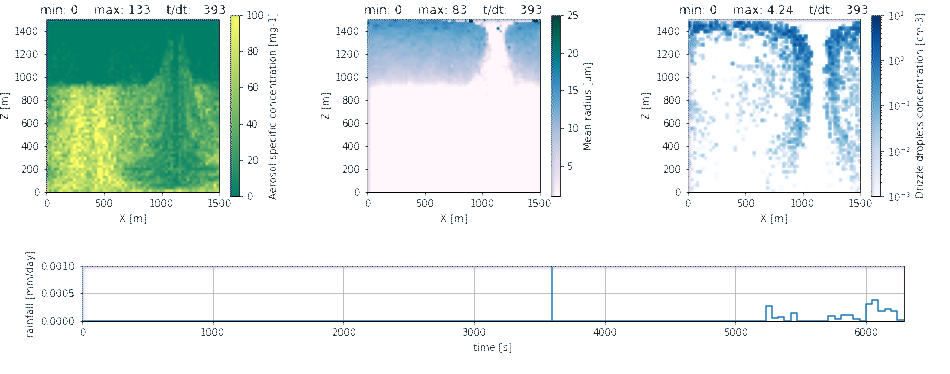
In the following example, a condensation-only setup is used with the adiabatic
Parcel environment.
An initial Lognormal
spectrum of dry aerosol particles is first initialised to equilibrium wet size for the given
initial humidity.
Subsequent particle growth due to Condensation of water vapour (coupled with the release of latent heat)
causes a subset of particles to activate into cloud droplets.
Results of the simulation are plotted against vertical
ParcelDisplacement
and depict the evolution of
PeakSupersaturation,
EffectiveRadius,
ParticleConcentration
and the
WaterMixingRatio .
Julia (click to expand)
using PyCall
using Plots; plotlyjs()
si = pyimport("PySDM.physics").si
spectral_sampling = pyimport("PySDM.initialisation.sampling").spectral_sampling
discretise_multiplicities = pyimport("PySDM.initialisation").discretise_multiplicities
Lognormal = pyimport("PySDM.initialisation.spectra").Lognormal
equilibrate_wet_radii = pyimport("PySDM.initialisation").equilibrate_wet_radii
CPU = pyimport("PySDM.backends").CPU
AmbientThermodynamics = pyimport("PySDM.dynamics").AmbientThermodynamics
Condensation = pyimport("PySDM.dynamics").Condensation
Parcel = pyimport("PySDM.environments").Parcel
Builder = pyimport("PySDM").Builder
Formulae = pyimport("PySDM").Formulae
products = pyimport("PySDM.products")
env = Parcel(
dt=.25 * si.s,
mass_of_dry_air=1e3 * si.kg,
p0=1122 * si.hPa,
initial_water_vapour_mixing_ratio=20 * si.g / si.kg,
T0=300 * si.K,
w= 2.5 * si.m / si.s
)
spectrum = Lognormal(norm_factor=1e4/si.mg, m_mode=50*si.nm, s_geom=1.4)
kappa = .5 * si.dimensionless
cloud_range = (.5 * si.um, 25 * si.um)
output_interval = 4
output_points = 40
n_sd = 256
formulae = Formulae()
builder = Builder(backend=CPU(formulae), n_sd=n_sd, environment=env)
builder.add_dynamic(AmbientThermodynamics())
builder.add_dynamic(Condensation())
r_dry, specific_concentration = spectral_sampling.Logarithmic(spectrum).sample(n_sd)
v_dry = formulae.trivia.volume(radius=r_dry)
r_wet = equilibrate_wet_radii(r_dry=r_dry, environment=env, kappa_times_dry_volume=kappa * v_dry)
attributes = Dict()
attributes["multiplicity"] = discretise_multiplicities(specific_concentration * env.mass_of_dry_air)
attributes["dry volume"] = v_dry
attributes["kappa times dry volume"] = kappa * v_dry
attributes["volume"] = formulae.trivia.volume(radius=r_wet)
particulator = builder.build(attributes, products=[
products.PeakSupersaturation(name="S_max", unit="%"),
products.EffectiveRadius(name="r_eff", unit="um", radius_range=cloud_range),
products.ParticleConcentration(name="n_c_cm3", unit="cm^-3", radius_range=cloud_range),
products.WaterMixingRatio(name="liquid water mixing ratio", unit="g/kg", radius_range=cloud_range),
products.ParcelDisplacement(name="z")
])
cell_id=1
output = Dict()
for (_, product) in particulator.products
output[product.name] = Array{Float32}(undef, output_points+1)
output[product.name][1] = product.get()[cell_id]
end
for step = 2:output_points+1
particulator.run(steps=output_interval)
for (_, product) in particulator.products
output[product.name][step] = product.get()[cell_id]
end
end
plots = []
ylbl = particulator.products["z"].unit
for (_, product) in particulator.products
if product.name != "z"
append!(plots, [plot(output[product.name], output["z"], ylabel=ylbl, xlabel=product.unit, title=product.name)])
end
global ylbl = ""
end
plot(plots..., layout=(1, length(output)-1))
savefig("parcel.svg")Matlab (click to expand)
si = py.importlib.import_module('PySDM.physics').si;
spectral_sampling = py.importlib.import_module('PySDM.initialisation.sampling').spectral_sampling;
discretise_multiplicities = py.importlib.import_module('PySDM.initialisation').discretise_multiplicities;
Lognormal = py.importlib.import_module('PySDM.initialisation.spectra').Lognormal;
equilibrate_wet_radii = py.importlib.import_module('PySDM.initialisation').equilibrate_wet_radii;
CPU = py.importlib.import_module('PySDM.backends').CPU;
AmbientThermodynamics = py.importlib.import_module('PySDM.dynamics').AmbientThermodynamics;
Condensation = py.importlib.import_module('PySDM.dynamics').Condensation;
Parcel = py.importlib.import_module('PySDM.environments').Parcel;
Builder = py.importlib.import_module('PySDM').Builder;
Formulae = py.importlib.import_module('PySDM').Formulae;
products = py.importlib.import_module('PySDM.products');
env = Parcel(pyargs( ...
'dt', .25 * si.s, ...
'mass_of_dry_air', 1e3 * si.kg, ...
'p0', 1122 * si.hPa, ...
'initial_water_vapour_mixing_ratio', 20 * si.g / si.kg, ...
'T0', 300 * si.K, ...
'w', 2.5 * si.m / si.s ...
));
spectrum = Lognormal(pyargs('norm_factor', 1e4/si.mg, 'm_mode', 50 * si.nm, 's_geom', 1.4));
kappa = .5;
cloud_range = py.tuple({.5 * si.um, 25 * si.um});
output_interval = 4;
output_points = 40;
n_sd = 256;
formulae = Formulae();
builder = Builder(pyargs('backend', CPU(formulae), 'n_sd', int32(n_sd), 'environment', env));
builder.add_dynamic(AmbientThermodynamics());
builder.add_dynamic(Condensation());
tmp = spectral_sampling.Logarithmic(spectrum).sample(int32(n_sd));
r_dry = tmp{1};
v_dry = formulae.trivia.volume(pyargs('radius', r_dry));
specific_concentration = tmp{2};
r_wet = equilibrate_wet_radii(pyargs(...
'r_dry', r_dry, ...
'environment', env, ...
'kappa_times_dry_volume', kappa * v_dry...
));
attributes = py.dict(pyargs( ...
'multiplicity', discretise_multiplicities(specific_concentration * env.mass_of_dry_air), ...
'dry volume', v_dry, ...
'kappa times dry volume', kappa * v_dry, ...
'volume', formulae.trivia.volume(pyargs('radius', r_wet)) ...
));
particulator = builder.build(attributes, py.list({ ...
products.PeakSupersaturation(pyargs('name', 'S_max', 'unit', '%')), ...
products.EffectiveRadius(pyargs('name', 'r_eff', 'unit', 'um', 'radius_range', cloud_range)), ...
products.ParticleConcentration(pyargs('name', 'n_c_cm3', 'unit', 'cm^-3', 'radius_range', cloud_range)), ...
products.WaterMixingRatio(pyargs('name', 'liquid water mixing ratio', 'unit', 'g/kg', 'radius_range', cloud_range)) ...
products.ParcelDisplacement(pyargs('name', 'z')) ...
}));
cell_id = int32(0);
output_size = [output_points+1, length(py.list(particulator.products.keys()))];
output_types = repelem({'double'}, output_size(2));
output_names = [cellfun(@string, cell(py.list(particulator.products.keys())))];
output = table(...
'Size', output_size, ...
'VariableTypes', output_types, ...
'VariableNames', output_names ...
);
for pykey = py.list(keys(particulator.products))
get = py.getattr(particulator.products{pykey{1}}.get(), '__getitem__');
key = string(pykey{1});
output{1, key} = get(cell_id);
end
for i=2:output_points+1
particulator.run(pyargs('steps', int32(output_interval)));
for pykey = py.list(keys(particulator.products))
get = py.getattr(particulator.products{pykey{1}}.get(), '__getitem__');
key = string(pykey{1});
output{i, key} = get(cell_id);
end
end
i=1;
for pykey = py.list(keys(particulator.products))
product = particulator.products{pykey{1}};
if string(product.name) ~= "z"
subplot(1, width(output)-1, i);
plot(output{:, string(pykey{1})}, output.z, '-o');
title(string(product.name), 'Interpreter', 'none');
xlabel(string(product.unit));
end
if i == 1
ylabel(string(particulator.products{"z"}.unit));
end
i=i+1;
end
saveas(gcf, "parcel.png");Python (click to expand)
from matplotlib import pyplot
from PySDM.physics import si
from PySDM.initialisation import discretise_multiplicities, equilibrate_wet_radii
from PySDM.initialisation.spectra import Lognormal
from PySDM.initialisation.sampling import spectral_sampling
from PySDM.backends import CPU
from PySDM.dynamics import AmbientThermodynamics, Condensation
from PySDM.environments import Parcel
from PySDM import Builder, Formulae, products
env = Parcel(
dt=.25 * si.s,
mass_of_dry_air=1e3 * si.kg,
p0=1122 * si.hPa,
initial_water_vapour_mixing_ratio=20 * si.g / si.kg,
T0=300 * si.K,
w=2.5 * si.m / si.s
)
spectrum = Lognormal(norm_factor=1e4 / si.mg, m_mode=50 * si.nm, s_geom=1.5)
kappa = .5 * si.dimensionless
cloud_range = (.5 * si.um, 25 * si.um)
output_interval = 4
output_points = 40
n_sd = 256
formulae = Formulae()
builder = Builder(backend=CPU(formulae), n_sd=n_sd, environment=env)
builder.add_dynamic(AmbientThermodynamics())
builder.add_dynamic(Condensation())
r_dry, specific_concentration = spectral_sampling.Logarithmic(spectrum).sample(n_sd)
v_dry = formulae.trivia.volume(radius=r_dry)
r_wet = equilibrate_wet_radii(r_dry=r_dry, environment=env, kappa_times_dry_volume=kappa * v_dry)
attributes = {
'multiplicity': discretise_multiplicities(specific_concentration * env.mass_of_dry_air),
'dry volume': v_dry,
'kappa times dry volume': kappa * v_dry,
'volume': formulae.trivia.volume(radius=r_wet)
}
particulator = builder.build(attributes, products=[
products.PeakSupersaturation(name='S_max', unit='%'),
products.EffectiveRadius(name='r_eff', unit='um', radius_range=cloud_range),
products.ParticleConcentration(name='n_c_cm3', unit='cm^-3', radius_range=cloud_range),
products.WaterMixingRatio(name='liquid water mixing ratio', unit='g/kg', radius_range=cloud_range),
products.ParcelDisplacement(name='z')
])
cell_id = 0
output = {product.name: [product.get()[cell_id]] for product in particulator.products.values()}
for step in range(output_points):
particulator.run(steps=output_interval)
for product in particulator.products.values():
output[product.name].append(product.get()[cell_id])
fig, axs = pyplot.subplots(1, len(particulator.products) - 1, sharey="all")
for i, (key, product) in enumerate(particulator.products.items()):
if key != 'z':
axs[i].plot(output[key], output['z'], marker='.')
axs[i].set_title(product.name)
axs[i].set_xlabel(product.unit)
axs[i].grid()
axs[0].set_ylabel(particulator.products['z'].unit)
pyplot.savefig('parcel.svg')The resultant plot (generated with the Matlab code) looks as follows:
Submitting new code to the project, please preferably use GitHub pull requests - it helps to keep record of code authorship, track and archive the code review workflow and allows to benefit from the continuous integration setup which automates execution of tests with the newly added code.
Code contributions are assumed to imply transfer of copyright. Should there be a need to make an exception, please indicate it when creating a pull request or contributing code in any other way. In any case, the license of the contributed code must be compatible with GPL v3.
Developing the code, we follow The Way of Python and the KISS principle. The codebase has greatly benefited from PyCharm code inspections and Pylint, Black and isort code analysis (which are all part of the CI workflows).
We also use pre-commit hooks.
In our case, the hooks modify files and re-format them.
The pre-commit hooks can be run locally, and then the resultant changes need to be staged before committing.
To set up the hooks locally, install pre-commit via pip install pre-commit and
set up the git hooks via pre-commit install (this needs to be done every time you clone the project).
To run all pre-commit hooks, run pre-commit run --all-files.
The .pre-commit-config.yaml file can be modified in case new hooks are to be added or
existing ones need to be altered.
Further hints addressed at PySDM developers are maintained in the open-atmos/python-dev-hints Wiki.
Issues regarding any incorrect, unintuitive or undocumented bahaviour of PySDM are best to be reported on the GitHub issue tracker. Feature requests are recorded in the "Ideas..." PySDM wiki page.
We encourage to use the GitHub Discussions feature (rather than the issue tracker) for seeking support in understanding, using and extending PySDM code.
We look forward to your contributions and feedback.
The development and maintenance of PySDM is led by Sylwester Arabas. Piotr Bartman had been the architect and main developer of technological solutions in PySDM. PySDM includes contributions from researchers from Jagiellonian University departments of computer science, physics and chemistry; from Caltech's Climate Modelling Alliance, from University of Warsaw (dept. physics), and from AGH University of Krakow (dept. physics & applied computer science) where release maintenance takes place currently.
Development of PySDM had been initially supported by the EU through a grant of the Foundation for Polish Science) (grant no. POIR.04.04.00-00-5E1C/18) realised at the Jagiellonian University. The immersion freezing support in PySDM was developed with support from the US Department of Energy Atmospheric System Research programme through a grant (no. DE-SC0021034) realised at the University of Illinois at Urbana-Champaign. Development of isotopic fractionation representation and mixed-phase support is carried out with support from the Polish National Science Centre (grant no. 2020/39/D/ST10/01220).
copyright: Jagiellonian University (2019-2023) & AGH University of Krakow (2023-...)
licence: GPL v3
- https://patents.google.com/patent/US7756693B2
- https://patents.google.com/patent/EP1847939A3
- https://patents.google.com/patent/JP4742387B2
- https://patents.google.com/patent/CN101059821B
- SCALE-SDM (Fortran):
https://github.com/Shima-Lab/SCALE-SDM_BOMEX_Sato2018/blob/master/contrib/SDM/sdm_coalescence.f90 - Pencil Code (Fortran):
https://github.com/pencil-code/pencil-code/blob/master/src/particles_coagulation.f90 - PALM LES (Fortran):
https://palm.muk.uni-hannover.de/trac/browser/palm/trunk/SOURCE/lagrangian_particle_model_mod.f90 - libcloudph++ (C++):
https://github.com/igfuw/libcloudphxx/blob/master/src/impl/particles_impl_coal.ipp - LCM1D (Python)
https://github.com/SimonUnterstrasser/ColumnModel - superdroplet (Cython/Numba/C++11/Fortran 2008/Julia)
https://github.com/darothen/superdroplet - NTLP (FORTRAN)
https://github.com/Folca/NTLP/blob/SuperDroplet/les.F - CLEO (C++)
https://yoctoyotta1024.github.io/CLEO/ - droplets.jl (Julia)
https://github.com/emmacware/droplets.jl
- PartMC (Fortran):
https://github.com/compdyn/partmc
- pyrcel: https://github.com/darothen/pyrcel
- PyBox: https://github.com/loftytopping/PyBox
- py-cloud-parcel-model: https://github.com/emmasimp/py-cloud-parcel-model
- CloudMicrophysics.jl: https://github.com/CliMA/CloudMicrophysics.jl