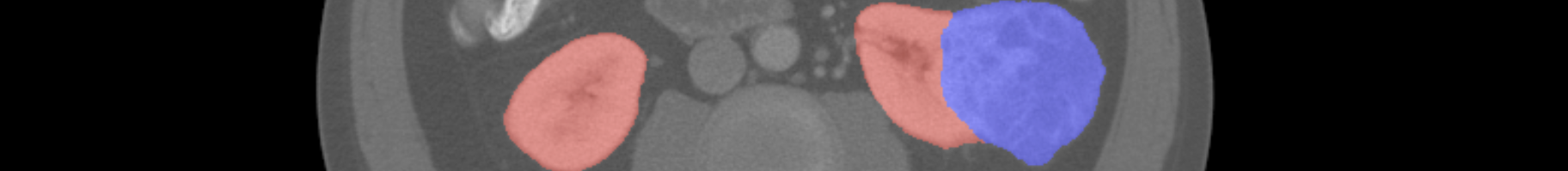
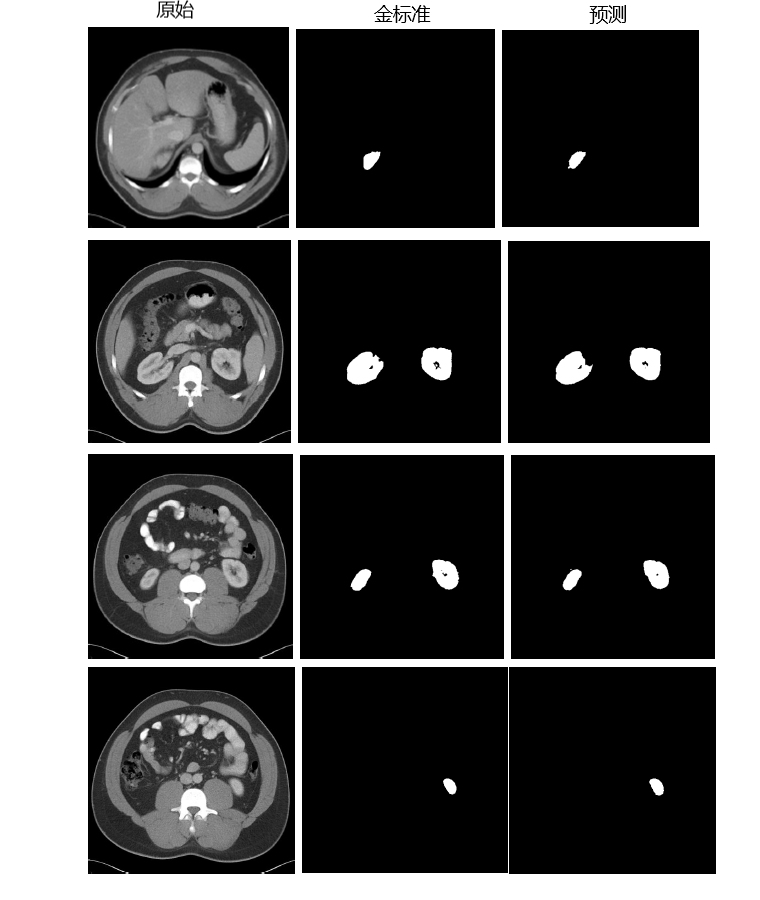
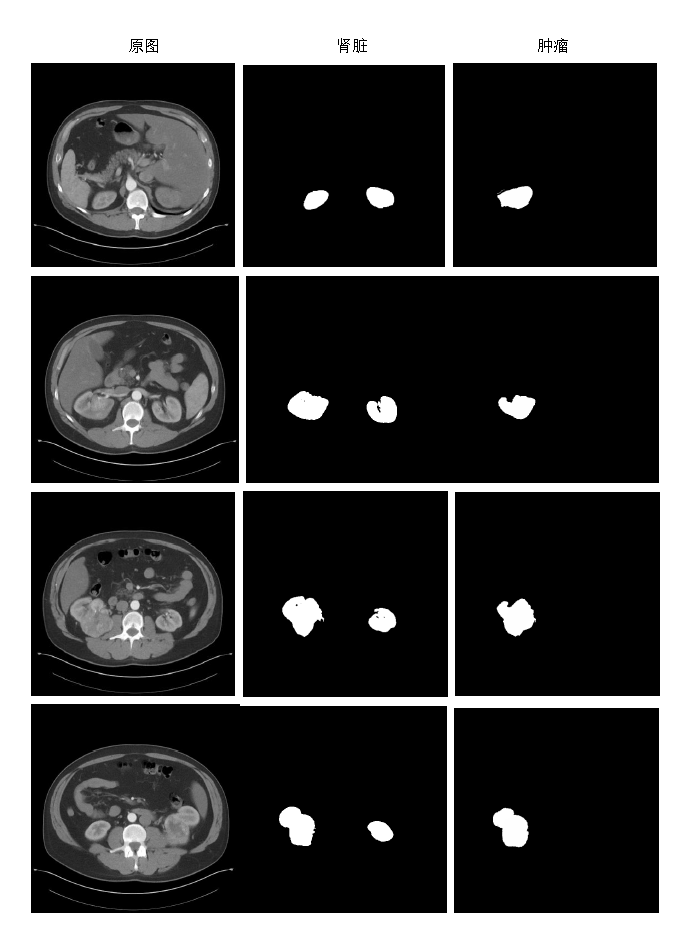
This is an example of the CT images Kidney Tumor Segmentation
The following dependencies are needed:
- python == 3.5.5
- numpy >= 1.11.1
- SimpleITK >= 1.0.1
- opencv-python >= 3.3.0
- tensorflow-gpu == 1.8.0
- pandas >=0.20.1
- scikit-learn >= 0.17.1
- json >=2.0.9
1、Preprocess
- analyze the ct image,and get the slice thickness and window width and position:run the dataAnaly.py
1.1 Preprocess Kidney
- keep kidney region into fixed size(512,512,64) for Corse Kidney Segmentation:run the corsedata2dprepare.py
- generate patch(128,128,64) kidney image and mask for Corse Kidney Segmentation:run the corsedata3dprepare.py
- keep Kidney region range image for fine Kidney Segmentation:run the finedata2dprepare.py
- generate patch(128,128,64) kidney image and mask for fine Kidney Segmentation:run the finedata3dprepare.py
- save patch image and mask into csv file: run the utils.py,like file trainSegmentation.csv
- split trainSegmentation.csv into training set and test set:run subset.py
1.2 Preprocess Kidney Tumor
- generate tumor image and mask for 2d Kidney Tumor Segmentation:run the tumordata2dprepare.py
- generate tumor image and mask for 3d Kidney Tumor Segmentation:run the finedata3dprepare.py
- save tumor image and mask path into csv file: run the utils.py,like file traintumorSegmentation.csv
- split traintumorSegmentation.csv into training set and test set
2、Kidney Segmentation
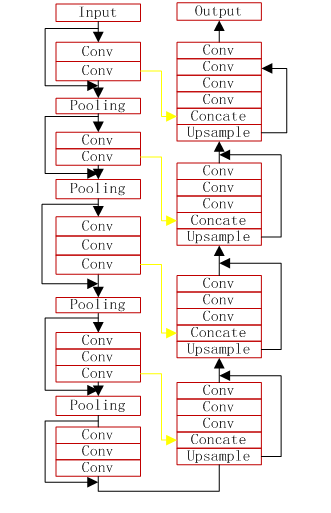
- the VNet model
2.1 Corse Kidney Segmentation
- Corse Kidney Segmentation training:run the train_vnet3d_kidney_corse.py
- Corse Kidney Segmentation inference:run the inference_vnet3d_kidney_corse.py
- this step get Corse Kidney range,can find the start and end pos in the kidneyrang.txt
2.2 Fine Kidney Segmentation
- Fine Kidney Segmentation training:run the train_vnet3d_kidney_fine.py
- Fine Kidney Segmentation inference:run the inference_vnet3d_kidney_fine.py
- this step following the 2.1 result,get fine Kidney result
2.3 Fine Kidney Segmentation
- remove Kidney Segmentation small object:run the segresultprocess.py removekidneysmallobj function
3、Kidney Tumor Segmentation
- the VNet2d model
3.1 2d Kidney Tumor Segmentation
-
2d Kidney Tumor Segmentation training:run the train_vnet2d_tumor.py
-
2d Kidney Tumor Segmentation inference:run the inference_vnet2d_tumor.py
-
this step get 2d slice tumor result
-
the VNet3d model
3.2 3d Kidney Tumor Segmentation
- 3d Kidney Tumor Segmentation training:run the train_vnet3d_tumor.py
- 3d Kidney Tumor Segmentation inference:run the inference_vnet3d_tumor.py
- this step get 3d tumor result
3.3 Kidney Tumor Result Process
- remove Kidney Tumor Segmentation small object:run the segresultprocess.py remove2d3dtumorsmallobj function
- calculate overlap between 2d tumor and 3d tumor reslut.
- save the region of 2d tumor and 3d tumor reslut that connect overlap region.
- save the region of 2d tumor and 3d tumor within Kidney result.
- merge the above two result and get final tumor result
- all above step can find in the segresultprocess.py
1、Kidney Segmentation
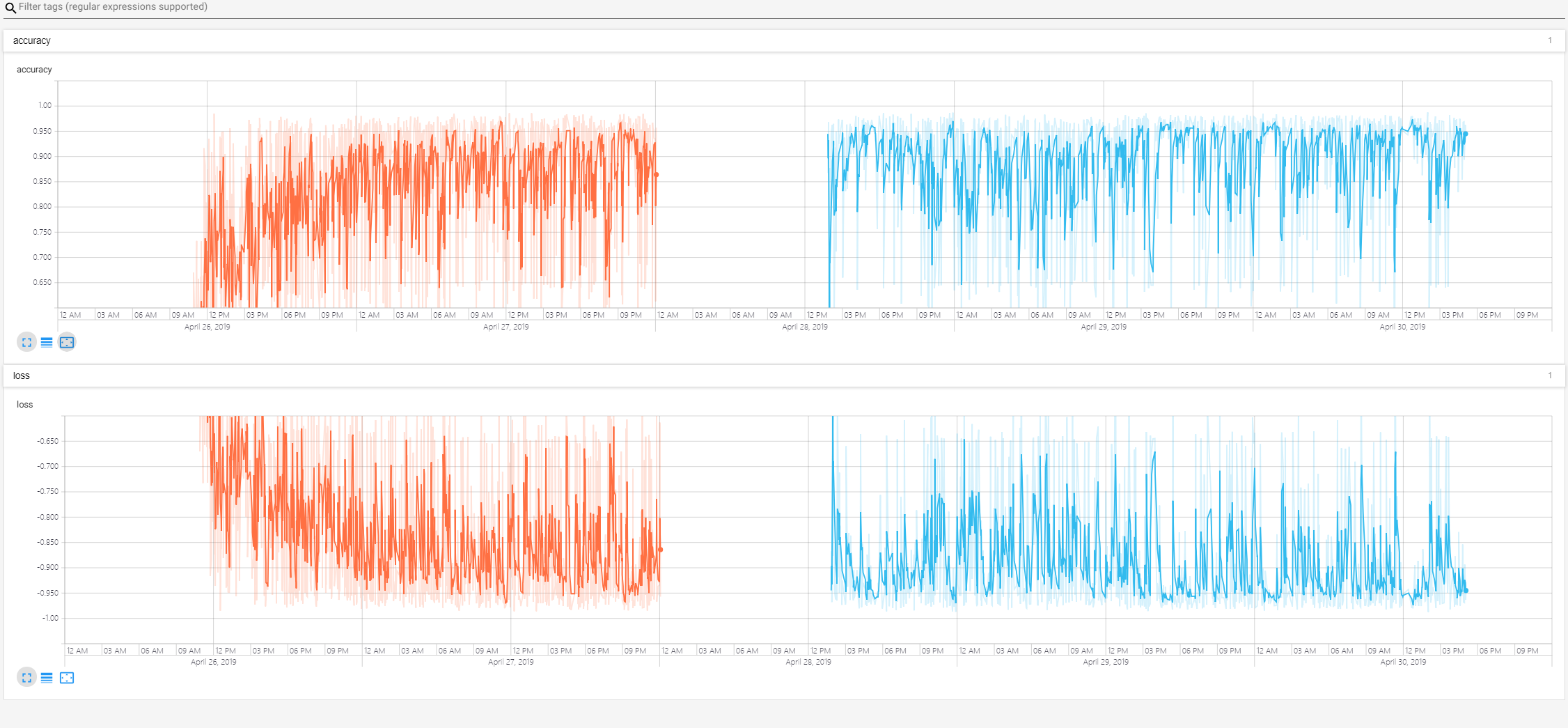
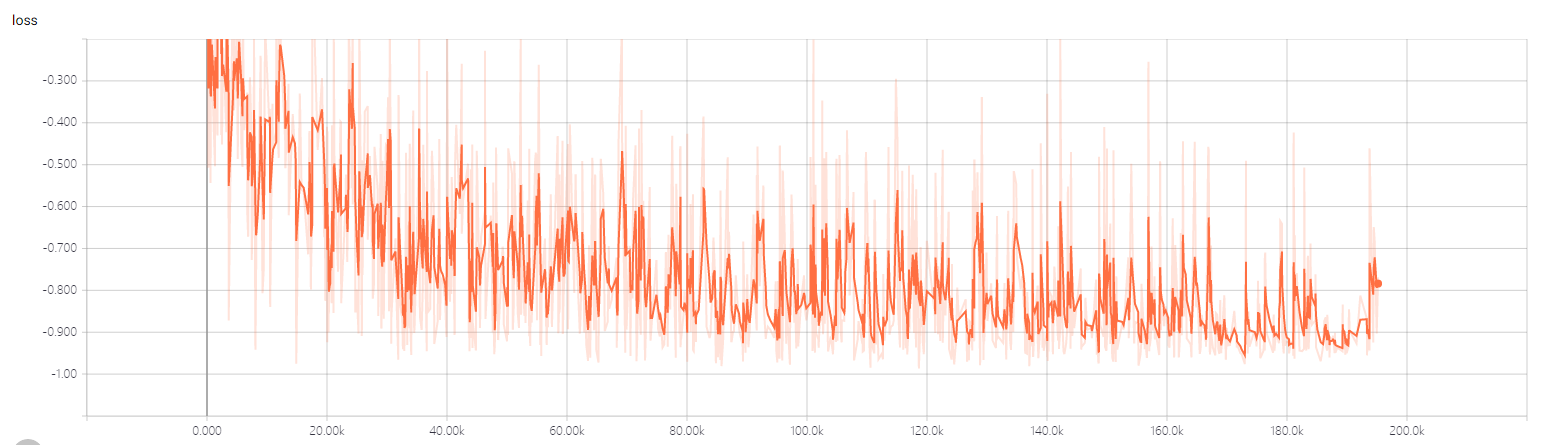
- the train loss
- 200-209case dice value and result
2、Kidney Tumor Segmentation
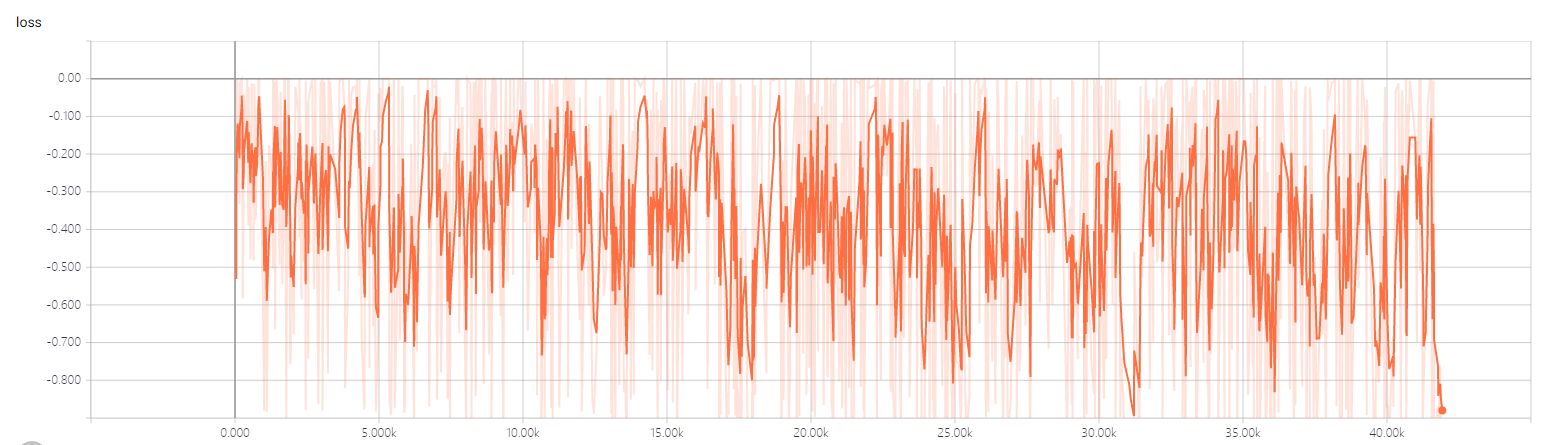
- the 2dtrain loss
- the 3dtrain loss
3、Lead Board
- https://github.com/junqiangchen
- email: 1207173174@qq.com,1259389904@qq.com
- Contact: junqiangChen,junMa(马骏)
- WeChat Number: 1207173174
- WeChat Public number: 最新医学影像技术