FIELDimageR.Extra: An package with new tools to support FIELDimageR software on evaluating GIS images from agriculture field trials using R.
This package was developed by Popat Pawar and Filipe Matias to help FIELDimageR users with new functions to analyze orthomosaic images from research fields. Cropping and rotating images are not necessary anymore, and there are amazing zoom capabilities provided by newly released GIS R packages. Come along with us and try this tutorial out ... We hope it helps with your research!
Step 2. Loading mosaics and visualizing
Step 3. Building the plot shapefile (Zoom)
Step 4. Editing the plot shapefile (Zoom)
Step 5. Building vegetation indices and removing the soil effect
Step 6. Extracting data from field images
Step 7. Vizualizing extracted data
Step 8. Saving output files and opening them in the QGIS
Step 9. Cropping individual plots and saving
First of all, install R and RStudio. Then, in order to install R/FIELDimageR.Extra from GitHub GitHub repository, you need to install the following packages in R. For Windows users who have an R version higher than 4.0, you need to install RTools, tutorial RTools For Windows.
Now install R/FIELDimageR.Extra using the
install_githubfunction from devtools package. If necessary, use the argument type="source".
install.packages("devtools")
devtools::install_github("filipematias23/FIELDimageR.Extra")If the method above doesn't work, use the next lines by downloading the FIELDimageR-master.zip file
setwd("~/FIELDimageR.Extra-main.zip") # ~ is the path from where you saved the file.zip
unzip("FIELDimageR.Extra-main.zip")
file.rename("FIELDimageR.Extra-main", "FIELDimageR.Extra")
shell("R CMD build FIELDimageR.Extra") # or system("R CMD build FIELDimageR.Extra")
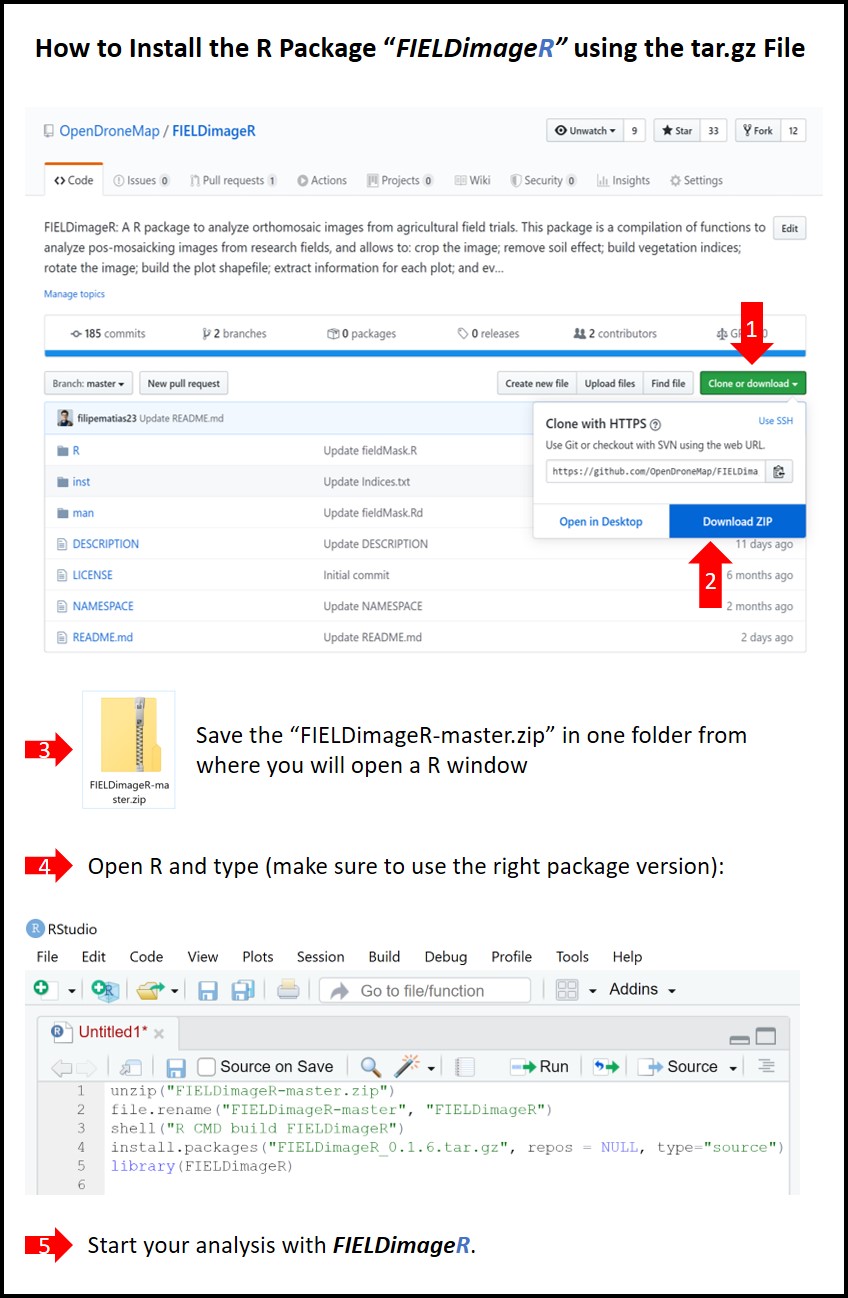
install.packages("FIELDimageR.Extra_0.0.1.tar.gz", repos = NULL, type="source") # Make sure to use the right version (e.g. 0.0.1)The image below from R/FIELDimageR is highlighting how to do the steps described above and install R/FIELDimageR.Extra using source code.
Reading necessary packages:
# Install:
install.packages(c('terra','mapview','sf','stars'))
devtools::install_github("OpenDroneMap/FIELDimageR")
# Packages:
library(FIELDimageR)
library(FIELDimageR.Extra)
library(terra)
library(mapview)
library(sf)
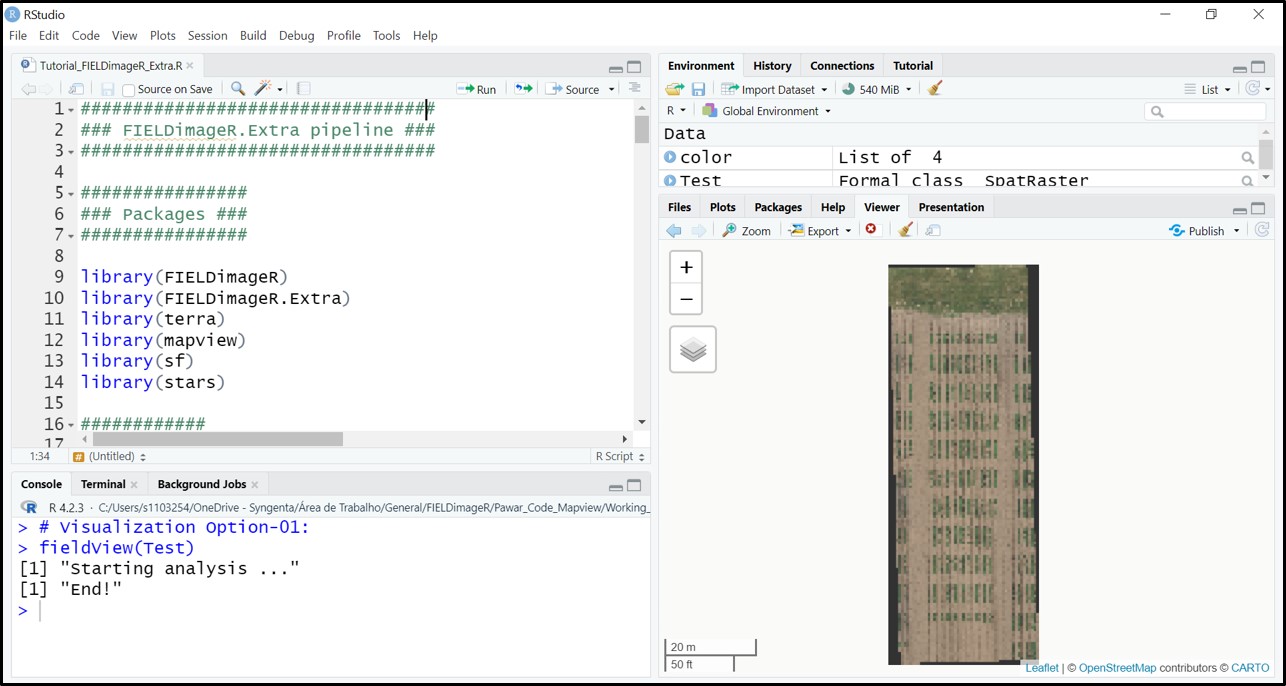
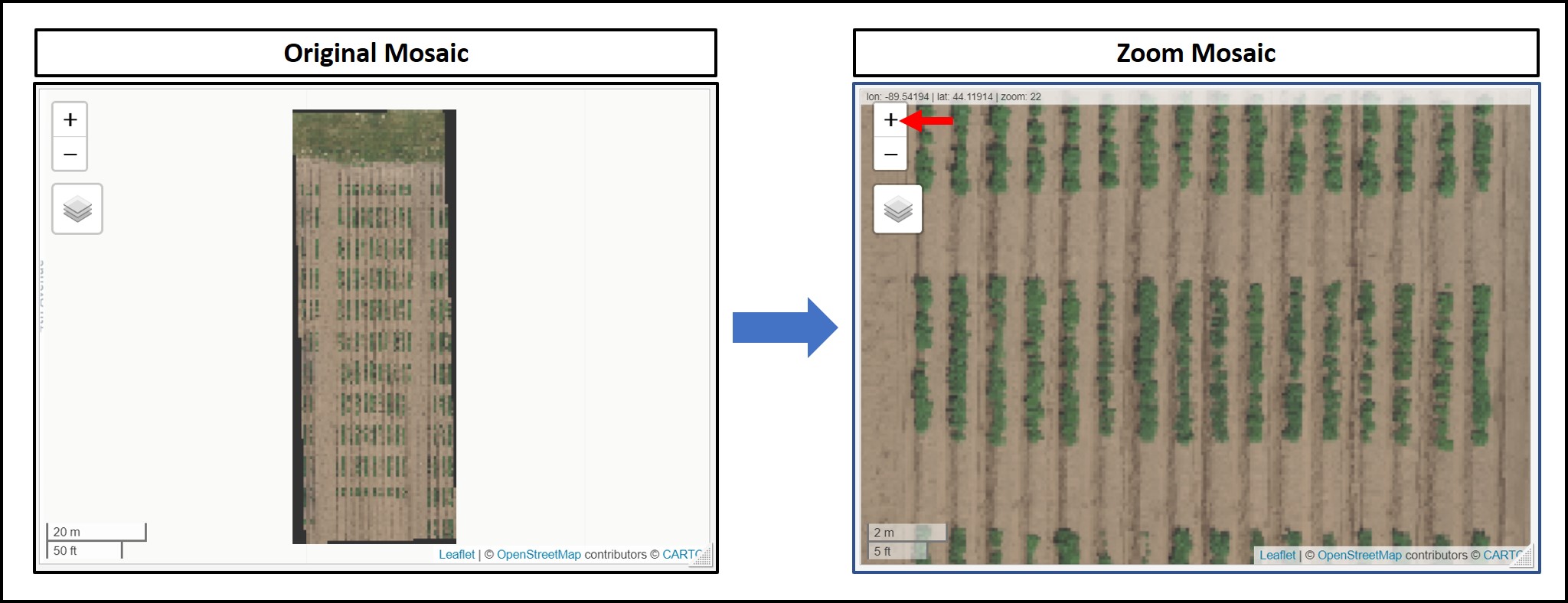
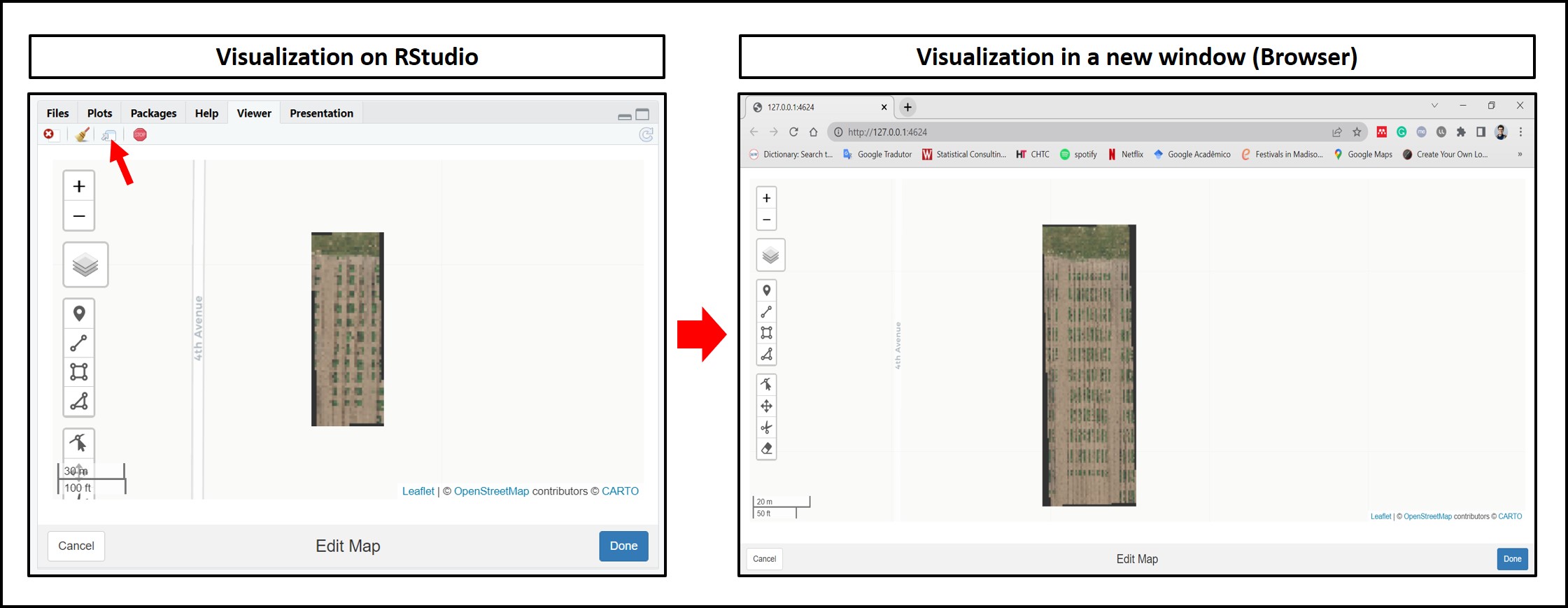
library(stars)The following example uses an image available to download here: EX1_RGB.tif. The first R/FIELDimageR.Extra function is
fieldViewused to visualize GIS images and grid shapefiles and zoom it out.
# Uploading an example mosaic
Test <- rast("EX1_RGB.tif")
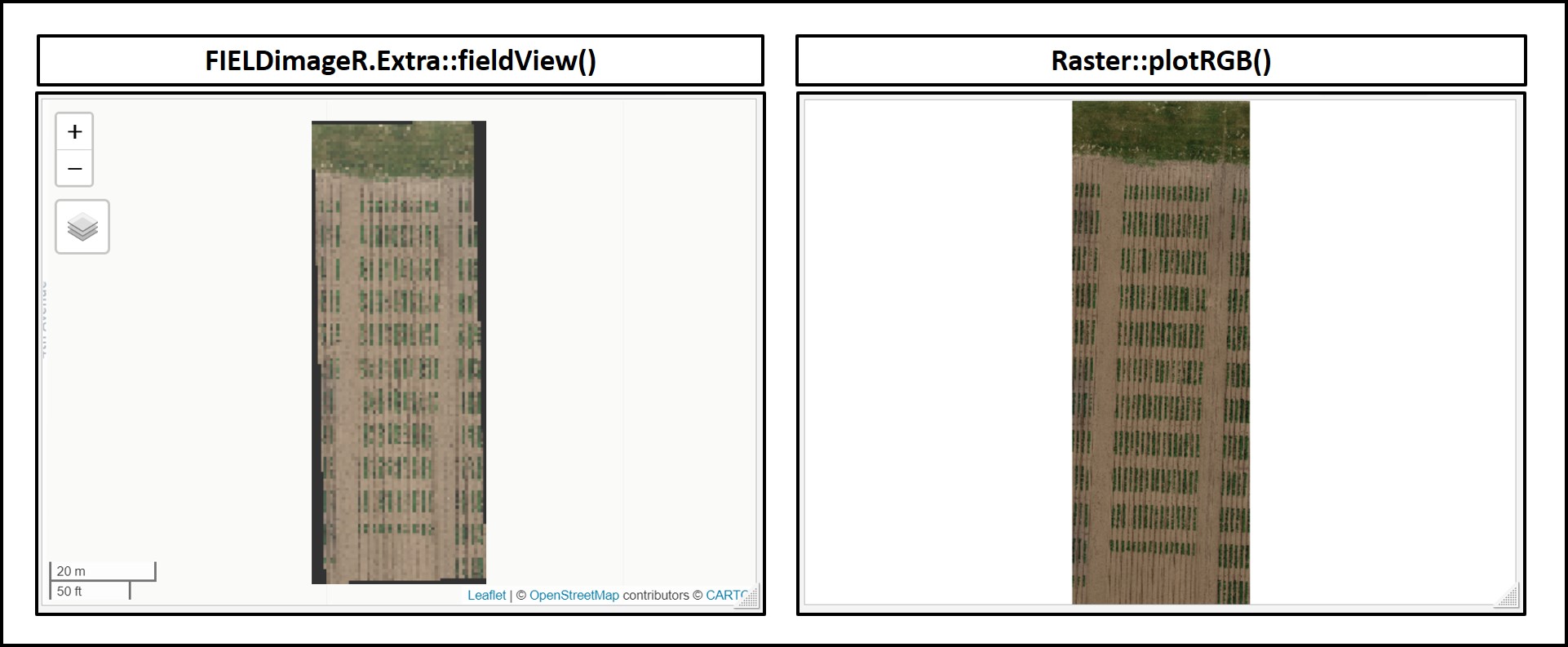
# Visualization Option-01 (FIELDimageR.Extra):
fieldView(Test)
# Visualization Option-02 (raster):
plotRGB(Test)
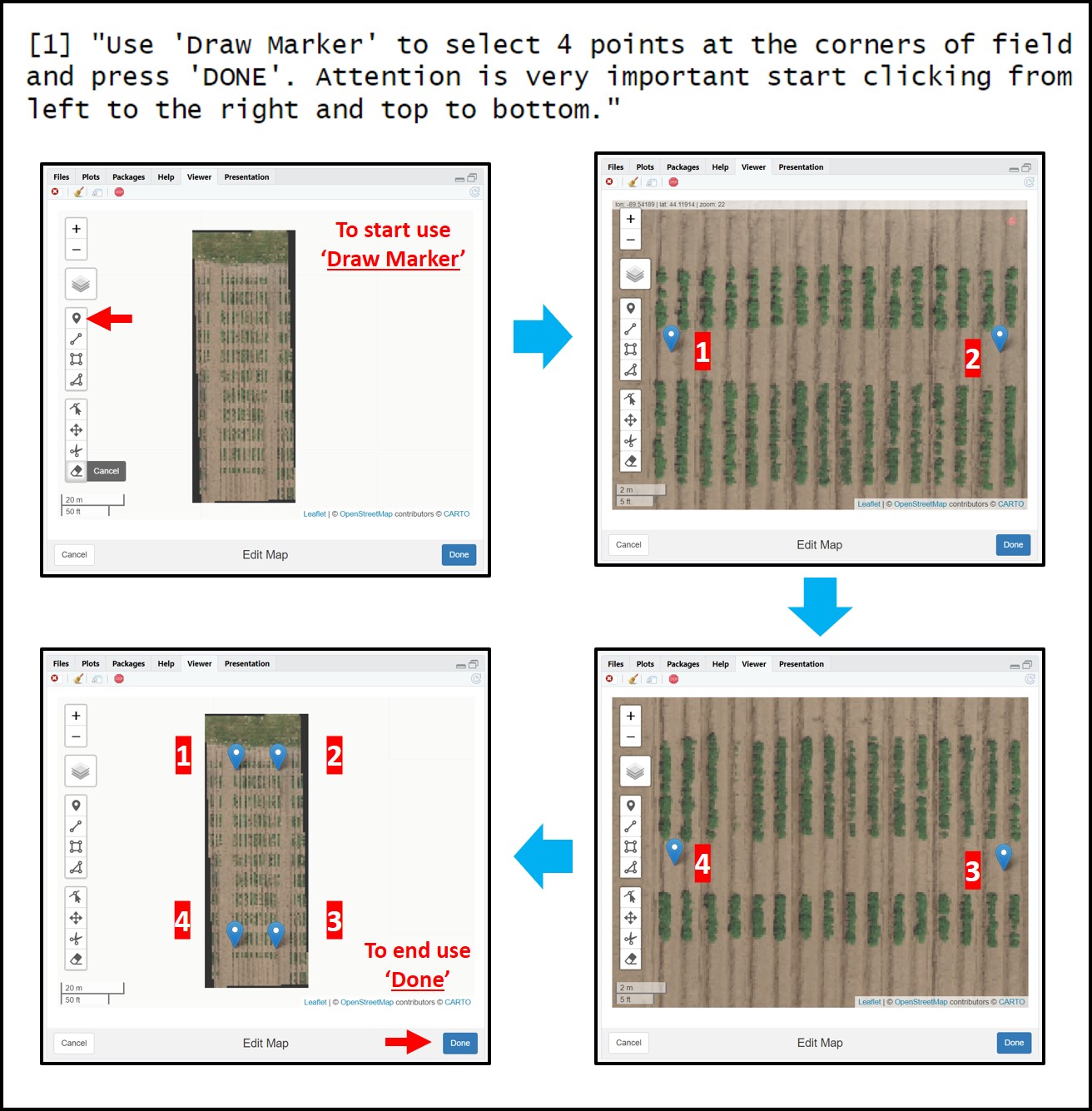
FIELDimageR.Extra allows drawing the plot shape file using the function
fieldShape_render. Different fromFIELDimageR::fieldShapethis new function does not request straight fields and the experimental trial can be used in any direction according to the original GIS position. It is very important to highlight that four points need to be set at the corners of the trial according to the following sequence (1st point) left superior corner, (2nd point) right superior corner, (3rd point) right inferior corner, and (4th point) left inferior corner. The mosaic will be presented for visualization with the North part on the superior part (top) and the south in )the inferior part (bottom). The number of columns and rows must be informed. At this point, the experimental borders can be eliminated (check the example below.
plotShape<-fieldShape_render(mosaic = Test,
ncols=14,
nrows=9)
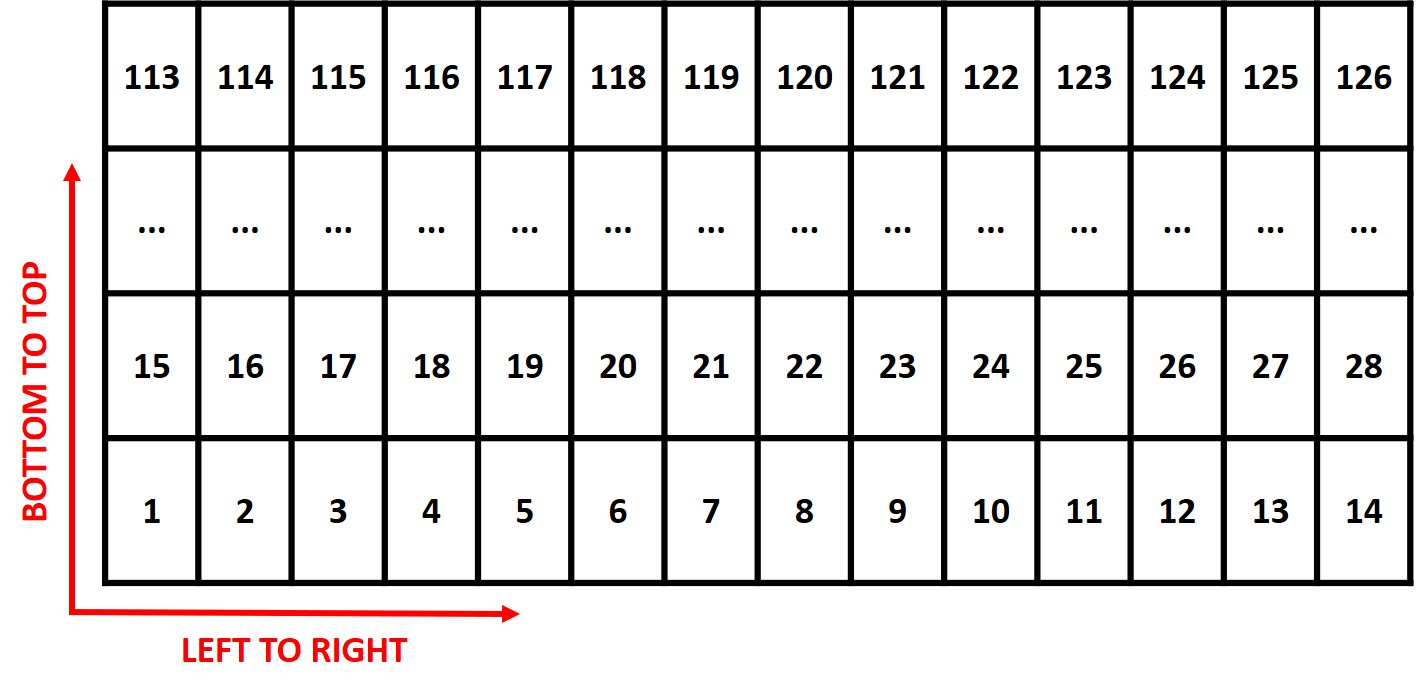
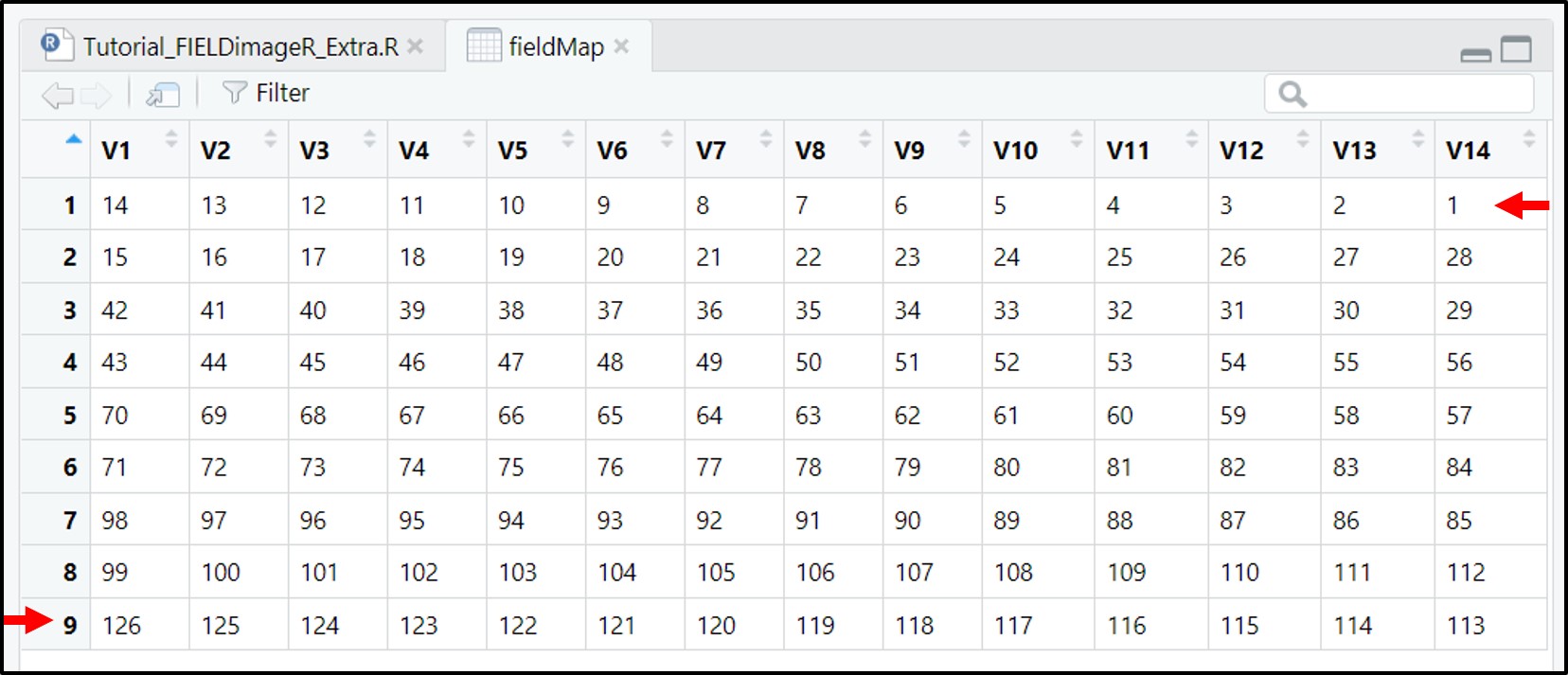
Attention: The plots are identified in ascending order from left to right and bottom to top (this is another difference from the
FIELDimageR::fieldShape) being evenly spaced and distributed inside the selected area independent of alleys.
One matrix can be used to identify the plots position according to the image above. The function
FIELDimageR::fieldMapcan be used to specify the plot ID automatic or any other matrix (manually built) also can be used. For instance, the new column PlotID will be the new identification. You can download an external table example here: DataTable.csv.
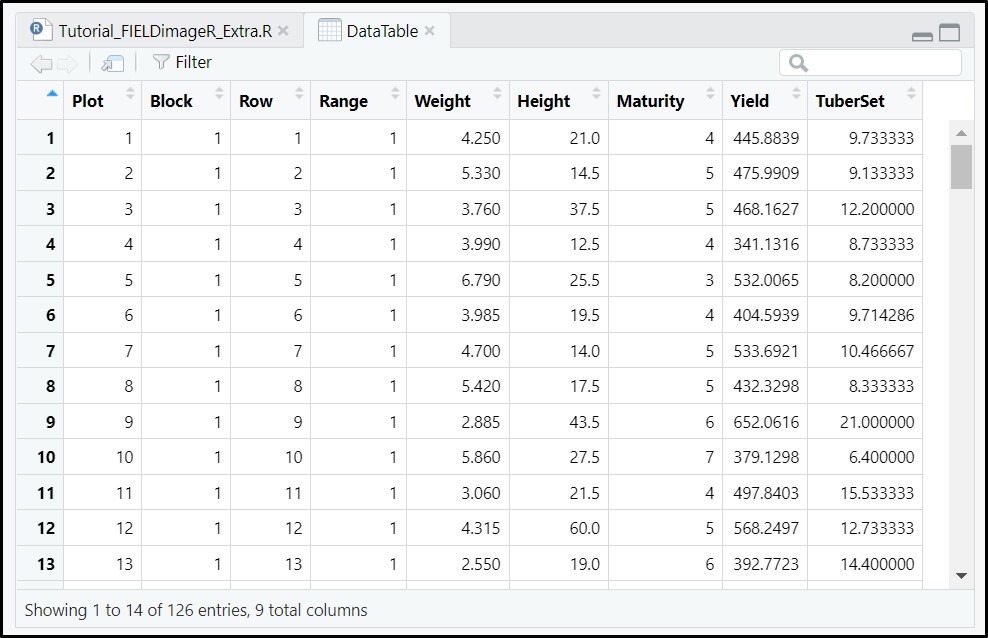
# Reading DataTable.csv
DataTable<-read.csv("DataTable.csv",header = T)
DataTable### Field map identification (name for each Plot). 'fieldPlot' argument can be a number or name.
fieldMap<-fieldMap(fieldPlot=DataTable$Plot, fieldColumn=DataTable$Row, fieldRow=DataTable$Range, decreasing=T)
fieldMap
# The new column PlotID is identifying the plots:
plotShape<-fieldShape_render(mosaic = Test,
ncols=14,
nrows=9,
fieldMap = fieldMap,
buffer = -0.05)### Joing all information from 'fieldData' in one "fieldShape" file using the column 'Plot' as reference:
plotShape<-fieldShape_render(mosaic = Test,
ncols=14,
nrows=9,
fieldData = DataTable,
fieldMap = fieldMap,
PlotID = "Plot",
buffer = -0.05)
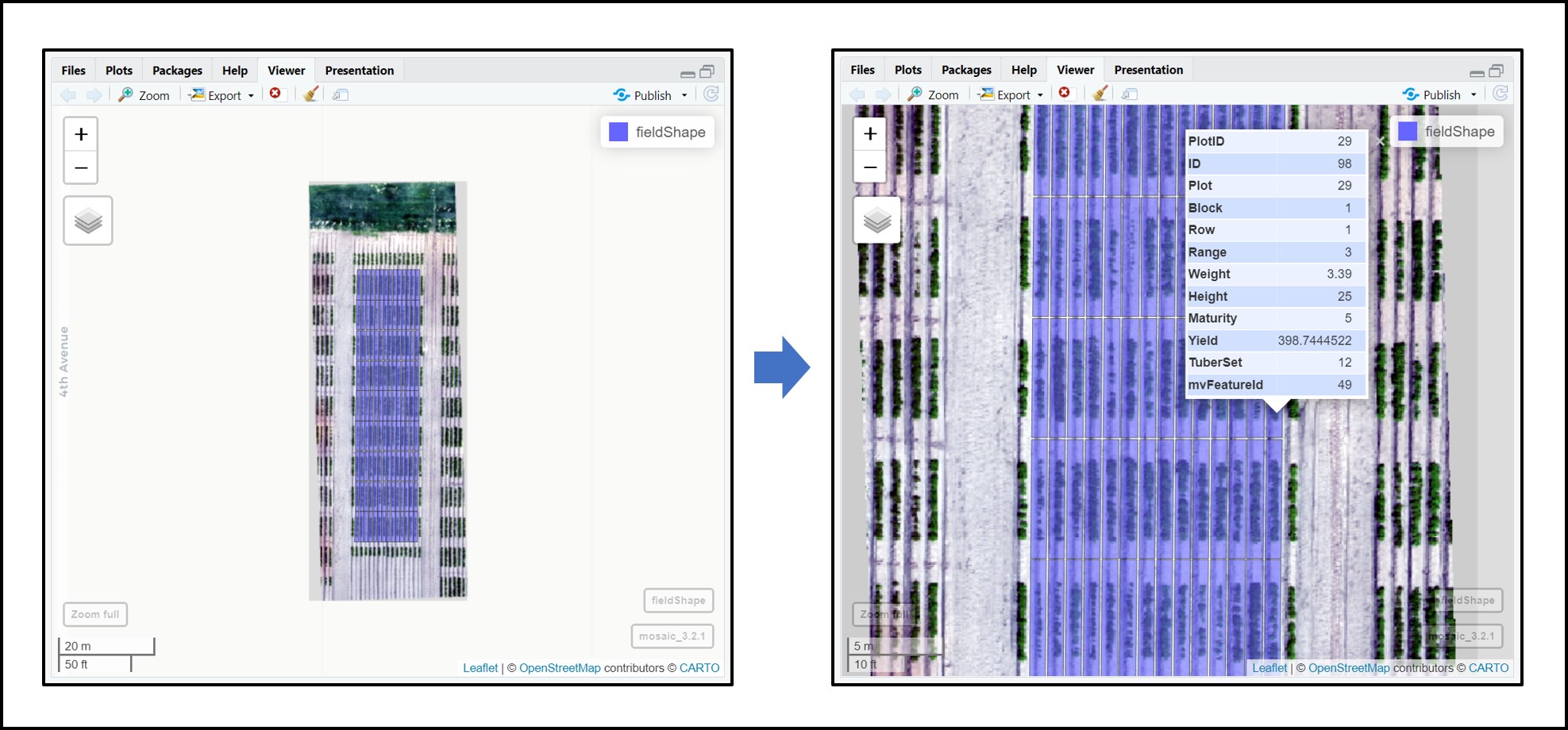
# The new column PlotID is identifying the plots:
plotShape
# Vizualizing new plot grid shape object with field data:
fieldView(mosaic = Test,
fieldShape = plotShape,
type = 2,
alpha = 0.2)
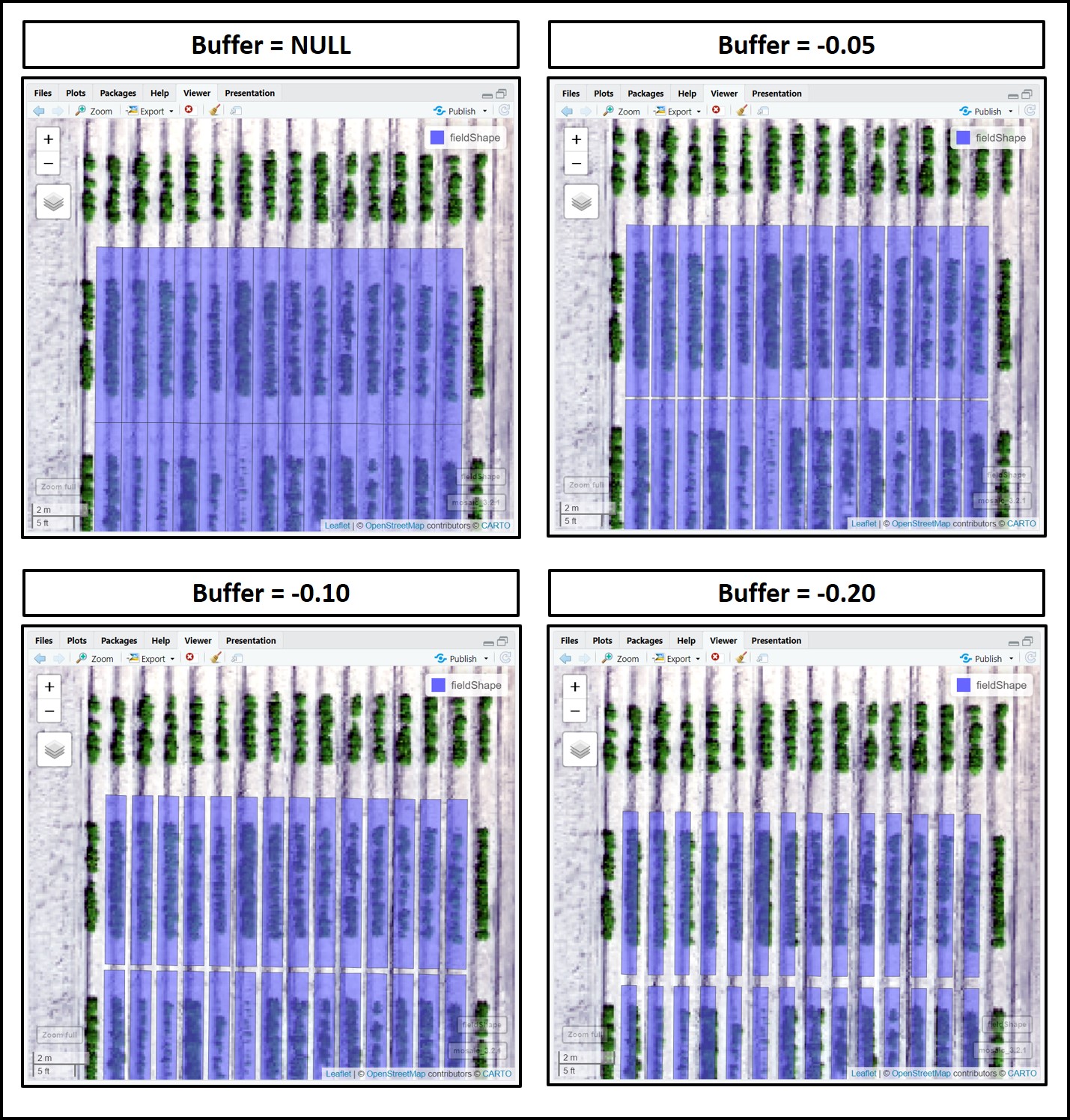
### Different plot dimensions using "Buffer":
# Buffer = NULL
Buffer.1<-fieldShape_render(mosaic = Test,
ncols=14,
nrows=9)
fieldView(mosaic = Test,
fieldShape = Buffer.1,
type = 2,
alpha = 0.2)
# Buffer = -0.05
Buffer.2<-fieldShape_render(mosaic = Test,
ncols=14,
nrows=9,
buffer = -0.05)
fieldView(mosaic = Test,
fieldShape = Buffer.2,
type = 2,
alpha = 0.2)
# Buffer = -0.10
Buffer.3<-fieldShape_render(mosaic = Test,
ncols=14,
nrows=9,
buffer = -0.10)
fieldView(mosaic = Test,
fieldShape = Buffer.3,
type = 2,
alpha = 0.2)
# Buffer = -0.20
Buffer.4<-fieldShape_render(mosaic = Test,
ncols=14,
nrows=9,
buffer = -0.20)
fieldView(mosaic = Test,
fieldShape = Buffer.4,
type = 2,
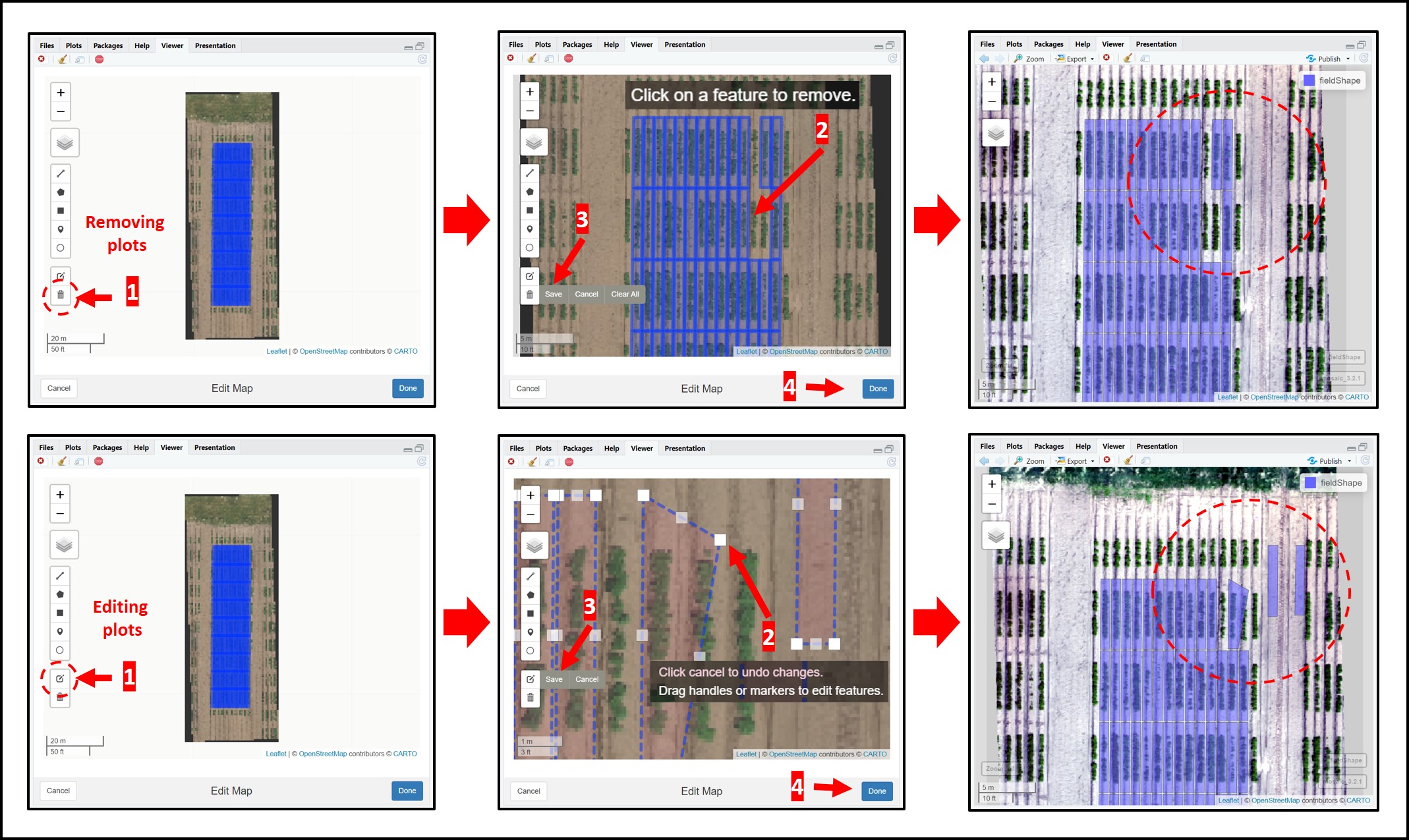
alpha = 0.2) Editing, moving, deleting, and changing single plot grids in fieldShape objects is now possible with the function
fieldShape_edit. Follow the image bellow to understand how this function works. Attention: do not forget to press SAVE before clicking on DONE.
# Editing plot grid shapefile:
editShape<- fieldShape_edit(mosaic=Test,
fieldShape=plotShape)
# Checking the edited plot grid shapefile:
fieldView(mosaic = Test,
fieldShape = editShape,
type = 2,
alpha = 0.2)
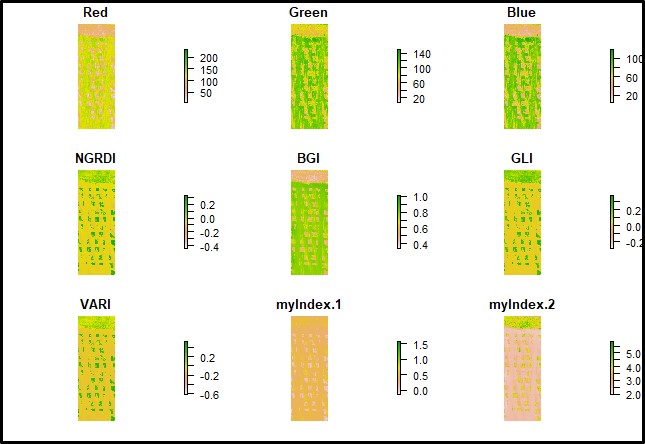
Vegetation indices still being calculated using the function
FIELDimageR::fieldIndexfrom FIELDimageR package.
# Vegetation indices:
Test.Indices<- fieldIndex(mosaic = Test,
Red = 1, Green = 2, Blue = 3,
index = c("NGRDI","BGI", "GLI","VARI"),
myIndex = c("(Red-Blue)/Green","2*Green/Blue"))
# To visualize single band layer:
single_layer<-Test.Indices$GLI
fieldView(single_layer,
fieldShape = editShape,
type = 2,
alpha_grid = 0.2)
fieldView(single_layer,
fieldShape = editShape,
plotCol = c("Yield"),
colorOptions = 'RdYlGn',
type = 1,
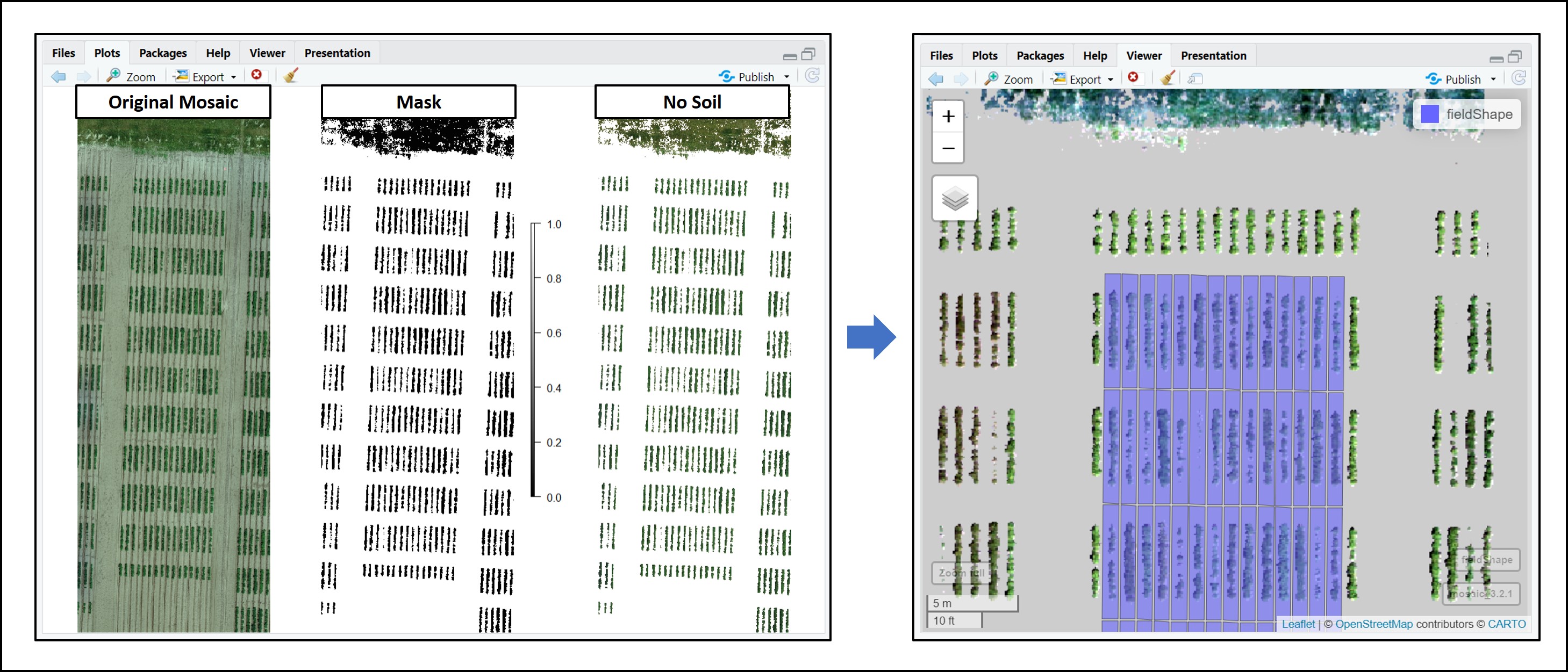
alpha_grid = 0.8)In the same way, removing soil step continues being made by the function
FIELDimageR::fieldMaskfrom FIELDimageR package.
# Removing soil and making a mask
Test.RemSoil<-fieldMask(Test.Indices)
fieldView(Test.RemSoil$newMosaic,
fieldShape = editShape,
type = 2,
alpha_grid = 0.2)
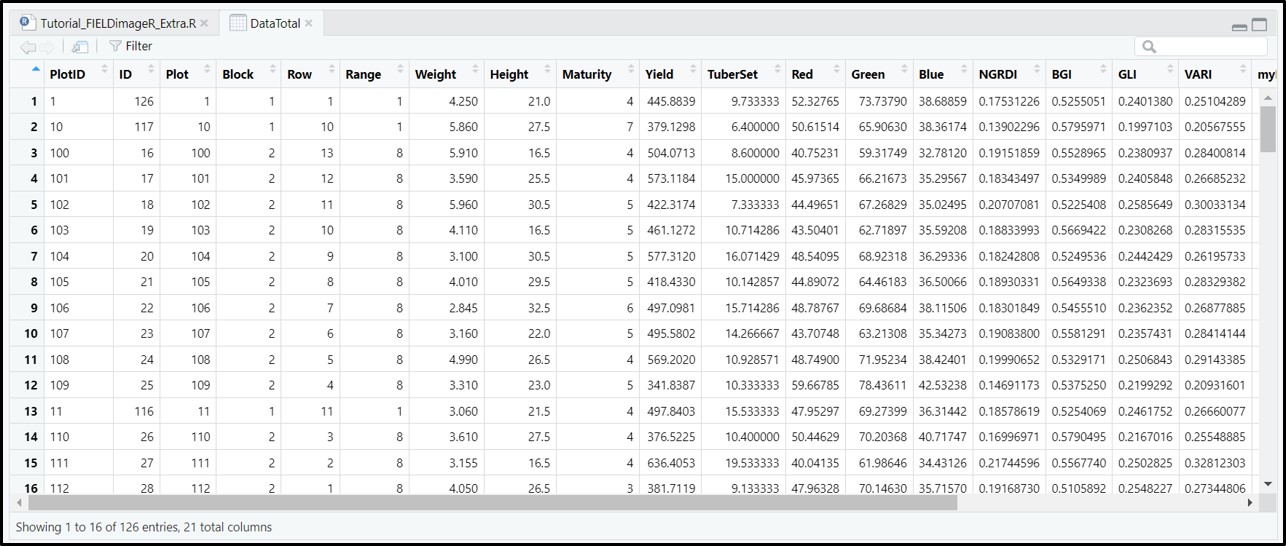
The function aggregate from stars was adapted for agricultural field experiments through function
fieldInfo_extra.
DataTotal<- fieldInfo_extra(mosaic = Test.RemSoil$newMosaic,
fieldShape = editShape,
fun = mean)
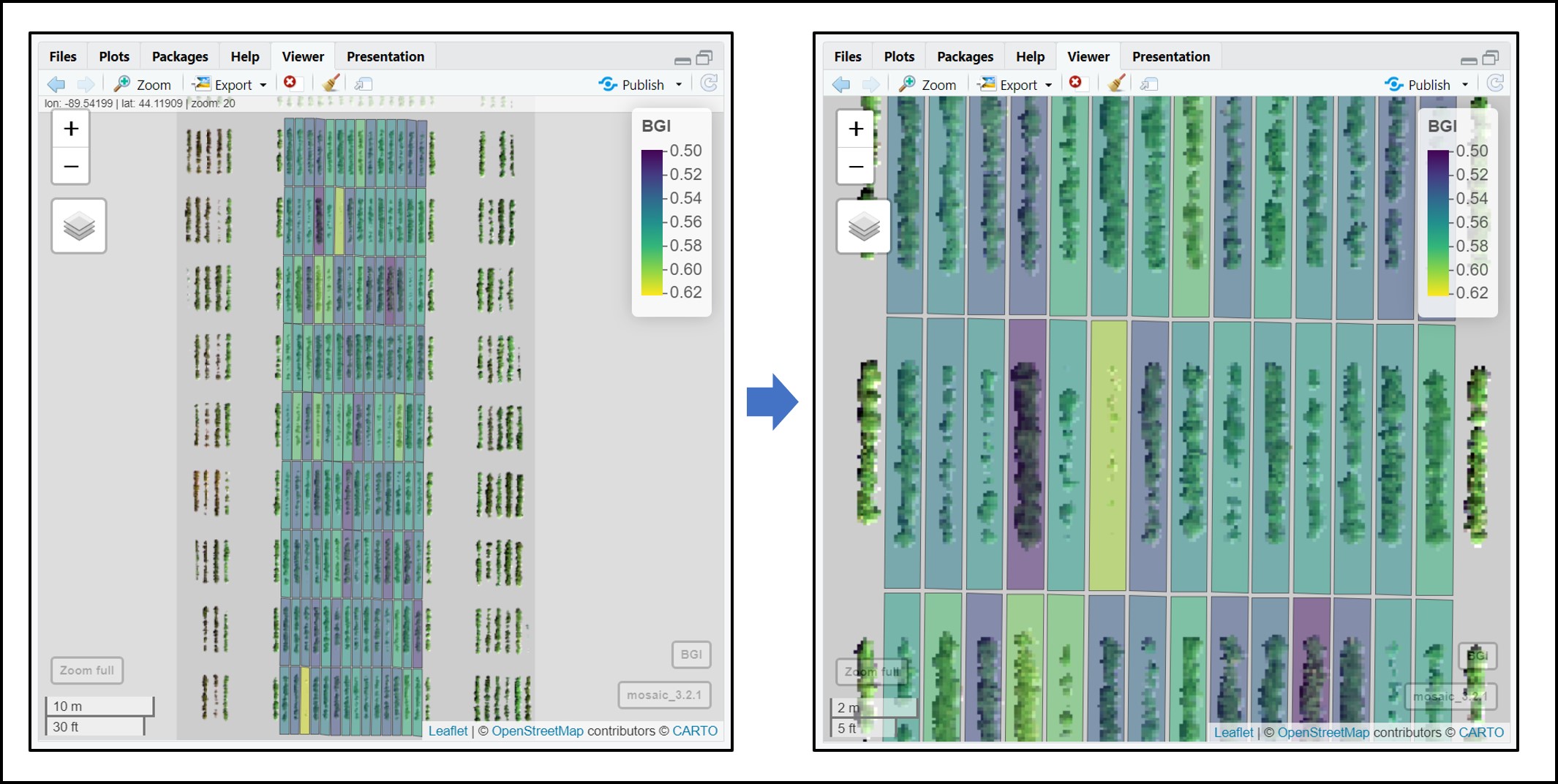
DataTotalThere are different ways to use the function
fieldViewto visualize and interpret extracted data.
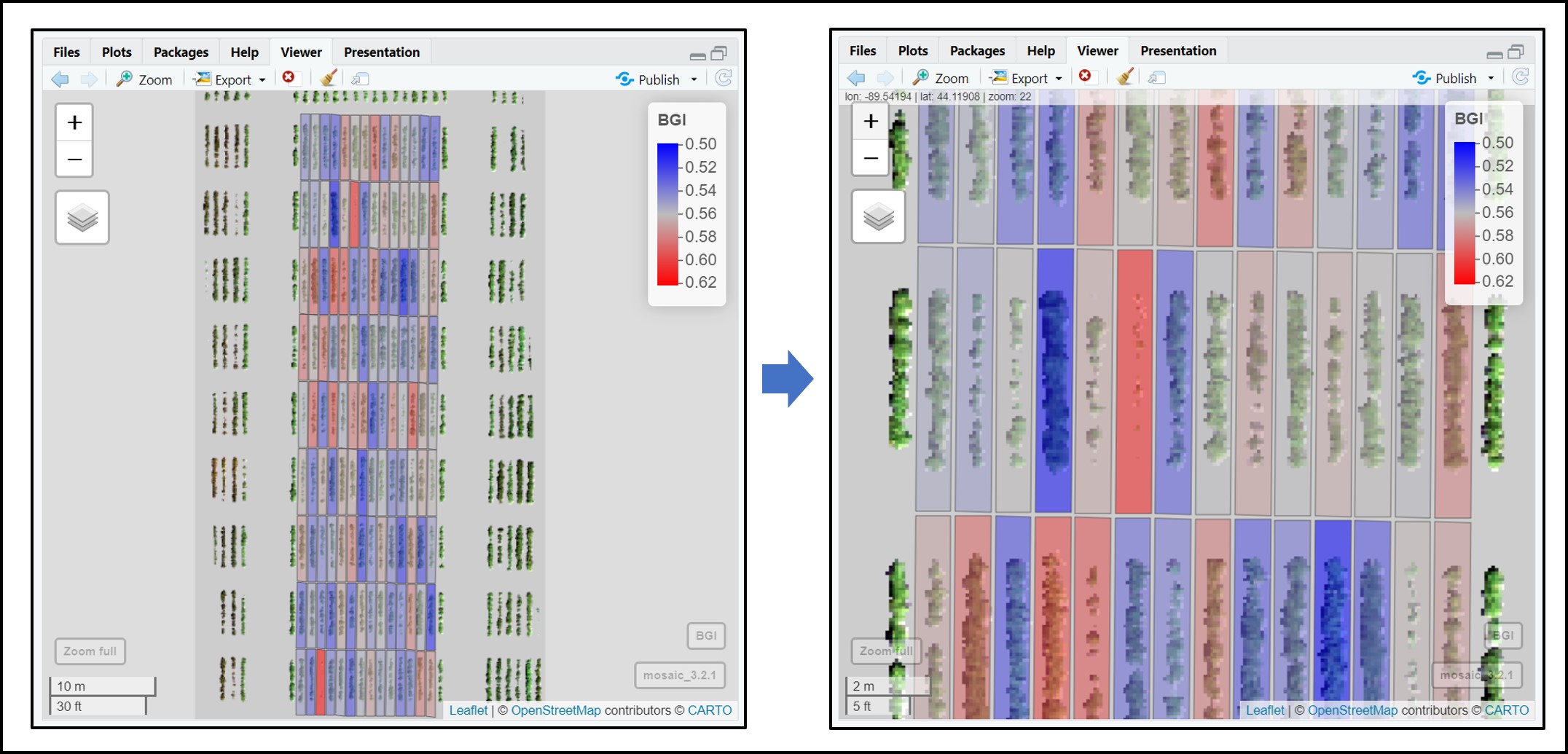
# Visualizing single layer vegetation index 'BGI':
fieldView(mosaic = Test.RemSoil$newMosaic,
fieldShape = DataTotal,
plotCol = "BGI",
type = 2,
alpha_grid = 0.6)# Applying different color schemes to paint plots in the fieldShape file. An example using extracted values for 'BGI':
fieldView(mosaic = Test.RemSoil$newMosaic,
fieldShape = DataTotal,
plotCol = "BGI",
col_grid = c("blue", "grey", "red"),
type = 2,
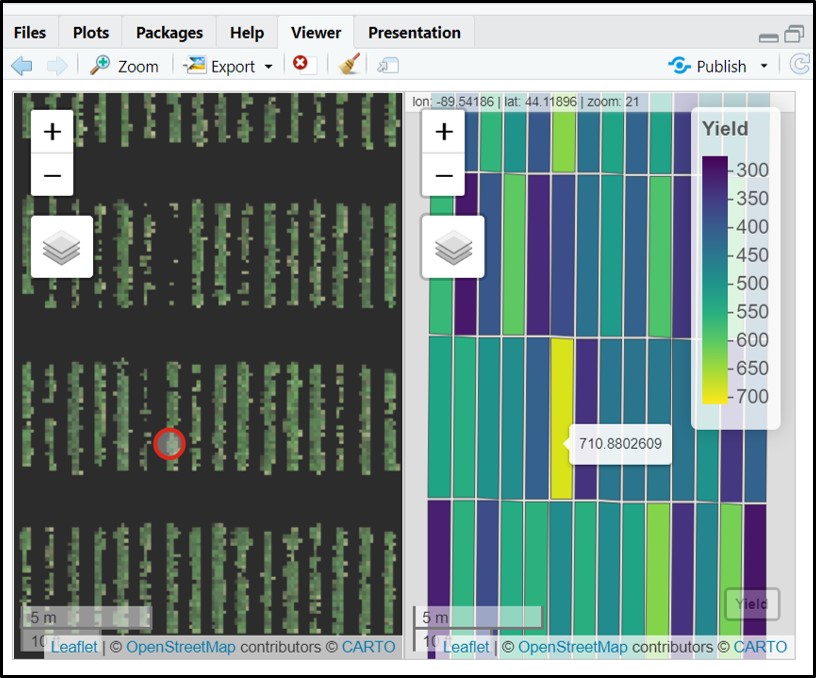
alpha_grid = 0.6)# Coloring fieldShape file using 'Yield' to highlight plots:
fieldView(mosaic = Test.RemSoil$newMosaic,
fieldShape = DataTotal,
plotCol = c("Yield"),
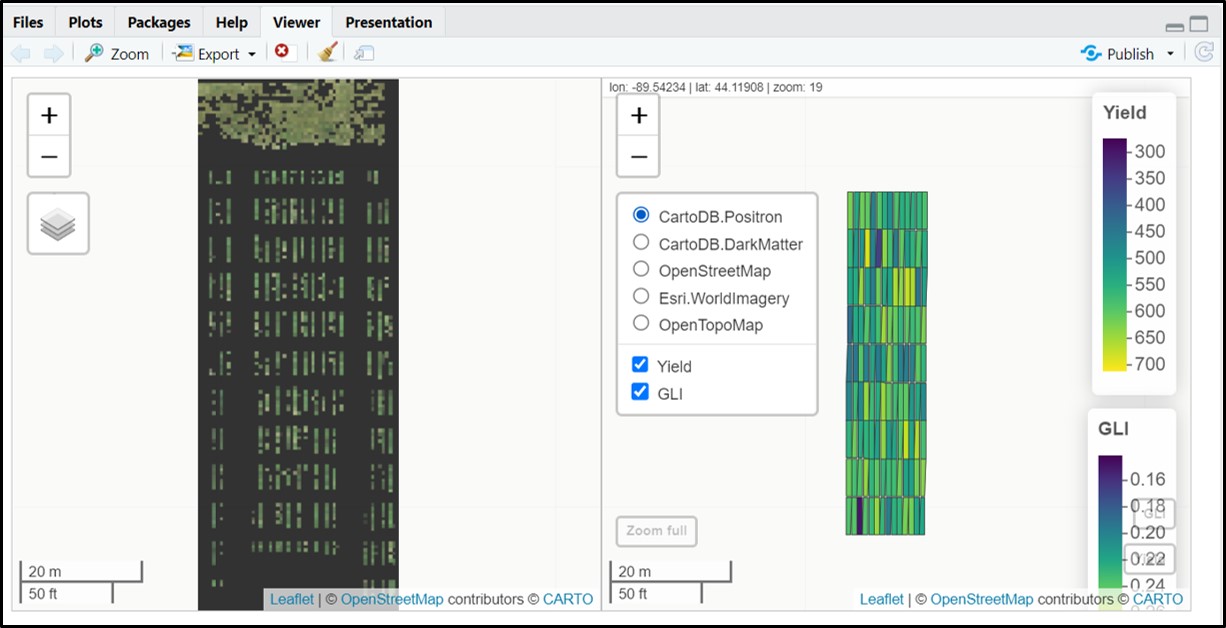
type = 1)# Coloring fieldShape file using more traits:
fieldView(mosaic = Test.RemSoil$newMosaic,
fieldShape = DataTotal,
plotCol = c("Yield","GLI"),
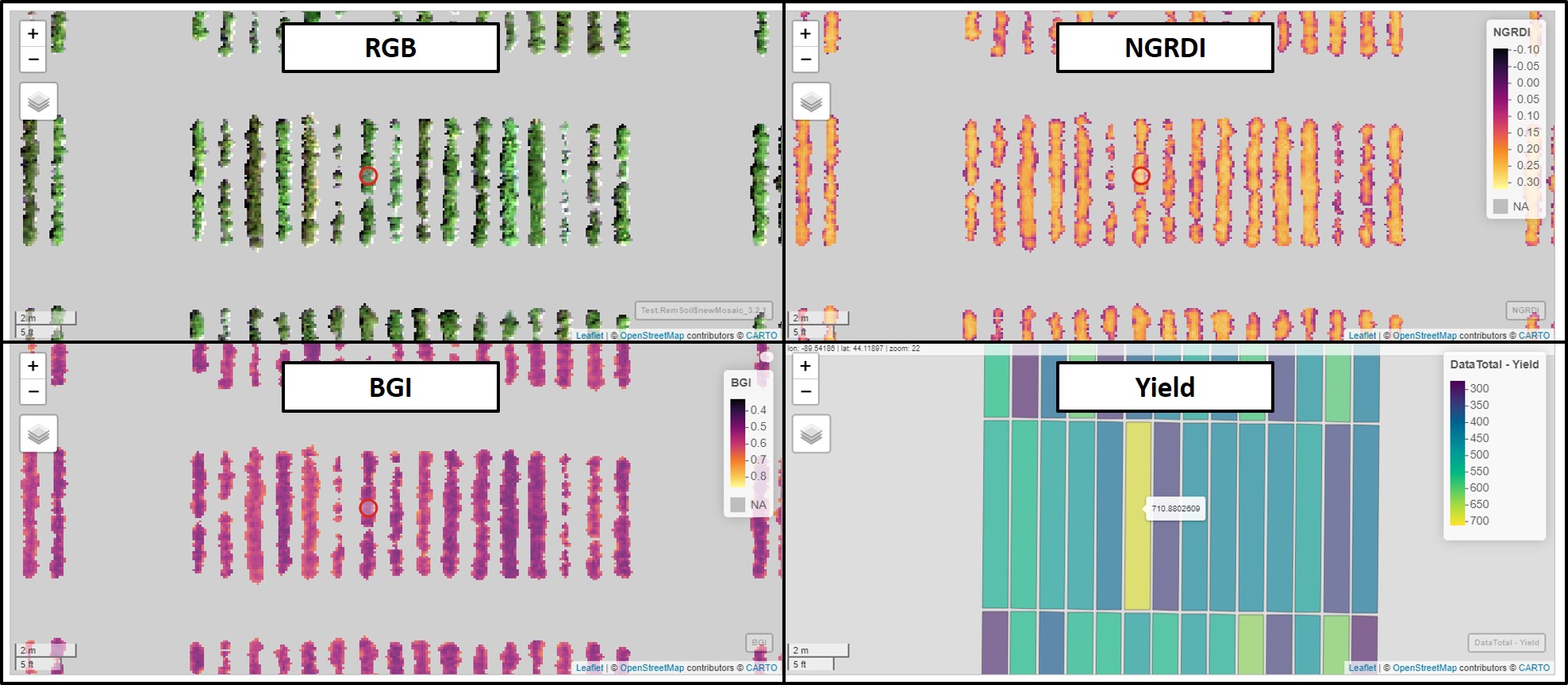
type = 1)# Syncronizing individual layers:
library(leafsync)
m1<-viewRGB(Test.RemSoil$newMosaic)
m2<-mapview(Test.RemSoil$newMosaic$NGRDI,layer.name="NGRDI")
m3<-mapview(Test.RemSoil$newMosaic$BGI,layer.name="BGI")
m4<-mapview(DataTotal,zcol="Yield")
sync(m1,m2,m3,m4)# Saving Data.csv (You can remove the 'geometry' info):
write.csv(as.data.frame(DataTotal)[,-dim(DataTotal)[2]],"DataTotal.csv",col.names = T, row.names = F)
# Saving grid fieldShape file:
st_write(DataTotal,"grid.shp")
# To read saved.shp object:
Saved_Grid= st_read("grid.shp")
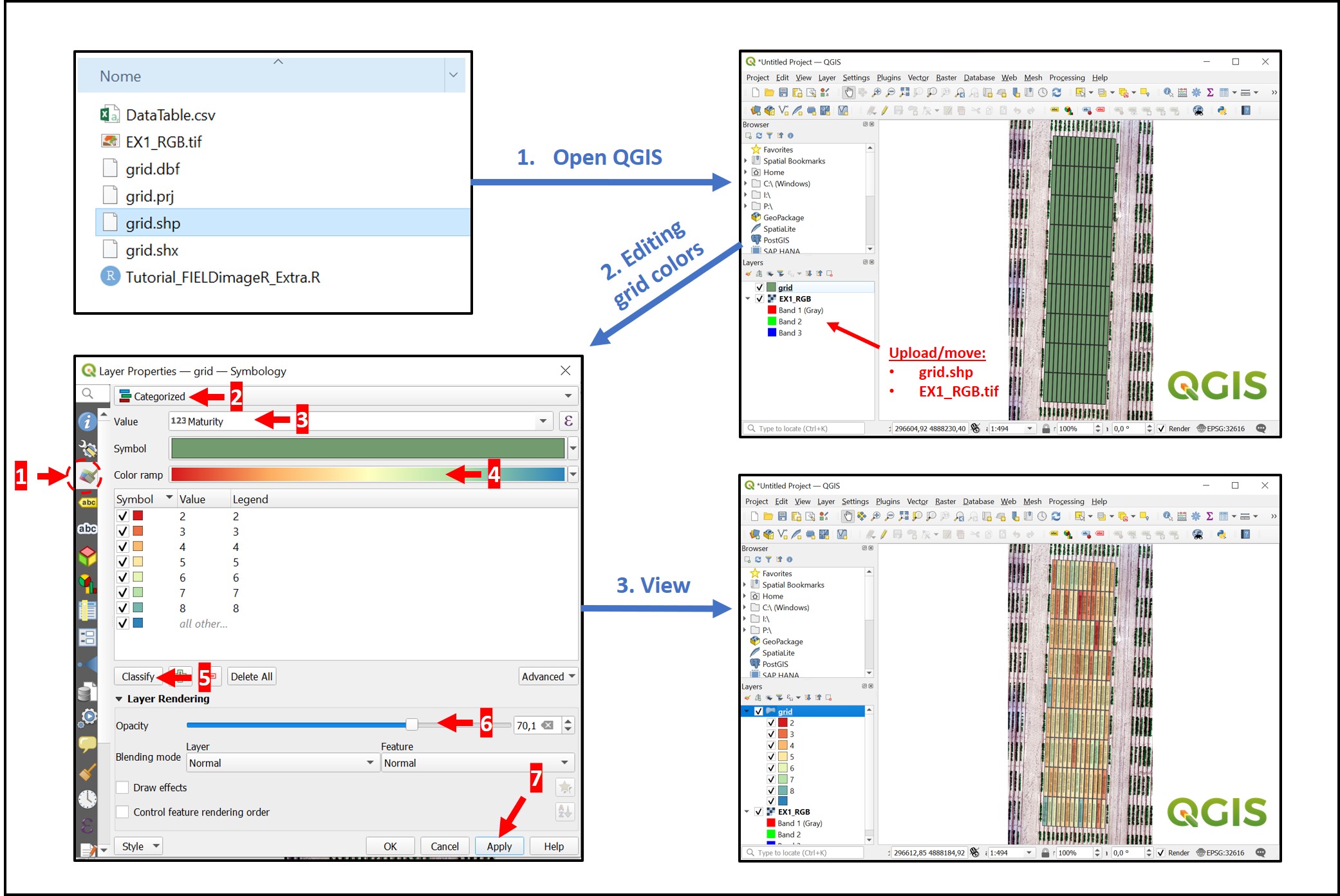
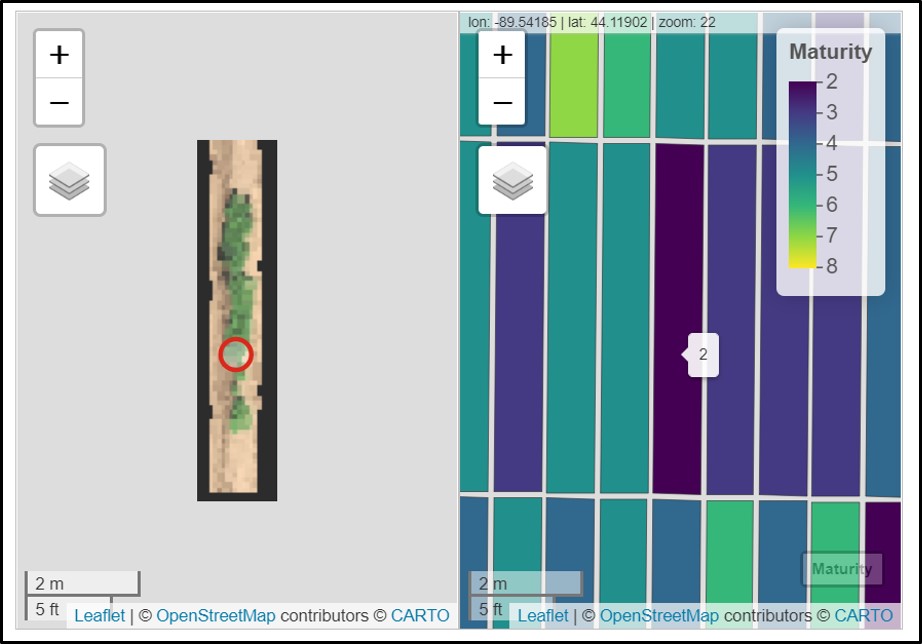
The saved "grid.shp" can be opened on others GIS softwares as QGIS. Users can easily upload the orthomosaic and grid.shp by holding the file from the folder and dragging it to a new project at QGIS. There are different visualization possibilities in QGIS. In the example below, each plot was identified according to the maturity score. However, any vegetation index calculated with FIELDimageR::fieldIndex can be visualized as well.
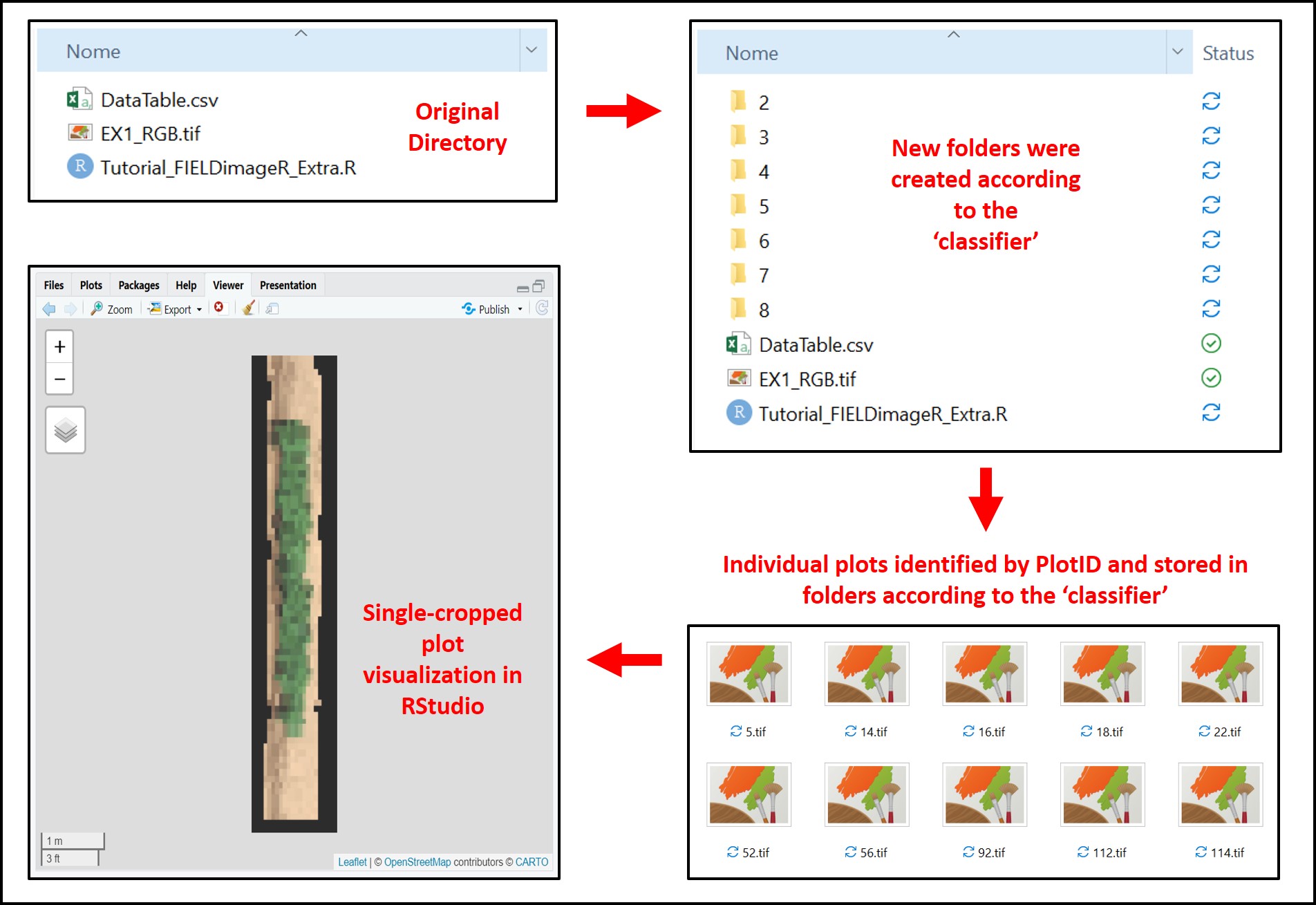
Many times when developing algorithms it is necessary to crop the mosaic using the plot fieldShape as a reference and sort/store cropped plot images on specific folders. For instance, the function
fieldCrop_gridallows cropping plots and identification by 'plotID'. The user also can save each plot according to a 'classifier' logic (Attention: a column in the 'fieldShape' with the desired classification must be informed). In the example below, each plot in the 'Test.Indices' mosaic is being cropped according to the 'editShape' grid file, identified by the information in the 'plot' column, and stored/saved in specific folders with different levels of Maturity the 'classifier'.
# Saving plot images format .jpg according to 'Maturity':
Field_plot_grids<- fieldCrop_grid(mosaic = rast(Test.Indices),
fieldShape = editShape,
classifier = "Maturity",
plotID = "Plot",
format = '.jpg',
output_dir = "./")
# Visualizing plot "#21"
fieldView(Field_plot_grids$'21')
# Saving plot images format .tif according to 'Maturity':
Field_plot_grids<- fieldCrop_grid(mosaic = rast(Test.Indices),
fieldShape = editShape,
classifier = "Maturity",
plotID = "Plot",
format = '.tif',
output_dir = "./")
# Reading .tif file and visualizing single plots with fieldView()
tif<-rast("./2/21.tif")
fieldView(mosaic = tif,
fieldShape = editShape,
plotCol ="Maturity")This discussion group provides an online source of information about the FIELDimageR package. Report a bug and ask a question at:
- https://groups.google.com/forum/#!forum/fieldimager
- https://community.opendronemap.org/t/about-the-fieldimager-category/4130
Help improve FIELDimageR.Extra pipeline. The easiest way to modify the package is by cloning the repository and making changes using R projects.
If you have questions, join the forum group at https://groups.google.com/forum/#!forum/fieldimager
Try to keep commits clean and simple
Submit a pull request with detailed changes and test results.
Let's work together and help more people (students, professors, farmers, etc) to have access to this knowledge. Thus, anyone anywhere can learn how to apply remote sensing in agriculture.
The R/FIELDimageR package as a whole is distributed under GPL-3 (GNU General Public License version 3).
-
Pawar P & Matias FI.* FIELDimageR.Extra. (2023) (Submitted)
-
Matias FI, Caraza-Harter MV, Endelman JB. FIELDimageR: An R package to analyze orthomosaic images from agricultural field trials. The Plant Phenome J. 2020; https://doi.org/10.1002/ppj2.20005
Popat Pawar
- GitHub
- E-mail: pspawar71@gmail.com
Filipe Matias
- GitHub
- E-mail: filipematias23@gmail.com