MimickNet
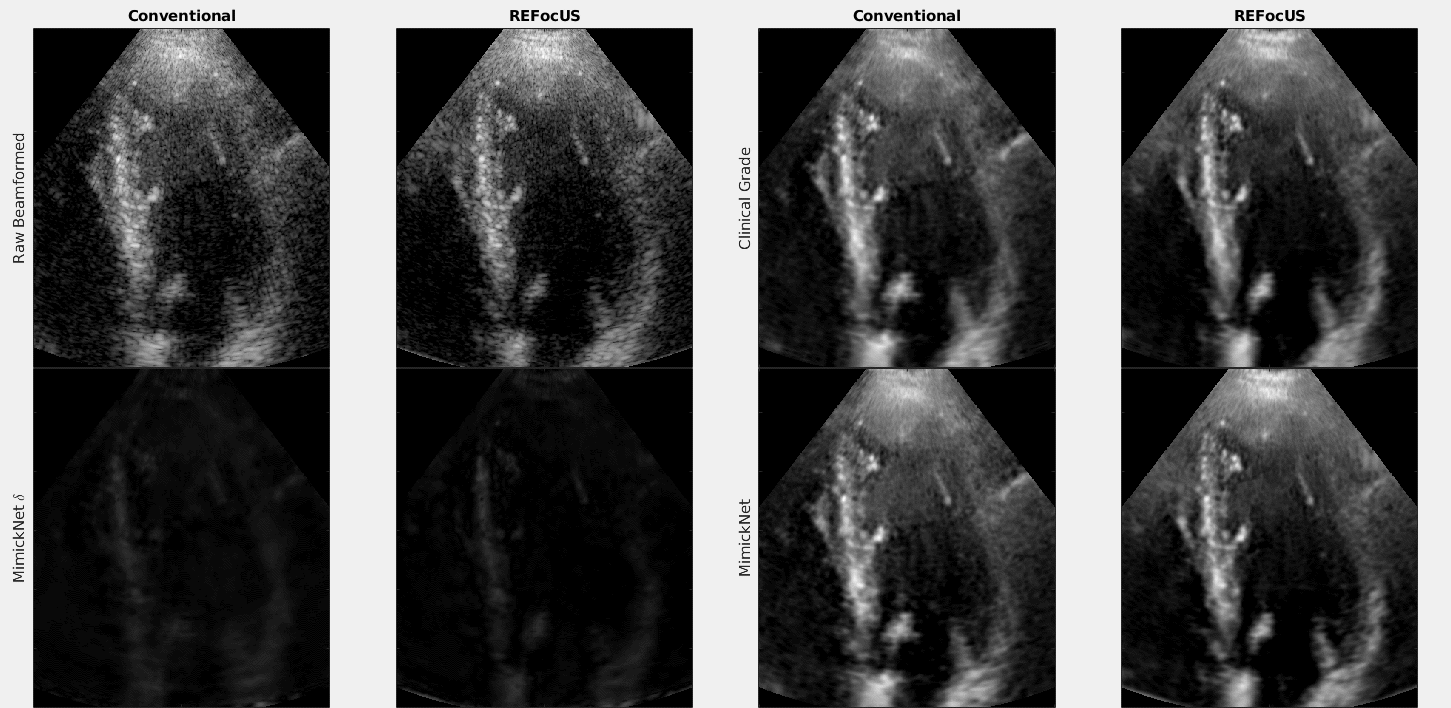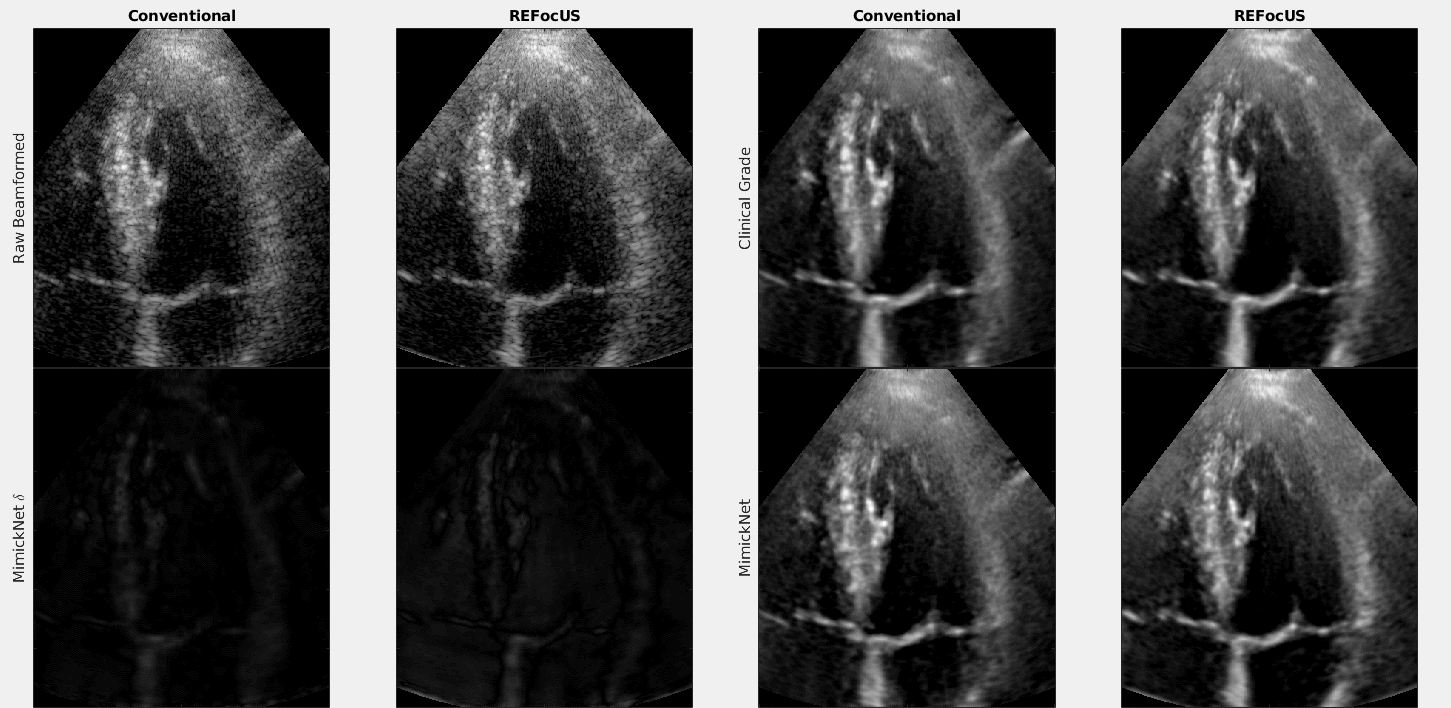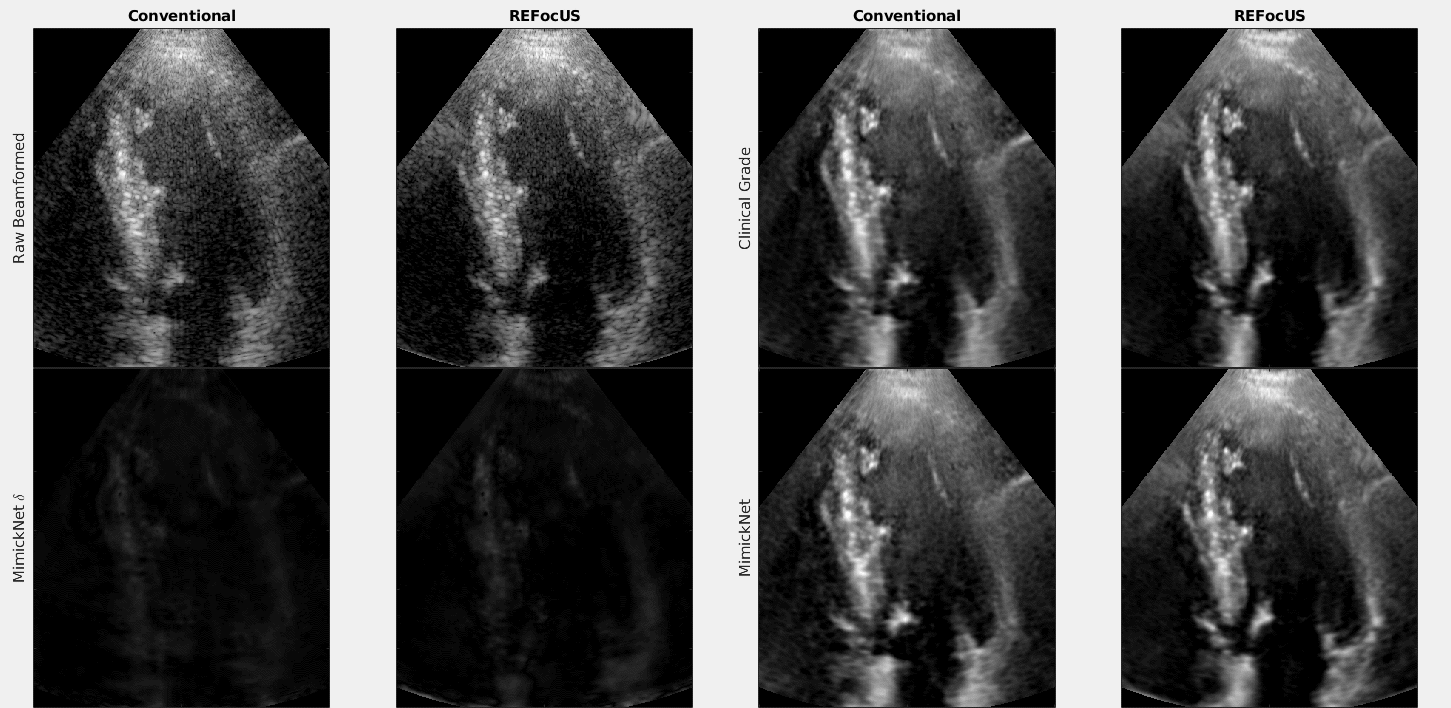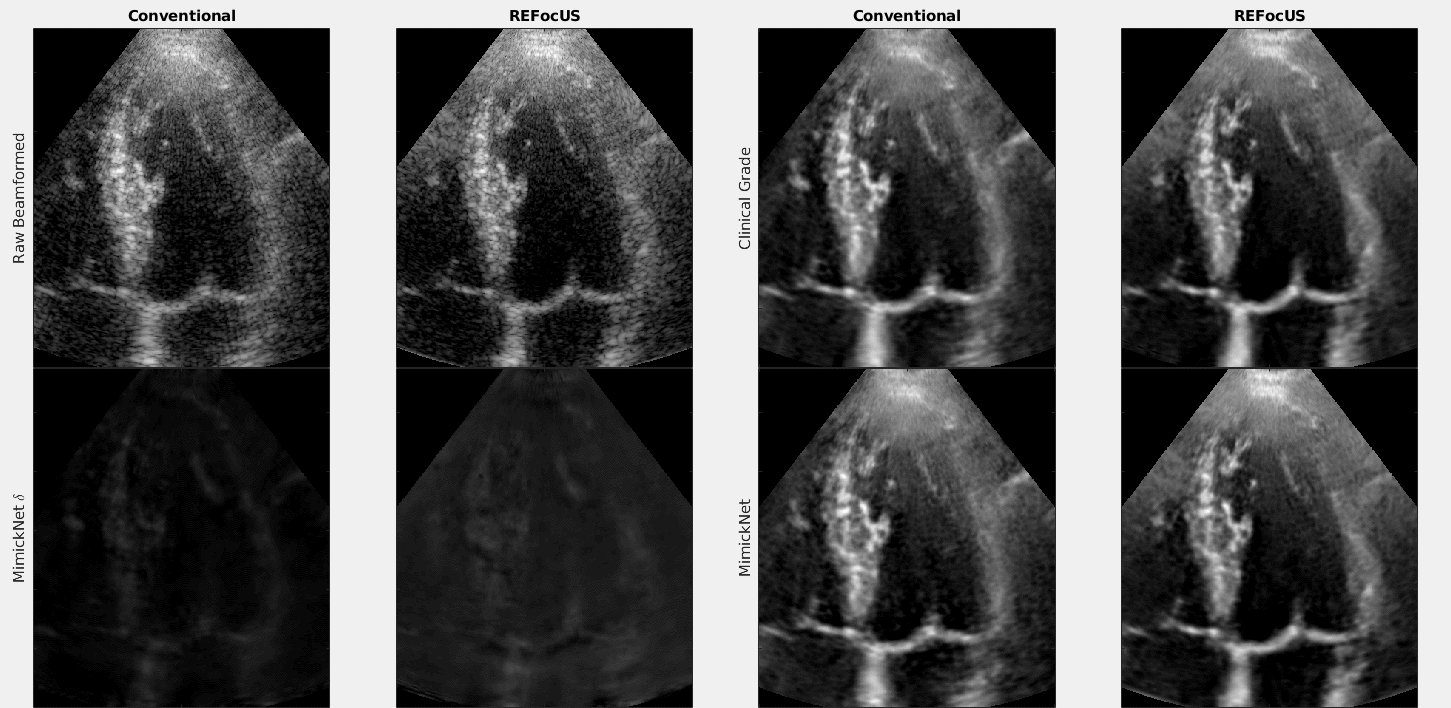
The above is a cineloop of cardiac data with conventional delay-and-sum beamforming and ReFoCUS beamforming. We apply clinical-grade post-processing, MimickNet post-processing, and show the unscaled difference between the two.
Quick Start
You can use the model immediately in this google colab notebook
The model from the paper is provided as mimicknet_legacy.h5. We also provide a luminance adjusted version with fewer weights with mimicknet_1568473738-210304.h5 and padded_mimicknet_1568473738-210304.h5. The padded prefix allows for padding to be apart of the model, so no 16 padding logic is required. However, the layer used is incompatible with matlab, thus the matlab version has no pre-padding.
The mimicknet_phantom_verasonics.h5 models were generated with the
A notebook and sample data is provided under examples for use in the following environments:
Dr. Mark Palmeri's liver and kidney images are provided as test examples (Thanks Mark!)
Metrics
| Model All | SSIM | PSNR |
|---|---|---|
| MimickNet (BlackBox) | 0.94 ± 0.014 | 31.95 ± 2.04 |
| GrayBox | 0.96 ± 0.012 | 32.86 ± 1.82 |
| MimickNet Phantom | 0.95 ± 0.007 | 33.50 ± 1.43 |
| MimickNet Mark | 0.96 ± 0.005 | 33.12 ± 0.92 |
Results above used non-public invivio + phantom data. Testing is on non-public invivo + phantom data. MimickNet Mark are results only on Mark's liver/cardiac data. MimickNet Phantom are results only on the public phantom data test split.
| Model Phantom | SSIM | PSNR |
|---|---|---|
| MimickNet Phantom | 0.90 ± 0.015 | 31.40 ± 2.76 |
| MimickNet Mark | 0.91 ± 0.005 | 31.43 ± 0.69 |
Results above used only public phantom data for training. MimickNet Phantom are results only on the public phantom data test data. MimickNet Mark are results only on Mark's liver/cardiac data.
Training the model from scratch
The following repo was designed to be run in Google Cloud and makes use of GCS for logging.
python3 -m trainer.blackbox_task
Different hyperparameters can be selected. Different tasks are shown in the root directory of the trainer. blackbox_task.py, and graybox_task.py are for running the blackbox and graybox tasks in the paper: MimickNet, Matching Clinical Post-Processing Under Realistic Black-Box Constraints. blackbox_paper_v1.py, is the exact code run for the first release.
Train the model in docker
To train this repo within a docker container, first clone the repo and run the following command in the root directory.
docker build -t mimicknet:latest .
Then you can run the docker container with the following command
docker run -it mimicknet:latest
If you have GPUs you can run docker with the --gpus flag.
Custom data
The current model uses the dataset loader found in ./trainer/utils/dataset.py. This dataset takes in two lists from different domains A and B. The dataset returns elements which are 4D tensors. Each dimension represents [batch, height, width, 1]. You can create your own dataset class, it simply needs to return a 4D tensor, and count the total number of elements. See here on how to use tensorflow datasets.
Contributing and Issues
Contributions are welcome in the form of pull requests. For issues, please provide a minimal example of code which results in an error.
License
Note that the model provided here is under a different license than the software code.
Model License
CC 4.0 Attribution-NonCommercial International
Software License
Copyright 2019 Ouwen Huang
Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
Thanks
This work was supported by the National Institute of Biomedical Imaging and Bioengineering under Grant R01- EB026574, and National Institutes of Health under Grant 5T32GM007171-44. The authors would like to thank Siemens Medical Inc. USA for in kind technical support.
Citing
@article{DBLP:journals/corr/abs-1908-05782,
author = {Ouwen Huang and
Will Long and
Nick Bottenus and
Gregg E. Trahey and
Sina Farsiu and
Mark L. Palmeri},
title = {MimickNet, Matching Clinical Post-Processing Under Realistic Black-Box
Constraints},
journal = {CoRR},
volume = {abs/1908.05782},
year = {2019},
url = {http://arxiv.org/abs/1908.05782},
archivePrefix = {arXiv},
eprint = {1908.05782},
timestamp = {Mon, 19 Aug 2019 13:21:03 +0200},
biburl = {https://dblp.org/rec/bib/journals/corr/abs-1908-05782},
bibsource = {dblp computer science bibliography, https://dblp.org}
}