pyGAM
Generalized Additive Models in Python.
Tutorial
pyGAM: Getting started with Generalized Additive Models in Python
Installation
pip install pygam
scikit-sparse
To speed up optimization on large models with constraints, it helps to have scikit-sparse installed because it contains a slightly faster, sparse version of Cholesky factorization. The import from scikit-sparse references nose, so you'll need that too.
The easiest way is to use Conda:
conda install -c conda-forge scikit-sparse nose
Contributing
Contributions are most welcome!
You can help pyGAM in many ways including:
- Trying it out and reporting bugs or what was difficult.
- Helping improve the documentation.
- Writing new distributions, and link functions.
- If you need some ideas, please take a look at the issues.
To start:
- fork the project and cut a new branch
- Now install the testing dependencies
conda install pytest numpy pandas scipy pytest-cov cython scikit-sparse nose
pip install --upgrade pip
pip install -r requirements.txt
It helps to add a sym-link of the forked project to your python path. To do this, you should install flit:
pip install flit- Then from main project folder (ie
.../pyGAM) do:flit install -s
Make some changes and write a test...
- Test your contribution (eg from the
.../pyGAM):py.test -s - When you are happy with your changes, make a pull request into the
masterbranch of the main project.
About
Generalized Additive Models (GAMs) are smooth semi-parametric models of the form:
where X.T = [X_1, X_2, ..., X_p] are independent variables, y is the dependent variable, and g() is the link function that relates our predictor variables to the expected value of the dependent variable.
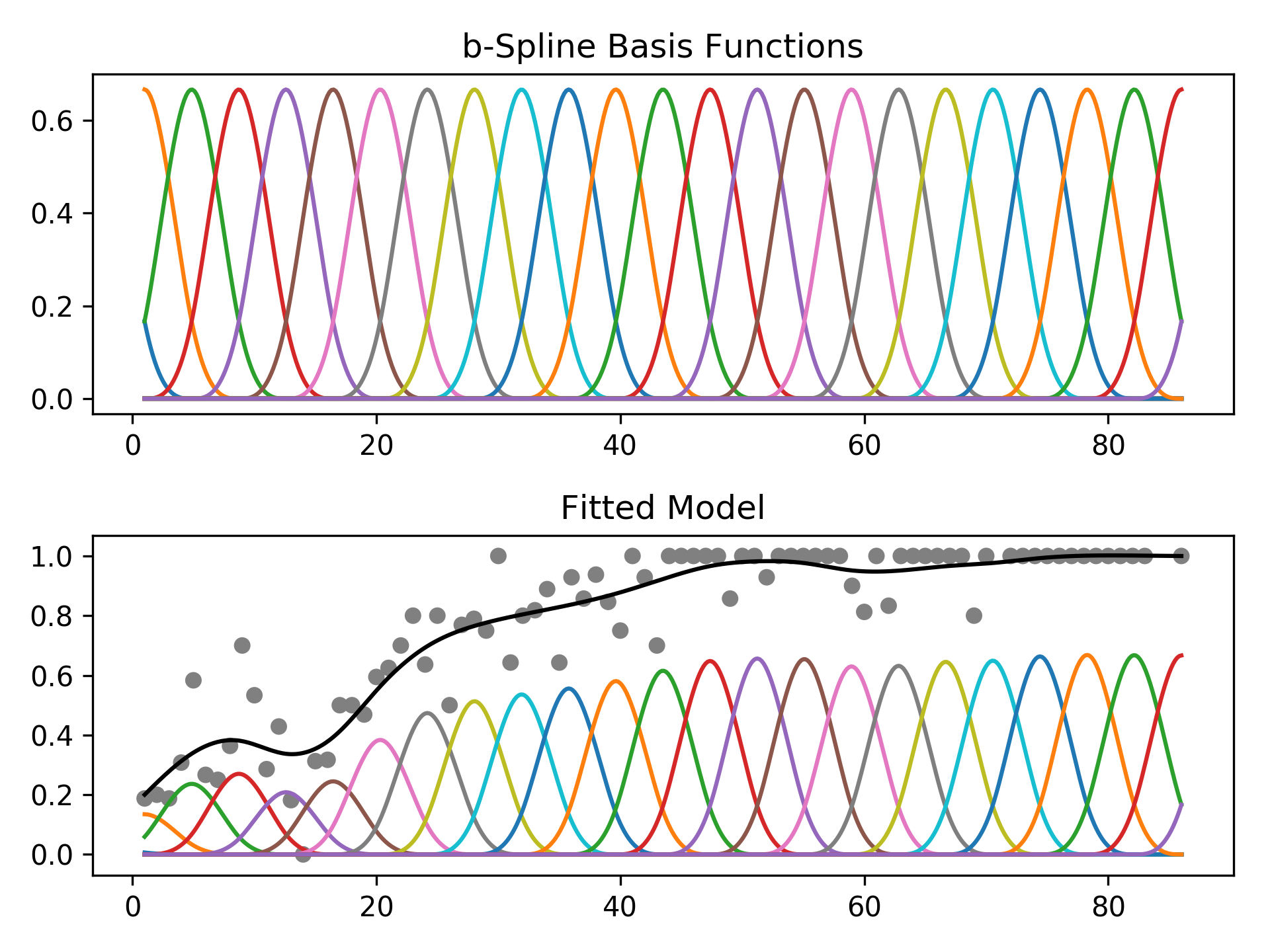
The feature functions f_i() are built using penalized B splines, which allow us to automatically model non-linear relationships without having to manually try out many different transformations on each variable.
GAMs extend generalized linear models by allowing non-linear functions of features while maintaining additivity. Since the model is additive, it is easy to examine the effect of each X_i on Y individually while holding all other predictors constant.
The result is a very flexible model, where it is easy to incorporate prior knowledge and control overfitting.
Regression
For regression problems, we can use a linear GAM which models:
from pygam import LinearGAM, s, f
from pygam.datasets import wage
X, y = wage(return_X_y=True)
gam = LinearGAM(s(0) + s(1) + f(2)).gridsearch(X, y)
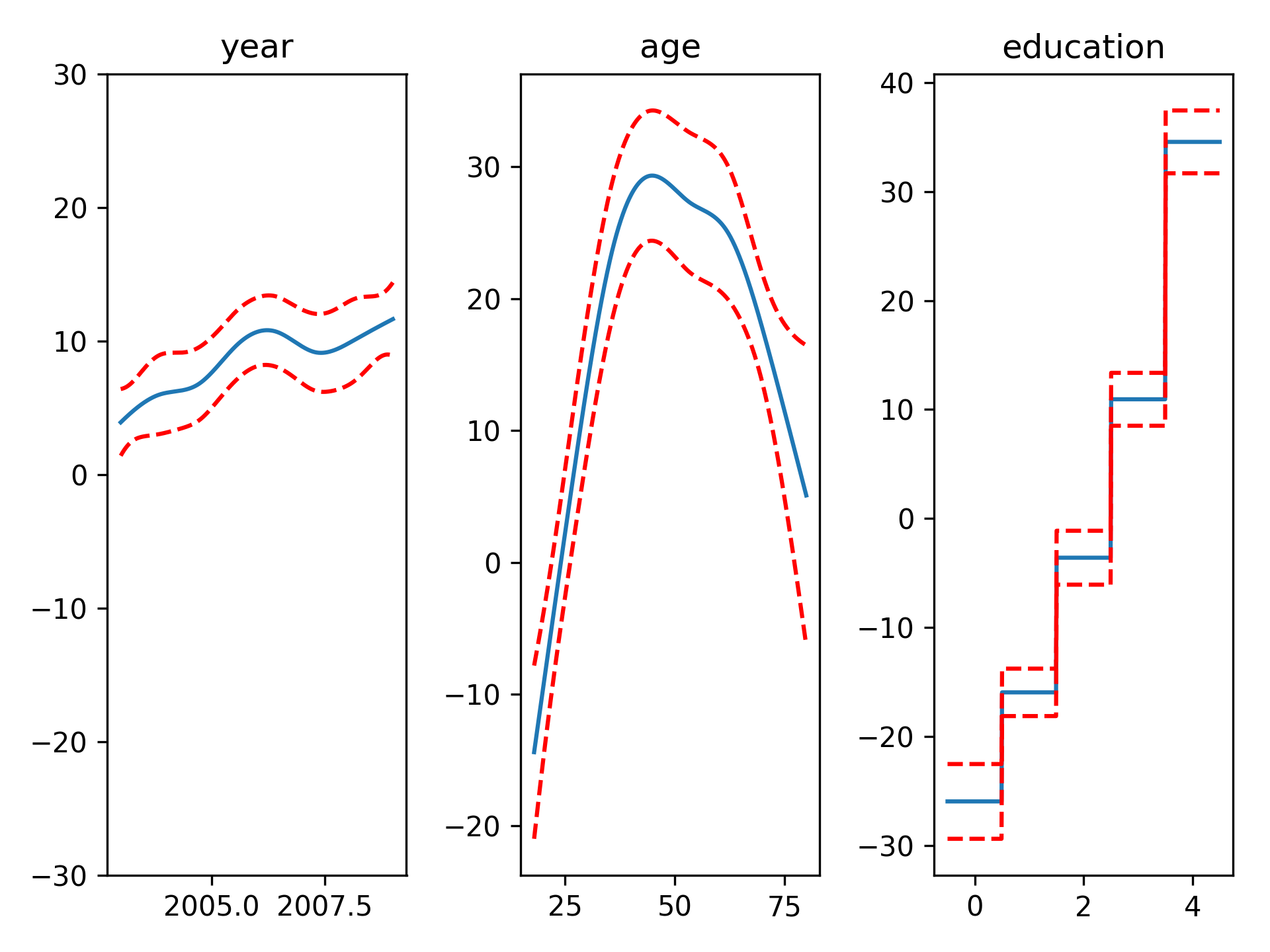
fig, axs = plt.subplots(1, 3)
titles = ['year', 'age', 'education']
for i, ax in enumerate(axs):
XX = gam.generate_X_grid(term=i)
pdep, confi = gam.partial_dependence(term=i, width=.95)
ax.plot(XX[:, i], pdep)
ax.plot(XX[:, i], confi, c='r', ls='--')
ax.set_title(titles[i])Even though we allowed n_splines=20 per numerical feature, our smoothing penalty reduces us to just 19 effective degrees of freedom:
gam.summary()
LinearGAM
=============================================== ==========================================================
Distribution: NormalDist Effective DoF: 19.2602
Link Function: IdentityLink Log Likelihood: -24116.7451
Number of Samples: 3000 AIC: 48274.0107
AICc: 48274.2999
GCV: 1250.3656
Scale: 1235.9245
Pseudo R-Squared: 0.2945
==========================================================================================================
Feature Function Lambda Rank EDoF P > x Sig. Code
================================= ==================== ============ ============ ============ ============
s(0) [15.8489] 20 6.9 5.52e-03 **
s(1) [15.8489] 20 8.5 1.11e-16 ***
f(2) [15.8489] 5 3.8 1.11e-16 ***
intercept 0 1 0.0 1.11e-16 ***
==========================================================================================================
Significance codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
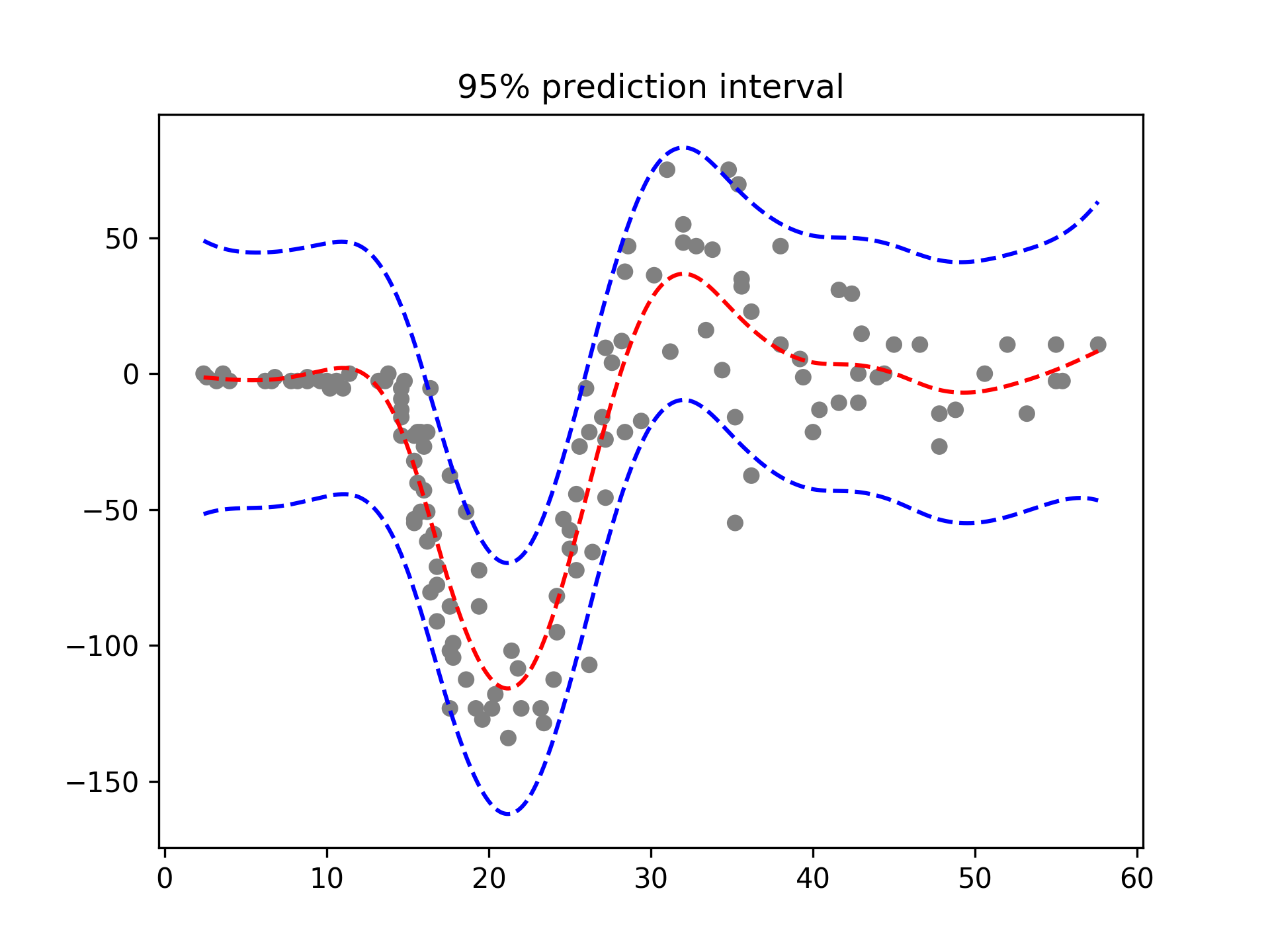
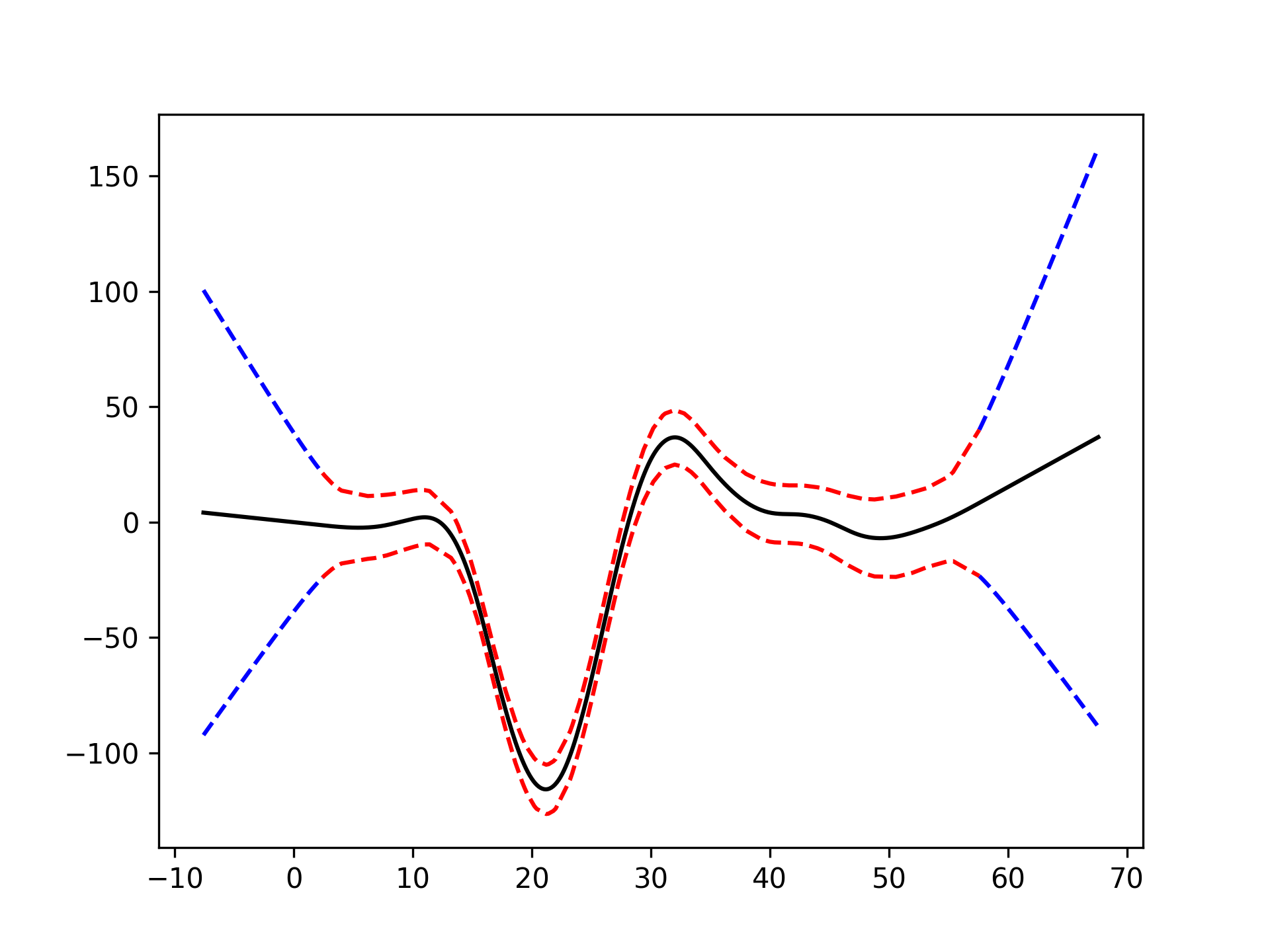
With LinearGAMs, we can also check the prediction intervals:
from pygam import LinearGAM
from pygam.datasets import mcycle
X, y = mcycle(return_X_y=True)
gam = LinearGAM().gridsearch(X, y)
XX = gam.generate_X_grid(term=0)
plt.plot(XX, gam.predict(XX), 'r--')
plt.plot(XX, gam.prediction_intervals(XX, width=.95), color='b', ls='--')
plt.scatter(X, y, facecolor='gray', edgecolors='none')
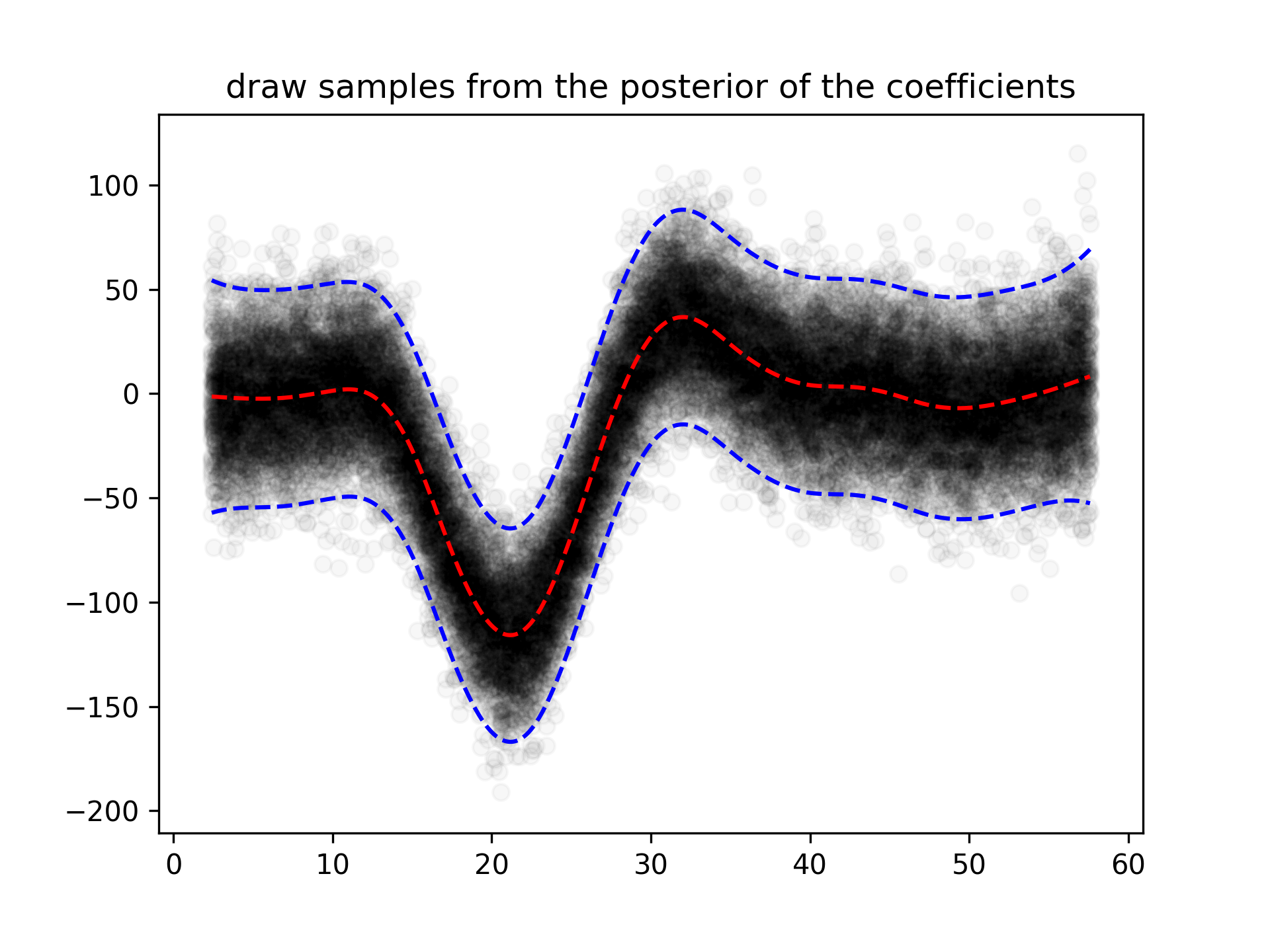
plt.title('95% prediction interval')And simulate from the posterior:
# continuing last example with the mcycle dataset
for response in gam.sample(X, y, quantity='y', n_draws=50, sample_at_X=XX):
plt.scatter(XX, response, alpha=.03, color='k')
plt.plot(XX, gam.predict(XX), 'r--')
plt.plot(XX, gam.prediction_intervals(XX, width=.95), color='b', ls='--')
plt.title('draw samples from the posterior of the coefficients')Classification
For binary classification problems, we can use a logistic GAM which models:
from pygam import LogisticGAM, s, f
from pygam.datasets import default
X, y = default(return_X_y=True)
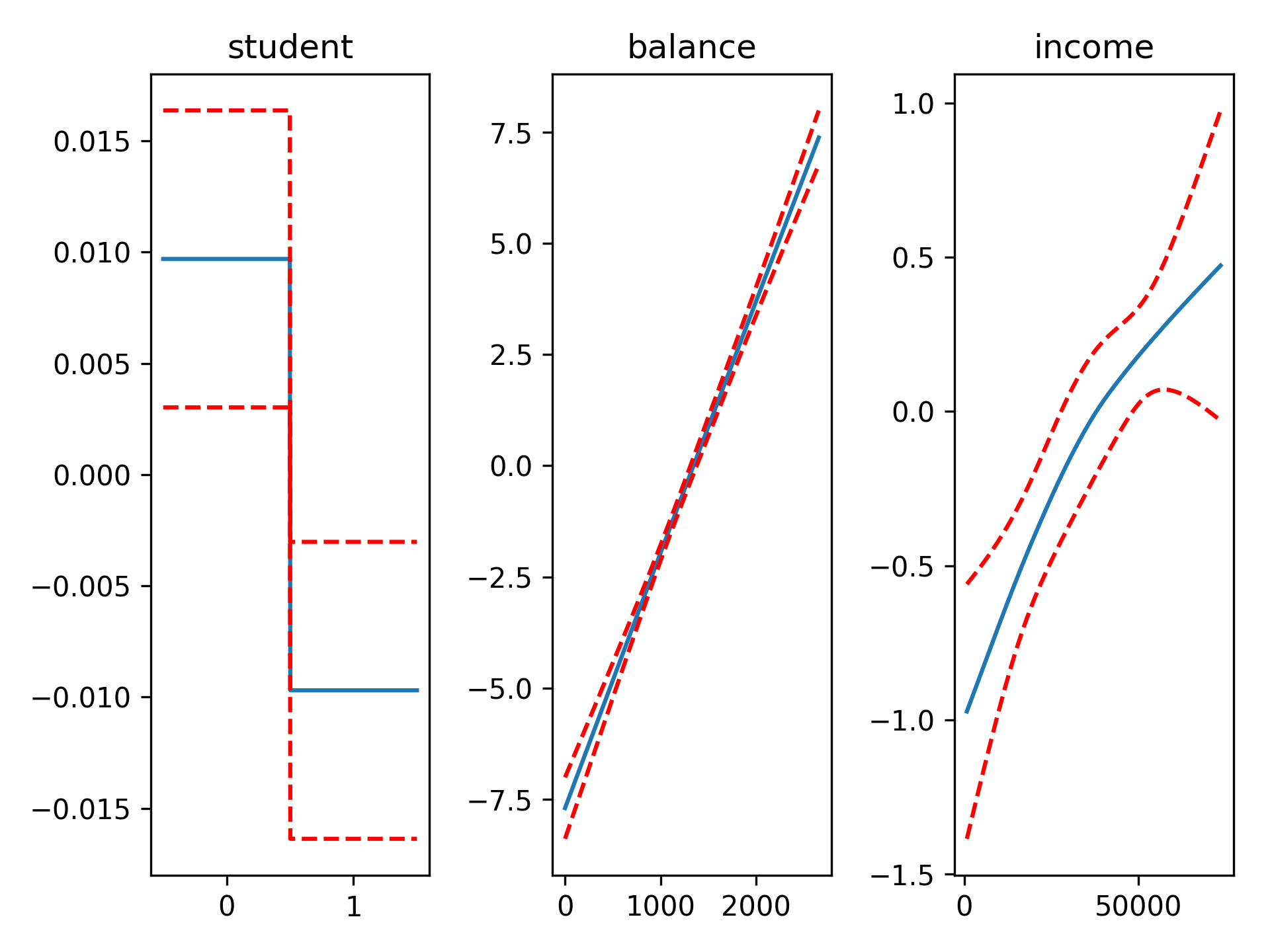
gam = LogisticGAM(f(0) + s(1) + s(2)).gridsearch(X, y)
fig, axs = plt.subplots(1, 3)
titles = ['student', 'balance', 'income']
for i, ax in enumerate(axs):
XX = gam.generate_X_grid(term=i)
pdep, confi = gam.partial_dependence(term=i, width=.95)
ax.plot(XX[:, i], pdep)
ax.plot(XX[:, i], confi, c='r', ls='--')
ax.set_title(titles[i])
# and check the accuracy
gam.accuracy(X, y)Since the scale of the Binomial distribution is known, our gridsearch minimizes the Un-Biased Risk Estimator (UBRE) objective:
gam.summary()
LogisticGAM
=============================================== ==========================================================
Distribution: BinomialDist Effective DoF: 3.8047
Link Function: LogitLink Log Likelihood: -788.877
Number of Samples: 10000 AIC: 1585.3634
AICc: 1585.369
UBRE: 2.1588
Scale: 1.0
Pseudo R-Squared: 0.4598
==========================================================================================================
Feature Function Lambda Rank EDoF P > x Sig. Code
================================= ==================== ============ ============ ============ ============
f(0) [1000.] 2 1.7 4.61e-03 **
s(1) [1000.] 20 1.2 0.00e+00 ***
s(2) [1000.] 20 0.8 3.29e-02 *
intercept 0 1 0.0 0.00e+00 ***
==========================================================================================================
Significance codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Poisson and Histogram Smoothing
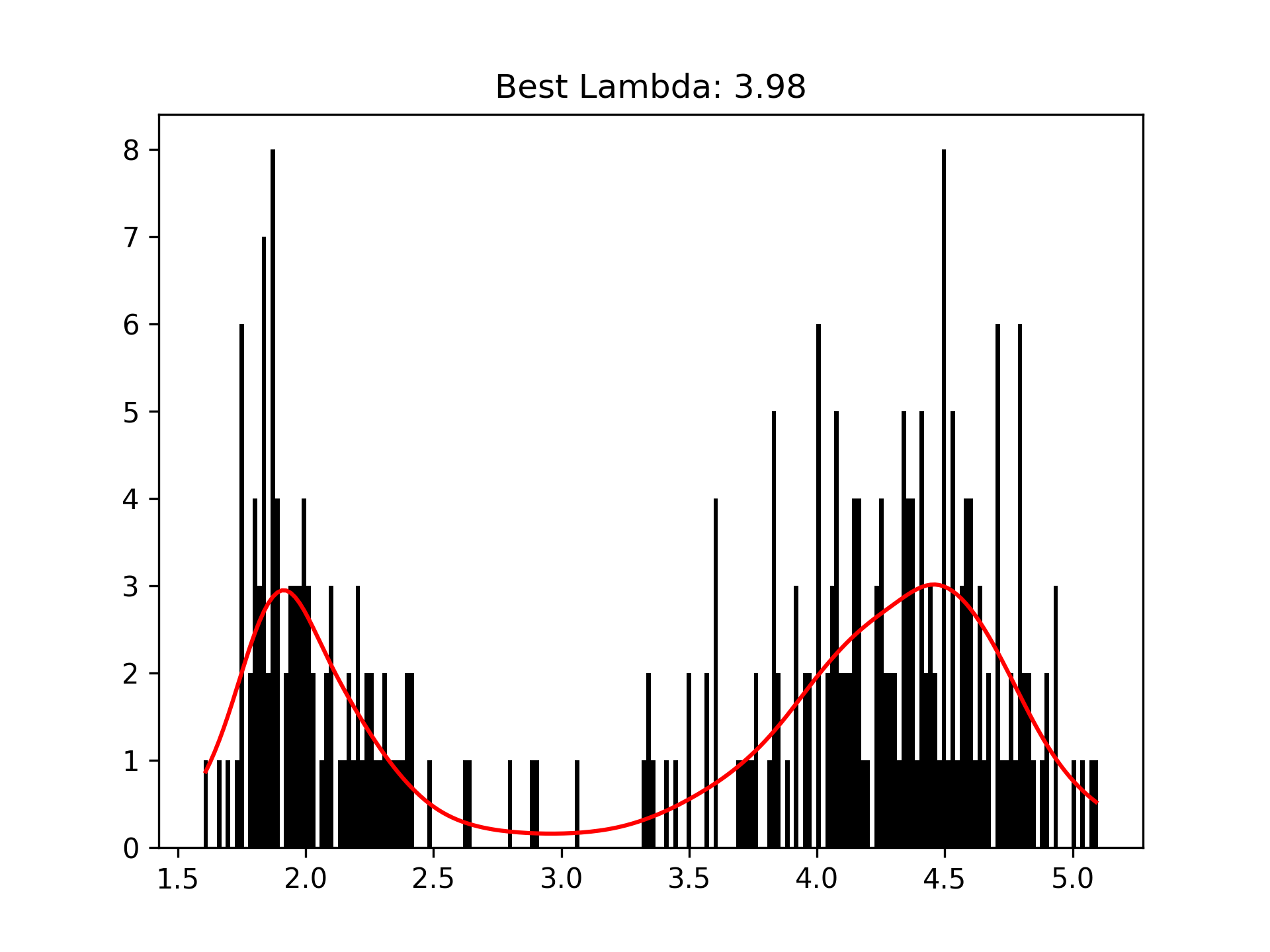
We can intuitively perform histogram smoothing by modeling the counts in each bin as being distributed Poisson via PoissonGAM.
from pygam import PoissonGAM
from pygam.datasets import faithful
X, y = faithful(return_X_y=True)
gam = PoissonGAM().gridsearch(X, y)
plt.hist(faithful(return_X_y=False)['eruptions'], bins=200, color='k');
plt.plot(X, gam.predict(X), color='r')
plt.title('Best Lambda: {0:.2f}'.format(gam.lam[0][0]));Terms and Interactions
pyGAM can also fit interactions using tensor products via te()
from pygam import LinearGAM, s, te
from pygam.datasets import chicago
X, y = chicago(return_X_y=True)
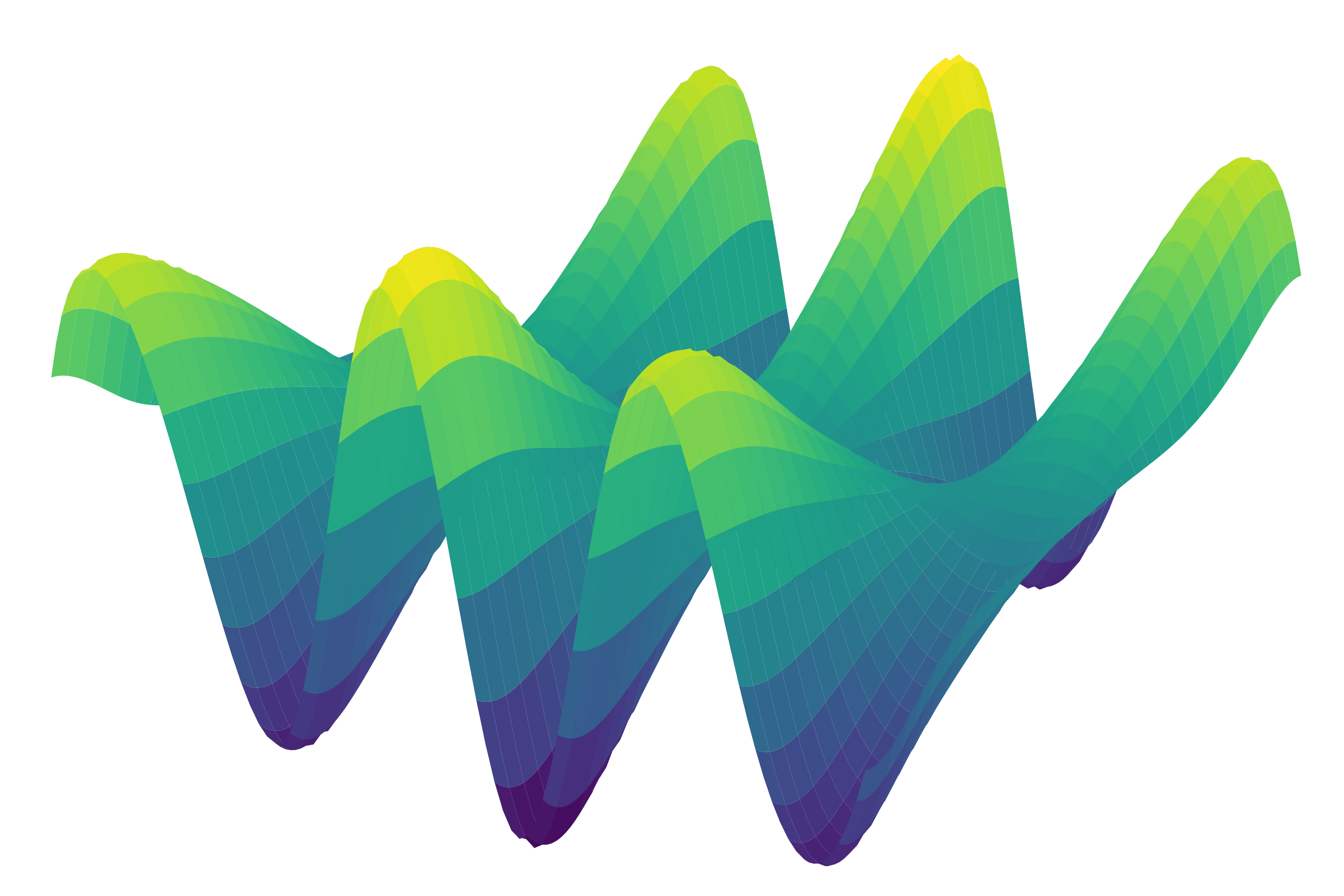
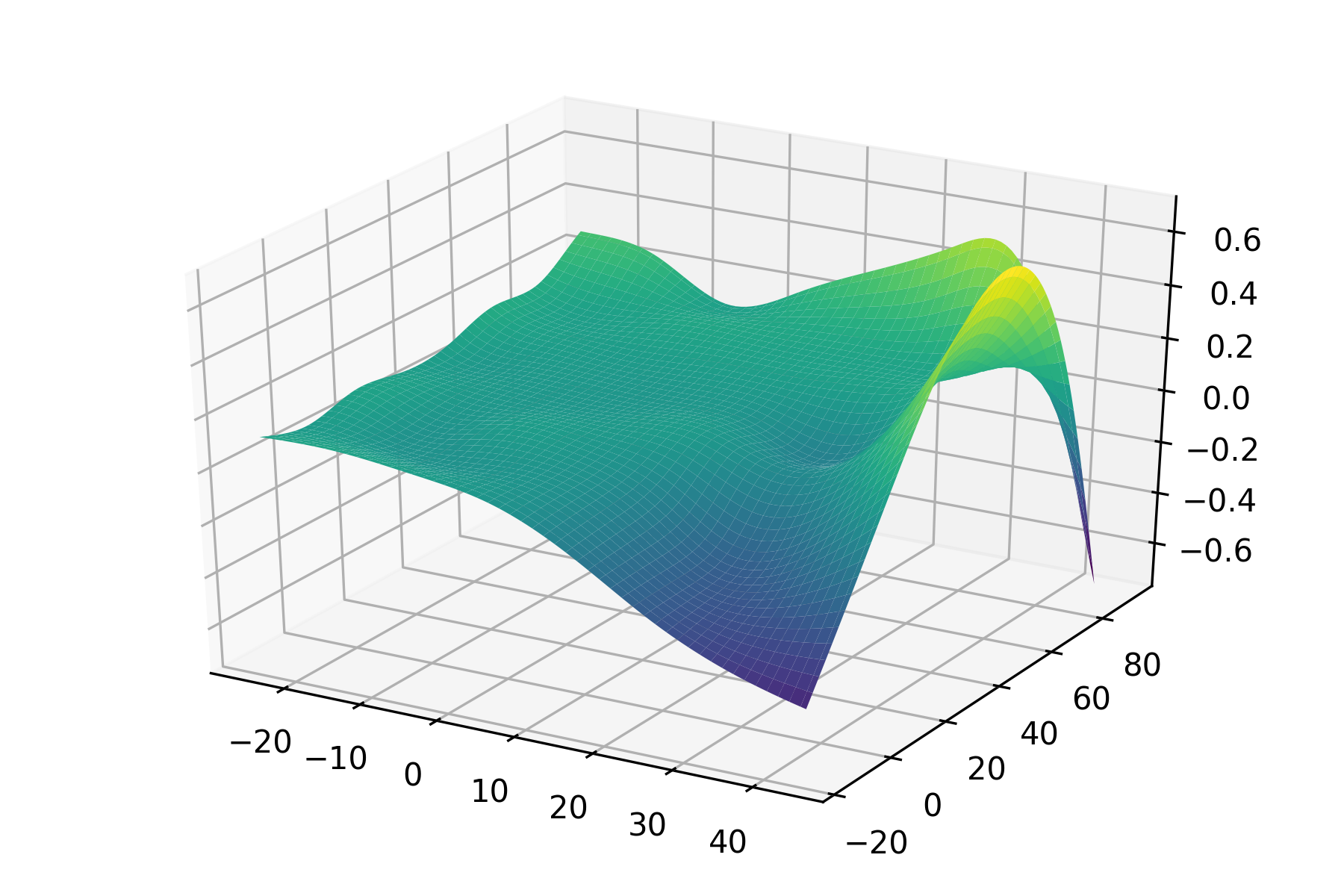
gam = PoissonGAM(s(0, n_splines=200) + te(3, 1) + s(2)).fit(X, y)and plot a 3D surface:
XX = gam.generate_X_grid(term=0, meshgrid=True)
Z = gam.partial_dependence(term=0, X=XX, meshgrid=True)
from mpl_toolkits import mplot3d
ax = plt.axes(projection='3d')
ax.plot_surface(XX[0], XX[1], Z, cmap='viridis')For simple interactions it is sometimes useful to add a by-variable to a term
from pygam import LinearGAM, s
from pygam.datasets import toy_interaction
X, y = toy_interaction(return_X_y=True)
gam = LinearGAM(s(0, by=1)).fit(X, y)
gam.summary()Available Terms
l()linear termss()spline termsf()factor termste()tensor productsintercept
Custom Models
It's also easy to build custom models, by using the base GAM class and specifying the distribution and the link function.
from pygam import GAM
from pygam.datasets import trees
X, y = trees(return_X_y=True)
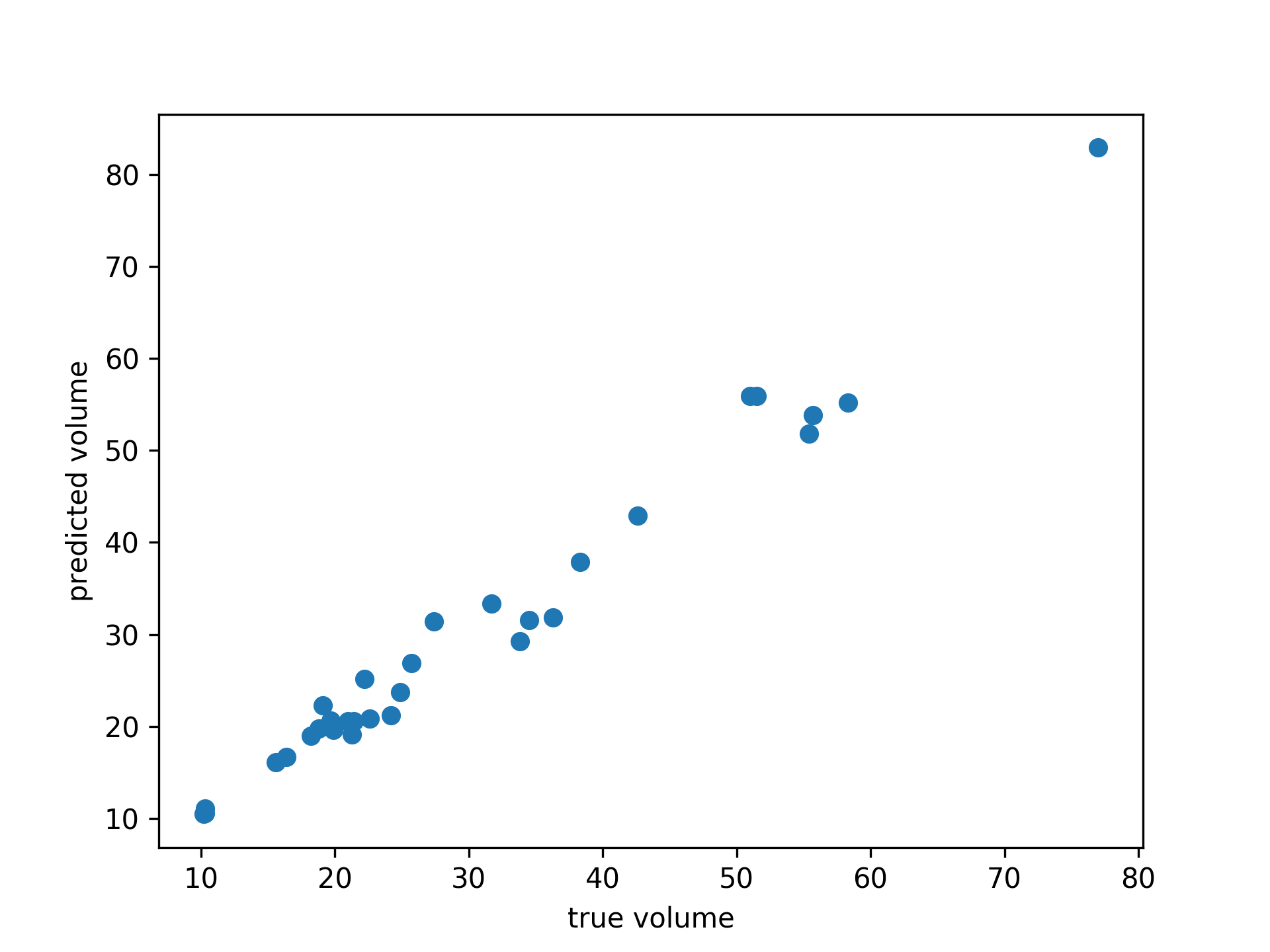
gam = GAM(distribution='gamma', link='log')
gam.gridsearch(X, y)
plt.scatter(y, gam.predict(X))
plt.xlabel('true volume')
plt.ylabel('predicted volume')We can check the quality of the fit by looking at the Pseudo R-Squared:
gam.summary()
GAM
=============================================== ==========================================================
Distribution: GammaDist Effective DoF: 25.3616
Link Function: LogLink Log Likelihood: -26.1673
Number of Samples: 31 AIC: 105.0579
AICc: 501.5549
GCV: 0.0088
Scale: 0.001
Pseudo R-Squared: 0.9993
==========================================================================================================
Feature Function Lambda Rank EDoF P > x Sig. Code
================================= ==================== ============ ============ ============ ============
s(0) [0.001] 20 2.04e-08 ***
s(1) [0.001] 20 7.36e-06 ***
intercept 0 1 4.39e-13 ***
==========================================================================================================
Significance codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Penalties / Constraints
With GAMs we can encode prior knowledge and control overfitting by using penalties and constraints.
Available penalties:
- second derivative smoothing (default on numerical features)
- L2 smoothing (default on categorical features)
Availabe constraints:
- monotonic increasing/decreasing smoothing
- convex/concave smoothing
- periodic smoothing [soon...]
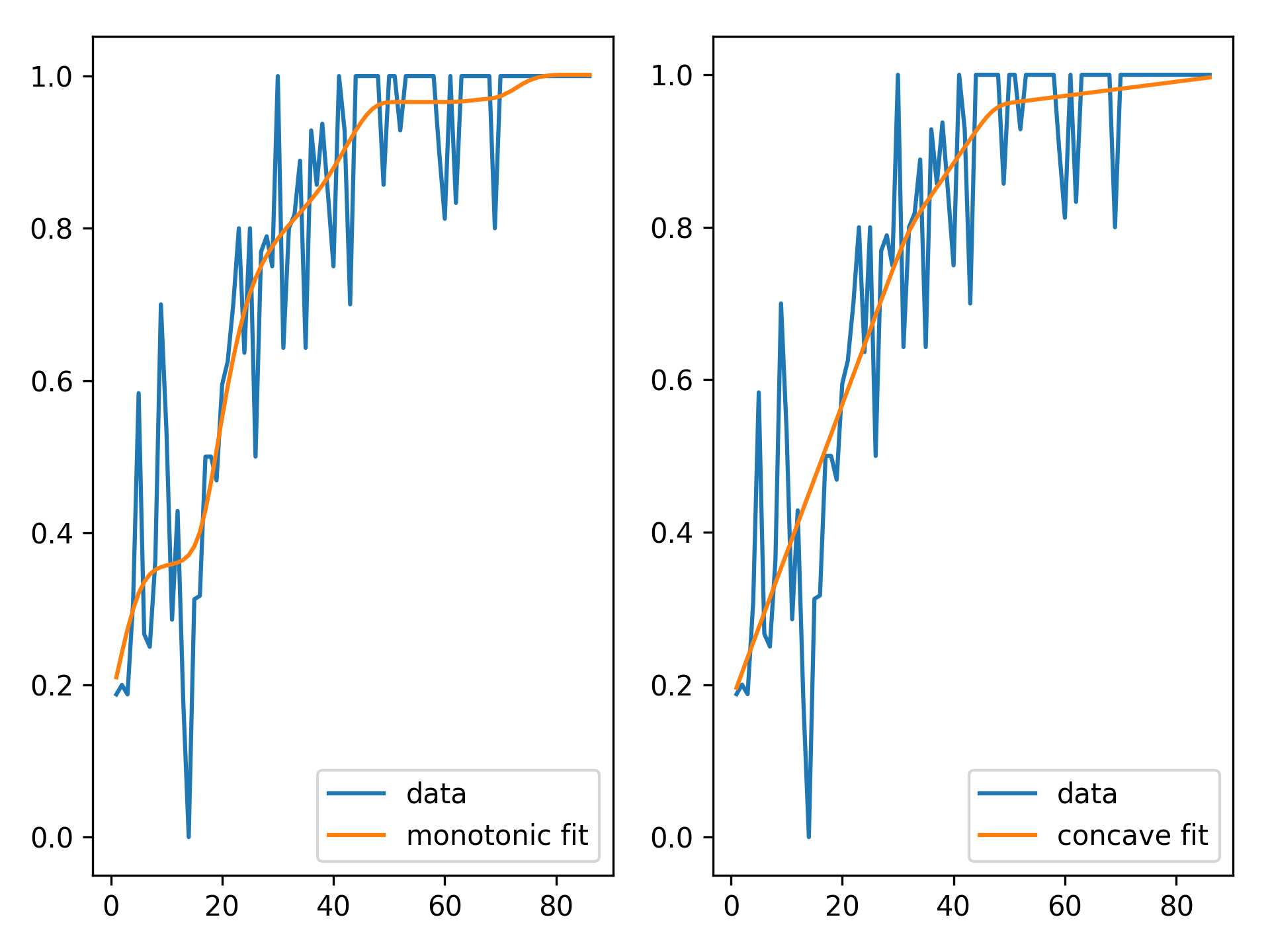
We can inject our intuition into our model by using monotonic and concave constraints:
from pygam import LinearGAM, s
from pygam.datasets import hepatitis
X, y = hepatitis(return_X_y=True)
gam1 = LinearGAM(s(0, constraints='monotonic_inc')).fit(X, y)
gam2 = LinearGAM(s(0, constraints='concave')).fit(X, y)
fig, ax = plt.subplots(1, 2)
ax[0].plot(X, y, label='data')
ax[0].plot(X, gam1.predict(X), label='monotonic fit')
ax[0].legend()
ax[1].plot(X, y, label='data')
ax[1].plot(X, gam2.predict(X), label='concave fit')
ax[1].legend()API
pyGAM is intuitive, modular, and adheres to a familiar API:
from pygam import LogisticGAM
from pygam.datasets import toy_classification
X, y = toy_classification(return_X_y=True)
gam = LogisticGAM(s(0) + s(1) + s(2) + s(3) + s(4) + f(5))
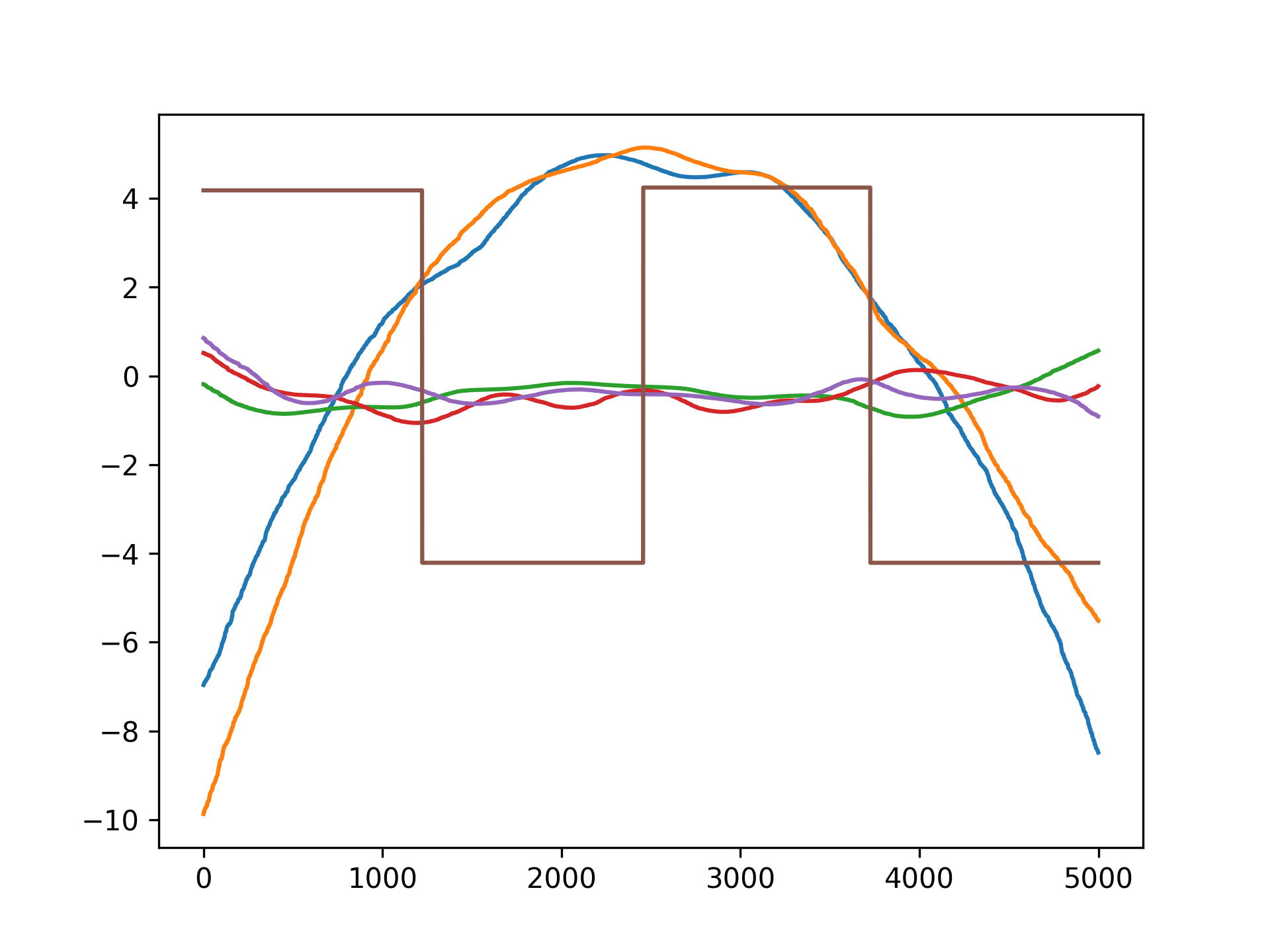
gam.fit(X, y)Since GAMs are additive, it is also super easy to visualize each individual feature function, f_i(X_i). These feature functions describe the effect of each X_i on y individually while marginalizing out all other predictors:
pdeps = gam.partial_dependence(X)
plt.plot(pdeps)Current Features
Models
pyGAM comes with many models out-of-the-box:
- GAM (base class for constructing custom models)
- LinearGAM
- LogisticGAM
- GammaGAM
- PoissonGAM
- InvGaussGAM
- ExpectileGAM
You can mix and match distributions with link functions to create custom models!
gam = GAM(distribution='gamma', link='inverse')Distributions
- Normal
- Binomial
- Gamma
- Poisson
- Inverse Gaussian
Link Functions
Link functions take the distribution mean to the linear prediction. These are the canonical link functions for the above distributions:
- Identity
- Logit
- Inverse
- Log
- Inverse-squared
Callbacks
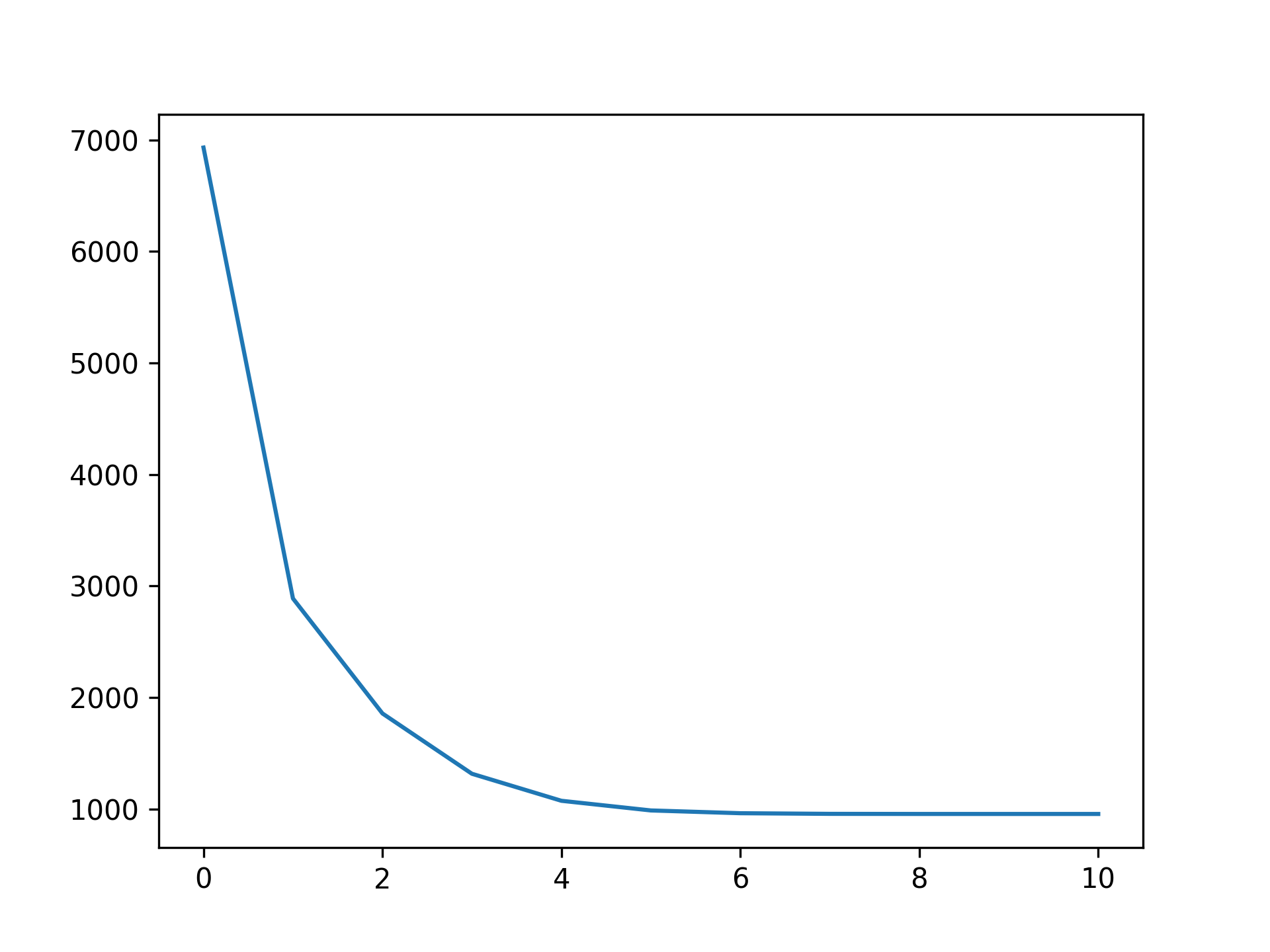
Callbacks are performed during each optimization iteration. It's also easy to write your own.
- deviance - model deviance
- diffs - differences of coefficient norm
- accuracy - model accuracy for LogisticGAM
- coef - coefficient logging
You can check a callback by inspecting:
plt.plot(gam.logs_['deviance'])Linear Extrapolation
Citing pyGAM
Please consider citing pyGAM if it has helped you in your research or work:
Daniel Servén, & Charlie Brummitt. (2018, March 27). pyGAM: Generalized Additive Models in Python. Zenodo. DOI: 10.5281/zenodo.1208723
BibTex:
@misc{daniel\_serven\_2018_1208723,
author = {Daniel Servén and
Charlie Brummitt},
title = {pyGAM: Generalized Additive Models in Python},
month = mar,
year = 2018,
doi = {10.5281/zenodo.1208723},
url = {https://doi.org/10.5281/zenodo.1208723}
}
References
-
Simon N. Wood, 2006
Generalized Additive Models: an introduction with R -
Hastie, Tibshirani, Friedman
The Elements of Statistical Learning
http://statweb.stanford.edu/~tibs/ElemStatLearn/printings/ESLII_print10.pdf -
James, Witten, Hastie and Tibshirani
An Introduction to Statistical Learning
http://www-bcf.usc.edu/~gareth/ISL/ISLR%20Sixth%20Printing.pdf -
Paul Eilers & Brian Marx, 1996 Flexible Smoothing with B-splines and Penalties http://www.stat.washington.edu/courses/stat527/s13/readings/EilersMarx_StatSci_1996.pdf
-
Kim Larsen, 2015
GAM: The Predictive Modeling Silver Bullet
http://multithreaded.stitchfix.com/assets/files/gam.pdf -
Deva Ramanan, 2008
UCI Machine Learning: Notes on IRLS
http://www.ics.uci.edu/~dramanan/teaching/ics273a_winter08/homework/irls_notes.pdf -
Paul Eilers & Brian Marx, 2015
International Biometric Society: A Crash Course on P-splines
http://www.ibschannel2015.nl/project/userfiles/Crash_course_handout.pdf -
Keiding, Niels, 1991
Age-specific incidence and prevalence: a statistical perspective