This repository provides a proof-of-concept implementation for the manuscript Deep Sets Are Viable Graph Learners.
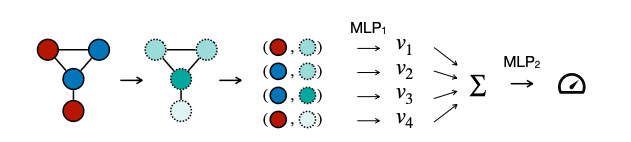
DSNN computes predictions on graphs, where the graph is represented as a multiset of nodes. Positional embeddings are computed using the following metrics:
- Centrality measures (e.g., betweenness, eigenvalue, laplacian centrality).
- The minimal distance to the node(s) with the highest/lowest centrality.
- The sum of neighboring nodes' features.
DSNN has two components (MLP_1 and MLP_2), each with nine layers. These components have a latent dimension of 64 and utilize residual connections.
For comparison, we employ models from PyTorch Geometric as our baselines, specifically GIN, PNA, and GCN. These models are configured with five layers and a latent dimension of 64.
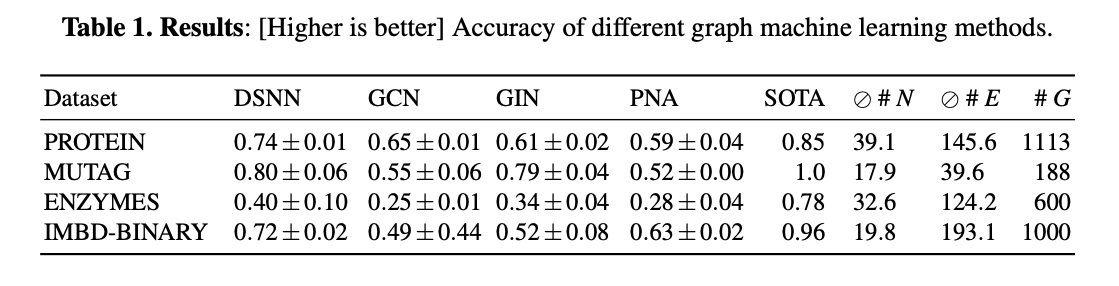
The number of parameters and the accuracy (higher is better) of each model are:
You can run DSNN locally using main.ipynb. First, install Anaconda, then create an environment with the Python dependencies (tested on OS X):
conda env create -f environment.yml -n dsnn
conda activate dsnn
jupyter labThen just run the notebook(s) from start to finish.
Install docker and then:
docker pull gerritgr/dsnn:latest
docker run -p 8888:8888 gerritgr/dsnn:latestYou need to manually copy the URL to your browser, navigate to the notebook, and activate the dsnnenv kernel (Kernel -> Change Kernel...).
- The positional encodings contain a sum of the values of all neighboring nodes (not only of their positional encodings).
- Accuracy decreased due to a calculation error in the split of train/val/test compared to the revision version.