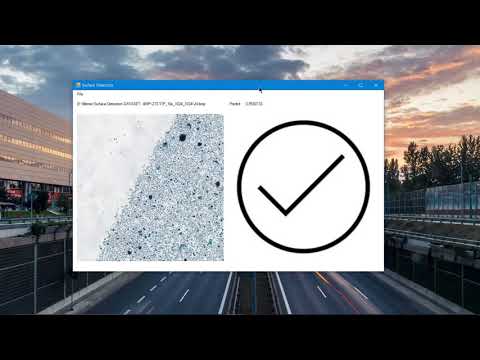
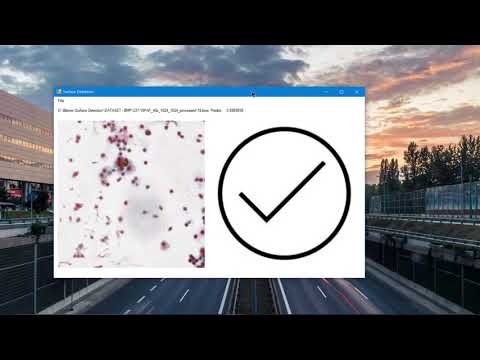
ANN model has been developed to make automatic surface scan. Our goal is to develop ANN that decides whether there is a tissue surface in the input image. If the input image contains the tissue surface, it returns 1 value.
Also, this repo is a reference for testing Keras model weights (file with .h5 extension) using C ++. Dense, Conv2D, MaxPooling2D, Flatten, BatchNormalization layers used in Keras library; ReLU, Sigmoid (0-1 range) activation functions are coded using C ++.
Keras model consists of Convolution, MaxPooling, Flatten, Dense and Batch Normalization layers. The model takes the of RGB image 512x512x3 from the input. Output Layer consists of 1 neuron and returns range of (0-1).
inputImage = new CpuMat(512, 512, 3, false); // Dense(input_image.Height, input_image.Width, input_image.Depth, useBias = false)
conv = new Conv2D(8, 3, 3, inputImage); // Conv2D(number of filters, filter_height, filter_width, inputImage)
maxPool = new MaxPooling2D(conv->Result, 2, 2, 2, 2); // Each layer gets the result values of the previous layer
conv1 = new Conv2D(16, 3, 3, maxPool->Result);
maxPool1 = new MaxPooling2D(conv1->Result, 2, 2, 2, 2);
flatten = new Flatten(maxPool1->Result);
dense = new Dense(16, flatten->Result->Rows, flatten->Result->Cols);
batchNorm = new BatchNormalization(dense->Result->Rows, dense->Result->Cols);
dense1 = new Dense(1, dense->Result->Rows, dense->Result->Cols, false); // Yeaaap, dense1 is end layerThen the weights of each layers are loaded. For example 10X scale model weights:
std::string weightFolder = "..\\database\\model_save_10X_BGR\\"
std::string conv2DKernel = weightFolder + "conv2d\\kernel.txt";
std::string conv2DBias = weightFolder + "conv2d\\bias.txt";
std::string conv2D1Kernel = weightFolder + "conv2d_1\\kernel.txt";
std::string conv2D1Bias = weightFolder + "conv2d_1\\bias.txt";
std::string denseKernel = weightFolder + "dense\\kernel.txt";
std::string denseBias = weightFolder + "dense\\bias.txt";
std::string dense1Kernel = weightFolder + "dense_1\\kernel.txt";
std::string dense1Bias = weightFolder + "dense_1\\bias.txt";
std::string batchNormBeta = weightFolder + "batch_normalization\\beta.txt";
std::string batchNormGamma = weightFolder + "batch_normalization\\gamma.txt";
std::string batchNormMovingMean = weightFolder + "batch_normalization\\moving_mean.txt";
std::string batchNormMovingVariance = weightFolder + "batch_normalization\\moving_variance.txt";
// load kernel and bias weights
conv->load(conv2DKernel, conv2DBias);
conv1->load(conv2D1Kernel, conv2D1Bias);
dense->load(denseKernel, denseBias);
dense1->load(dense1Kernel, dense1Bias);
// load batchnormalization layer weights
batchNorm->load(batchNormBeta, batchNormGamma, batchNormMovingMean, batchNormMovingVariance);
The image to be tested must be set to the InputImage-> CpuP pointer.
Then, each layer is applied to inputImage.
conv->apply(inputImage);
cpuRelu(conv->Result); // apply ReLU activation
maxPool->apply(conv->Result);
conv1->apply(maxPool->Result);
cpuRelu(conv1->Result);
maxPool1->apply(conv1->Result);
flatten->apply(maxPool1->Result);
dense->apply(flatten->Result);
cpuRelu(dense->Result);
batchNorm->apply(dense->Result);
dense1->apply(dense->Result);
cpuSigmoid(dense1->Result); // apply Sigmoid (Unipolar) activationFinally the predict value is read from pointer CpuP of end layer.
predict = (float*)dense1->Result->CpuP;Keras weights are in hdf5 file format. I assume you got the model record as .json and .h5. You can create your model training weights as follows:
# keras library import for Saving and loading model and weights
from keras.models import model_from_json
from keras.models import load_model
# serialize model to JSON
# the keras model which is trained is defined as 'model' in this example
model_json = model.to_json()
with open("model_save_json.json", "w") as json_file:
json_file.write(model_json)
# serialize weights to HDF5
model.save_weights("model_save_weight.h5")It is converted to a text file for use with the C environment. You can do it as follows:
python h5_to_txt.py model_save_weight.h5Each layer in the model will be saved in a folder and their weight in it. The text files will then be loaded into the model layers.
Tissue surface prediction results for 10X and 40X scale images