TreeTime: maximum likelihood dating and ancestral inference for phylogenetic trees
Overview
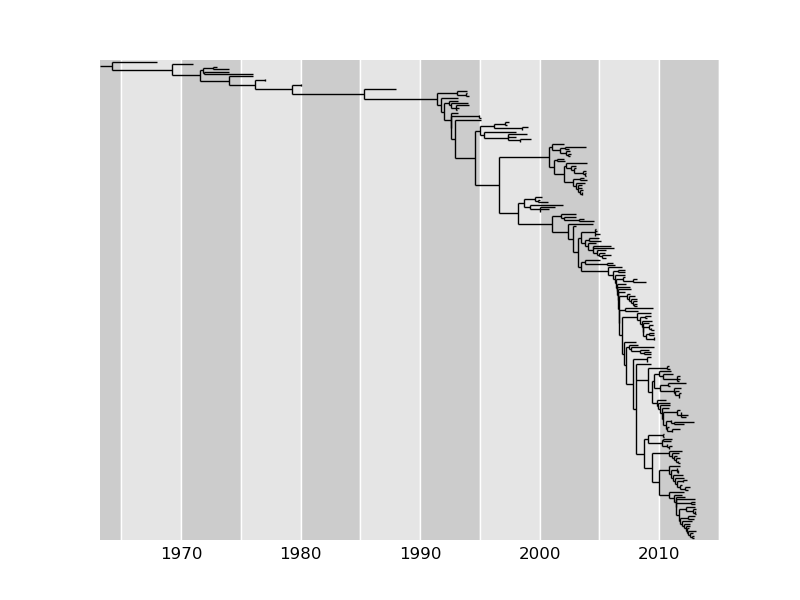
TreeTime provides routines for ancestral sequence reconstruction and the inference of molecular-clock phylogenies, i.e., a tree where all branches are scaled such that the locations of terminal nodes correspond to their sampling times and internal nodes are placed at the most likely time of divergence.
TreeTime is implemented in Python 2.7 -- an experimental port to Python 3 is available in a separate branch. TreeTime aims at striking a compromise between sophisticated probabilistic models of evolution and fast heuristics. It implements GTR models of ancestral inference and branch length optimization, but takes the tree topology as given. To optimize the likelihood of time-scaled phylogenies, treetime uses an iterative approach that first infers ancestral sequences given the branch length of the tree, then optimizes the positions of unconstrained nodes on the time axis, and then repeats this cycle. The only topology optimization are (optional) resolution of polytomies in a way that is most (approximately) consistent with the sampling time constraints on the tree. The package is designed to be used as a stand-alone tool or as a library used in larger phylogenetic analysis work flows.
Features
- ancestral sequence reconstruction (marginal and joint maximum likelihood)
- molecular clock tree inference (marginal and joint maximum likelihood)
- inference of GTR models
- rerooting to obtain best root-to-tip regression
- auto-correlated relaxed molecular clock (with normal prior)
Table of contents
- Installation and prerequisites
- Basic usage
- Example scripts
- Comparable Tools
- Projects using treetime
- Developer info
Installation and prerequisites
TreeTime is written in Python 2.7 and currently doesn't support Python 3.
-
The package depends on several python libraries:
-
numpy, scipy: for all kind of mathematical operations as matrix operations, numerical integration, interpolation, minimization, etc.
-
BioPython: for parsing multiple sequence alignments and all phylogenetic functionality
-
matplotlib: optional dependency for plotting If you do not have these libraries, you can install them by typing in the terminal:
$pip install numpy scipy biopython matplotlib -
-
To install the package, run
setup.pyscript from the terminal:$python setup.py install
You might need root privileges for system wide installation. Alternatively, you can simply use it TreeTime locally without installation. In this case, just download and unpack it, and then add the TreeTime folder to your $PYTHONPATH.
Building the documentation
The API documentation for the treetime package is generated created with Sphinx. The source code for the documentaiton is located in doc folder.
Requirements
- sphinx-build to generate static html pages from source. Installed as
$pip install Sphinx- basicstrap Html theme for sphinx:
$pip install sphinxjp.themes.basicstrapAfter required packages are installed, navifgate to doc directory, and build the docs by typing:
$make htmlInstead of html, another target as latex or epub can be specified to build the docs in the desired format.
Requirements
To build the documentation, sphinx-build tool should be installed. The doc pages are using basicstrap html theme to have the same design as the treetime web server. Therefore, the basicstrap theme should be also available in the system
Basic usage
TreeTime can be used as part of python programs that create and interact with tree time objects. How treetime can be used to address typical questions like ancestral sequence reconstruction, rerooting, timetree inference etc is illustrated by a collection of example scripts described below.
In addition, we provide scripts that can be run from the command line with arguments specifying input data and parameters.
Ancestral sequence reconstruction:
To perform ancestral sequence reconstruction, use the script ancestral_inference.py.
usage: ancestral_reconstruction.py [-h] --aln ALN --tree TREE [--marginal]
[--infer_gtr]
Reconstructs ancestral sequences and maps mutations to the tree. The output
consists of a file ending with _ancestral.fasta with ancestral sequences and a
tree ending with _mutation.newick with mutations appended to node names as
_A45G_... The inferred GTR model is written to stdout
optional arguments:
-h, --help show this help message and exit
--aln ALN fasta file with input sequences
--tree TREE newick file with tree
--marginal marginal instead of joint ML reconstruction
--infer_gtr infer substitution model
--keep_overhangs keep 5' and 3' gaps rather than filling them with ancestral sequence
Alternatively, you can directly interact with the class TreeAnc from treetime as follows
python from treetime import TreeAnc ta = TreeAnc(tree='my_tree.nwk', aln='my_seqs.nwk', gtr='JC69') ta.infer_ancestral_sequences(infer_gtr=True, marginal=False)
Every node of ta.tree now has a node.sequence attached. With the optional argument infer_gtr=True, a maximum likelihood GTR model is inferred and overwrites the initial one, the option marginal=True can be used to construct a marginal rather than joint maximum likelihood reconstruction.
The tree and alignment arguments can be either file names (newick and fasta) or Biopython tree and alignment objects.
After running ta.infer_ancestral_sequences(), the each node in the tree
will have a node.sequence attached and node.mutations contains the difference
of that sequence to the sequence of the parent node.
Other useful options to the constructor include fill_overhangs. If TreeAnc is
initialized with fill_overhangs=True, 5' or 3' gaps will be treated as missing data and filled with the sequence of the nearest ancestral node.
The inferred substitution model is accessible via print(tt.gtr) and the equilibrium character frequencies are stored in tt.gtr.pi, the symmetric substitution matrix in tt.gtr.W.
A longer example of the usage is available in examples/ancestral_inference.py
Molecular clock phylogenies
To infer molecular clock phylogenies, use the script timetree_inference.py:
usage: timetree_inference.py [-h] --aln ALN --tree TREE --dates DATES
[--infer_gtr] [--reroot REROOT]
[--optimize_branch_length] [--keep_polytomies]
[--relax [RELAX [RELAX ...]]]
[--max_iter MAX_ITER] [--verbose VERBOSE]
[--Tc TC] [--plot]
Reconstructs ancestral sequences and infers a molecular clock tree. The script
produces an alignment file ending on _ancestral.fasta which contains the
inferred ancestral sequences and a tree file ending on _timetree.nexus.
Inferred mutations are included as comments. The molecular clock, along with
the inferred GTR model, is written to stdout)
optional arguments:
-h, --help show this help message and exit
--aln ALN fasta file with input sequences
--tree TREE newick file with tree
--dates DATES csv with dates for nodes with 'node_name, date' where
date is float (as in 2012.15)
--infer_gtr infer substitution model
--reroot REROOT reroot the tree. Valid arguments are 'best',
'midpoint', or a node name
--optimize_branch_length
Reoptimize branch length. Note that branch length
optimized by treetime are only accurate at short
evolutionary distances.
--keep_polytomies Don't resolve polytomies using temporal information.
--relax [RELAX [RELAX ...]]
use an autocorrelated molecular clock. Prior strength
and coupling of parent and offspring rates can be
specified e.g. as --relax 1.0 0.5
--max_iter MAX_ITER maximal number of iterations the inference cycle is
run
--verbose VERBOSE verbosity of output 0-6
--Tc TC coalescent time scale -- sensible values are on the
order of the average hamming distance of
contemporaneous sequences. In addition, "opt"
"skyline" are valid options and estimate a constant
coalescent rate or a piecewise linear coalescent rate
history
--plot plot the tree on a time axis and save as pdf
Alternatively, you can interact directly with the TreeTime class from within a script or the python console. TreeTime is a subclass of TreeAnc, so all its methods are available.
```python
from treetime import TreeTime
tt = TreeTime(dates=mydates, tree='my_tree.nwk', aln='my_seqs.nwk', gtr='JC69')
tt.run(root='best', infer_gtr=True, Tc='skyline', resolve_polytomies=True, max_iter=2)
```
Every node of tt.tree will be assigned a numdate and time_before_present attribute.
The numdate attribute has units of years, the time_before_present attribute has units of (inverse) substitution rate. The molecular clock estimate can be obtained by print(tt.date2dist).
The additional argument resolve_polytomies specifies whether TreeTime will attempt to resolve multiple mergers using the temporal constraints on leaves.
The optional argument Tc specifies if TreeTime will use a coalescent prior. Tc can either be a float, which is interpreted as coalescence time scale in units of the inverse clock rate (values similar to average pairwise sequence distance are sensible), or Tc='opt' in which case TreeTime will optimize Tc to maximize the likelihood. Finally, Tc='skyline' will fit a piecewise linear function as coalescent rate history. To access the result, use
skyline = tt.merger_model.skyline_inferred(gen=50)
The result will be an interpolation object the return the "effective population size", its pivots are available as skyline.x and skyline.y.
The argument gen=50 specifies the number of generations per year to convert the rate estimate into population size.
The optional argument niqd takes a float and will activate a filter that marks all nodes that deviate more than niqd inter-quartile distance from the molecular clock. This is done via the method tt.clock_filter that optionally plots a root-to-tip distance vs time scatter plot.
In addition, an autocorrelated relaxed clocks can be used by passing a tuple of two numbers (slack, coupling). slack is the strength of the normal prior on rate variation, coupling penalizes rate variation between parents and children.
Longer examples of treetime usage are available in the scripts examples/relaxed_clock.py, examples/rerooting_and_timetrees.py, and examples/ebola.py.
Quantify temporal signal in phylogenies and reroot to the maximize "clock-i-ness"
The script temporal_signal.py provides functionality analogous to TempEst by Andrew Rambaut.
```
usage: temporal_signal.py [-h] --tree TREE --dates DATES [--aln ALN]
[--infer_gtr] [--reroot] [--plot]
[--verbose VERBOSE]
Calculates the root-to-tip regression and quantifies the 'clock-i-ness' of the
tree. It will optionally reroot the tree to maximize the clock-like signal and
recalculate branch length.
optional arguments:
-h, --help show this help message and exit
--tree TREE newick file with tree
--dates DATES csv with dates for nodes with 'node_name, date' where
date is float (as in 2012.15)
--aln ALN fasta file with input sequences
--infer_gtr infer substitution model
--reroot reroot the tree to maximize the correlation of root-to-
tip distance with sampling time
--plot
--verbose VERBOSE verbosity of output 0-6
```
The slope of the regression of root-to-tip distance vs sampling date will be printed to stdout along with the fraction of variance explained by the linear regression. By passing the flag --reroot, treetime will search for the root that maximizes the correlation of root-to-tip distance with time and reroot the tree. The option --plot will produce a scatter plot with the best regression and save it to file.
Example scripts
The following scripts illustrate how treetime can be used to solve common problem with short python scripts. They are meant to be used in an interactive ipython environment and run as run examples/ancestral_inference.py.
ancestral_inference.pyillustrates how ancestral sequences are inferred and likely mutations are assigned to branches in the tree,relaxed_clock.pywalks the user through the usage of relaxed molecular clock models.examples/rerooting_and_timetrees.pyillustrates the rerooting and root-to-tip regression scatter plots.ebola.pyuses about 300 sequences from the 2014-2015 Ebola virus outbreak to infer a timetree. This example takes a few minutes to run.
Comparable Tools
There are several other tools which estimate molecular clock phylogenies.
- Beast relies on the MCMC-type sampling of trees. It is hence rather slow for large data sets. But BEAST allows the flexible inclusion of prior distributions, complex evolutionary models, and estimation of parameters.
- Least-Square-Dating (LSD) emphasizes speed (it scales as O(N) as TreeTime), but provides limited scope for customization.
- treedater by Eric Volz and Simon Frost is an R package that implements time tree estimation and supports relaxed clocks.
Projects using treetime
Treetime is an integral part of the nextstrain.org project to track and analyze viral sequence data in real time.
Developer info
- Credits -- .
- Copyright and License: Pavel Sagulenko and Richard Neher, MIT Licence
- How to contribute
- References