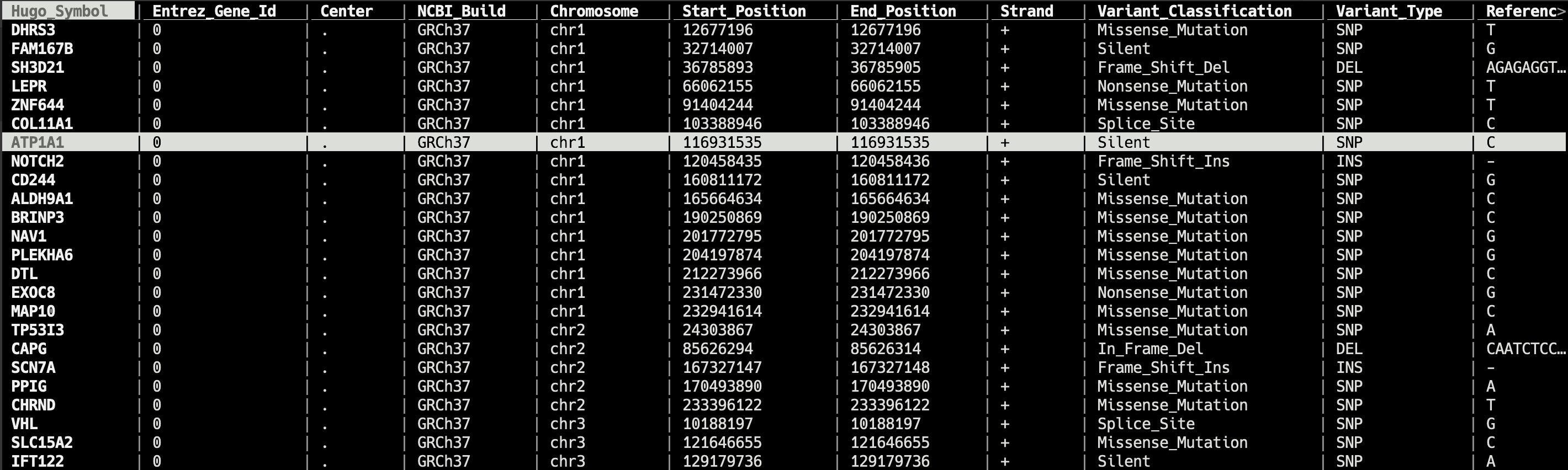
bv
Data Viewer in Terminal for Bioinformatician
Table of Contents
- bv
- Table of Contents - Description
- Feature
- To do
- Supported file types
- Installation
- Key binding
- Usage
Description
bv is a tool to view the common bioinformatics data file in terminal.The TUI of bv is modifyied from vdtui
Feature
- Spreadsheet-like view for biological delimited data
- Vim-like key binding
- Support for gzip compressed file
- Automatically identify unknown file type's delimiter
- User-defined format processing configure
To do
- support for bam and fastaq format
- view data from url
Supported file types
| File type | filename extension | description |
|---|---|---|
| csv | .csv | Delimited text file that uses a comma to separate values |
| tsv | .tsv | Delimited text file that uses a tab to separate values |
| excel | .xlsx | Microsoft Excel is a spreadsheet developed by Microsoft |
| vcf | .vcf | The Variant Call Format (VCF) specifies the format of a text file used in bioinformatics for storing gene sequence variations |
| bed | .bed | A BED file is a tab-delimited text file that defines a feature track |
| maf | .maf | Mutation Annotation Format (MAF) is a tab-delimited text file with aggregated mutation information from VCF Files and are generated on a project-level |
| gff | .gff | The GFF (General Feature Format) format consists of one line per feature, each containing 9 columns of data, plus optional track definition lines |
| gtf | .gtf | The Gene transfer format (GTF) is a file format used to hold information about gene structure |
Installation
Linux and macOS
pip
$ pip install bvconda
$ conda install -c codechenx bv Window
Not Support
Key binding
| Key | description |
|---|---|
| q | quit |
| h, left arrow | go one column left |
| l, right arrow | go one column right |
| j, down arrow | go one row down |
| k, up | go one row up |
| gg, gk | go to top row |
| G, gj | go to bottom row |
| gh | go to leftmost column |
| gl | go to rightmost column |
| ctrl-f, page down | scroll one page down |
| ctrl-b, page up | scroll one page up |
| < | move up to previous value in this column |
| > | move down to next value in this column |
| / | search this column forward for regex |
| ? | search this column backward for regex |
| g/ | search regex forward in all visible columns |
| g? | search regex backward in all visible columns |
| n | go to next match |
| p | go to previous match |
| s, space | select this row |
| u | unselect this row |
| gu | unselect all rows |
| { | move to previous selected row |
| } | move to next selected row |
| [ | sort by this column ascending |
| ] | sort by this column descending |
| - | hide this column |
| ctrl-l | redraw entire terminal screen |
| ctrl-g | show info for the current sheet |
| z?, F1 | open command help sheet |
Usage
usage: bv [-h] [-s S] [-ss SS] [-sn SN] [-rc RC [RC ...]] [-hc HC [HC ...]]
[-type {csv,tsv,vcf,maf,gff,gtf,bed,xlsx}] [--noheader] [--trans]
[--compressed]
filename
positional arguments:
filename file name
optional arguments:
-h, --help show this help message and exit
-s S delimiter
-ss SS ignore lines with specific prefix
-sn SN ignore first n lines
-rc RC [RC ...] only show columns(support for multiple arguments,
separated by space)
-hc HC [HC ...] hide columns(support for multiple arguments, separated
by space)
-type {csv,tsv,vcf,maf,gff,gtf,bed}
specify a file type to file manual
--noheader not to use fist line as header
--trans view transposed data
--compressed file is compressed?