Modern C++ framework for Symbolic Regression
Operon is a modern C++ framework for symbolic regression that uses genetic programming to explore a hypothesis space of possible mathematical expressions in order to find the best-fitting model for a given regression target. Its main purpose is to help develop accurate and interpretable white-box models in the area of system identification. More in-depth documentation available at https://operongp.readthedocs.io/.
How does it work?
Broadly speaking, genetic programming (GP) is said to evolve a population of "computer programs" ― AST-like structures encoding behavior for a given problem domain ― following the principles of natural selection. It repeatedly combines random program parts keeping only the best results ― the "fittest". Here, the biological concept of fitness is defined as a measure of a program's ability to solve a certain task.
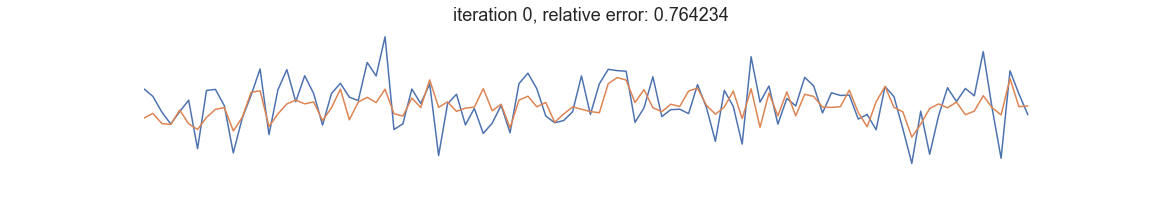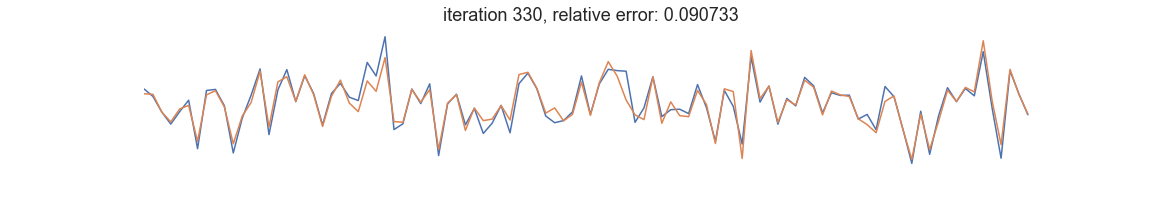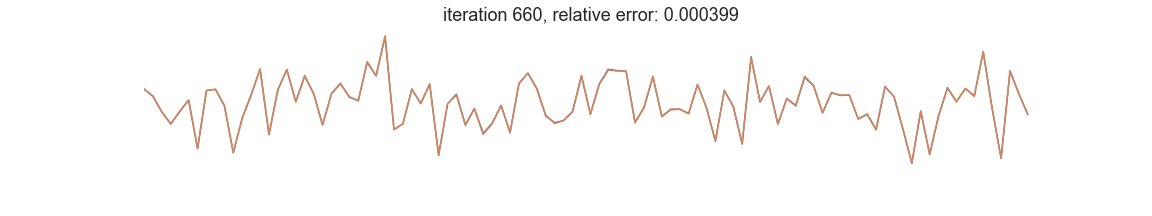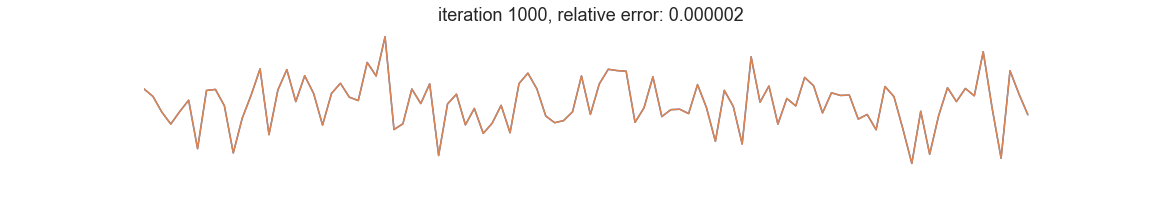
In symbolic regression, the programs represent mathematical expressions typically encoded as expression trees. Fitness is usually defined as goodness of fit between the dependent variable and the prediction of a tree-encoded model. Iterative selection of best-scoring models followed by random recombination leads naturally to a self-improving process that is able to uncover patterns in the data:
Build instructions
The project requires CMake and a C++17 compliant compiler. Using the git versions of Eigen and Ceres is recommended. On Windows we recommend building with MinGW or with your WSL distro.
Required dependencies
Optional dependencies
- cxxopts required for the cli app.
- doctest required for unit tests.
- python and pybind11 required to build the python bindings.
These libraries are well-known and should be available in your distribution's package repository. On Windows they can be easily managed using vcpkg. CMake will download the following header-only libraries during the build generation phase: microsoft-gsl, rapidcsv, nanobench and xxhash.
Build options
The following options can be passed to CMake:
| Option | Description |
|---|---|
-DUSE_SINGLE_PRECISION=ON |
Perform model evaluation using floats (single precision) instead of doubles. Great for reducing runtime, might not be appropriate for all purposes. |
-DUSE_OPENLIBM=ON |
Link against Julia's openlibm, a high performance mathematical library (recommended to improve consistency across compilers and operating systems). |
-DBUILD_TESTS=ON |
Build the unit tests. |
-DBUILD_PYBIND=ON |
Build the Python bindings. |
-DUSE_JEMALLOC=ON |
Link against jemalloc, a general purpose malloc(3) implementation that emphasizes fragmentation avoidance and scalable concurrency support (mutually exclusive with tcmalloc). |
-DUSE_TCMALLOC=ON |
Link against tcmalloc (thread-caching malloc), a malloc(3) implementation that reduces lock contention for multi-threaded programs (mutually exclusive with jemalloc). |
Windows / VCPKG
- Install vcpkg following the instructions from https://github.com/Microsoft/vcpkg
- Install the required dependencies:
vcpkg install <deps> cd <path/to/operon>mkdir build && cd buildcmake .. -G"Your Visual Studio Version" -DCMAKE_TOOLCHAIN_FILE=[vcpkg root]\scripts\buildsystems\vcpkg.cmakecmake --build . --config Release
GNU/Linux
- Install the required dependencies
mkdir build && cd buildcmake .. -DCMAKE_BUILD_TYPE=Release. UseDebugfor a debug build, or useCC=clang CXX=clang++to build with a different compiler.make. AddVERBOSE=1to get the full compilation output or-jfor parallel compilation.
Usage
- Run
operon-gp --helpto see the usage of the console client. This is the easiest way to just start modeling some data. The program expects a csv input file and assumes that the file has a header. - The Python script provided under
scriptswraps theoperon-gpbinary and can be used to run bigger experiments. Data can be provided ascsvorjsonfiles containing metadata (seedatafolder for examples). The script will run a grid search over a parameter space defined by the user. - Several examples (C++ and Python) are available here
Installing the Python bindings
Operon comes with Python bindings as well as a scikit learn estimator. To build the bindings the option -DBUILD_PYBIND=TRUE must be passed to CMake. The desired install path can be specified using the CMAKE_INSTALL_PREFIX variable (for example, -DCMAKE_INSTALL_PREFIX=/usr/local/lib/python3.8/site-packages). If an install prefix is not provided CMake will try to detect the default path as reported by Python.
Then, the Python module and package can be installed with cmake --install . or make install (with sudo if needed).
Usage
Sklearn estimator
from operon.sklearn import SymbolicRegressor
reg = SymbolicRegressor()
# usual sklearn stuff
reg.fit(X, y)Operon library
from operon import Dataset, RSquared, etc.