This is the repository for our benchmark paper MedIAnomaly: A comparative study of anomaly detection in medical images.
- Python 3.10
- PyTorch 2.1.2
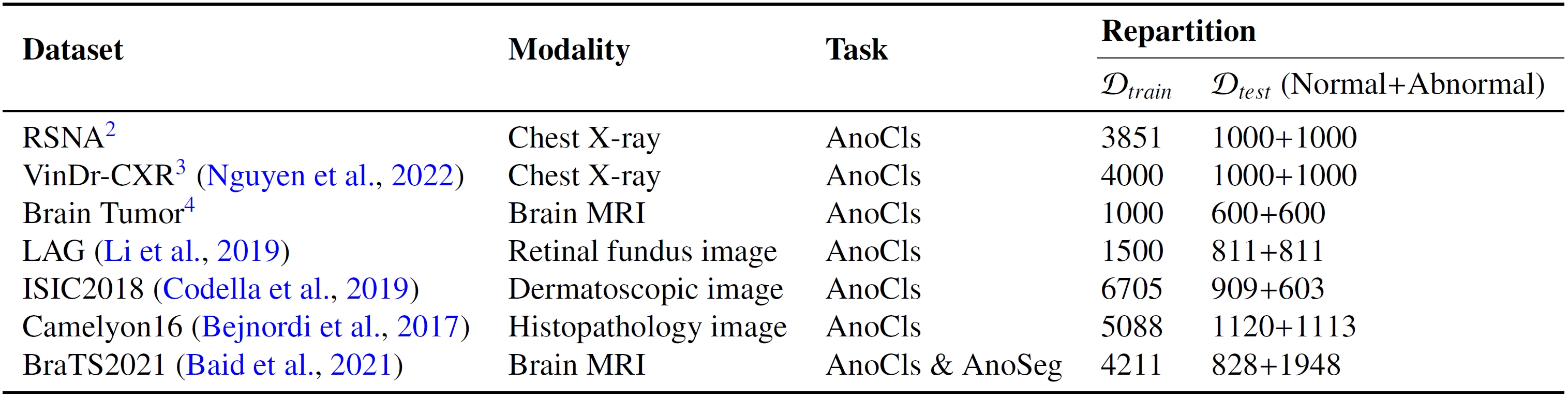
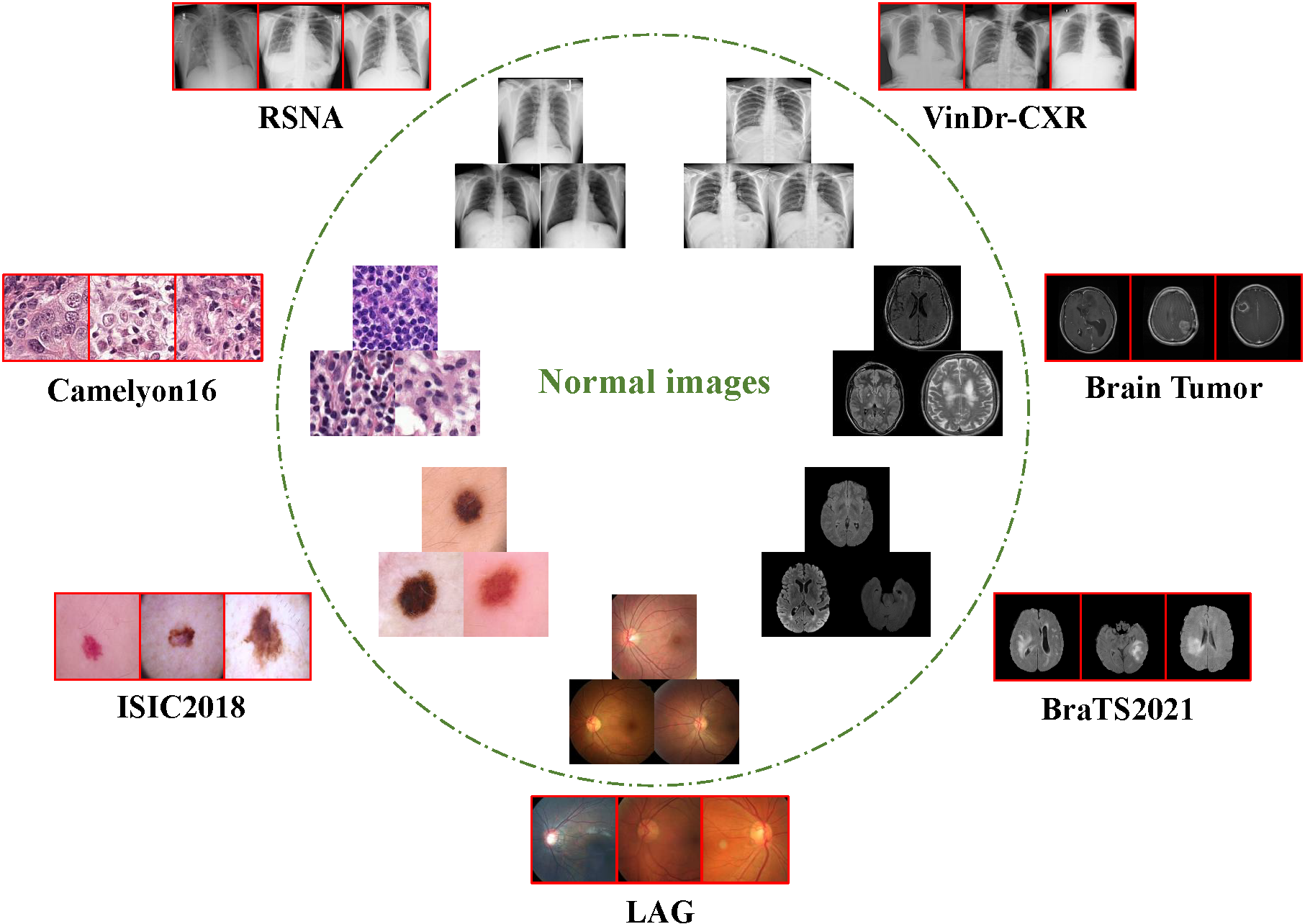
We provide the pre-processed seven datasets.
- Download the pre-processed datasets from: MedIAnomaly-Data
- Unzip the datasets via:
tar -zxvf RSNA.tar.gz
tar -zxvf VinCXR.tar.gz
tar -zxvf BrainTumor.tar.gz
tar -zxvf LAG.tar.gz
tar -zxvf ISIC2018_Task3.tar.gz
tar -zxvf Camelyon16.tar.gz
tar -zxvf BraTS2021.tar.gz- Place the
MedIAnomaly-Datadirectory in the user's home directory, i.e.,~/MedIAnomaly-Data/. (Otherwise, you need to modify the data root in your code.)
Finally, the data path should have the following structure:
~/MedIAnomaly-Data
├─RSNA
│ ├─images
│ └─data.json
├─VinCXR
│ ├─images
│ └─data.json
├─BrainTumor
│ ├─images
│ └─data.json
├─LAG
│ ├─images
│ └─data.json
├─ISIC2018_Task3
│ ├─ISIC2018_Task3_Training_Input
│ ├─ISIC2018_Task3_Training_GroundTruth
│ ├─ISIC2018_Task3_Test_Input
│ └─ISIC2018_Task3_Test_GroundTruth
├─Camelyon16
│ ├─train
│ │ ├─good
│ ├─test
│ │ ├─good
│ └─ └─Ungood
├─BraTS2021
│ ├─train
│ ├─test
│ │ ├─normal
│ │ ├─tumor
└─ └─ └─annotation
-
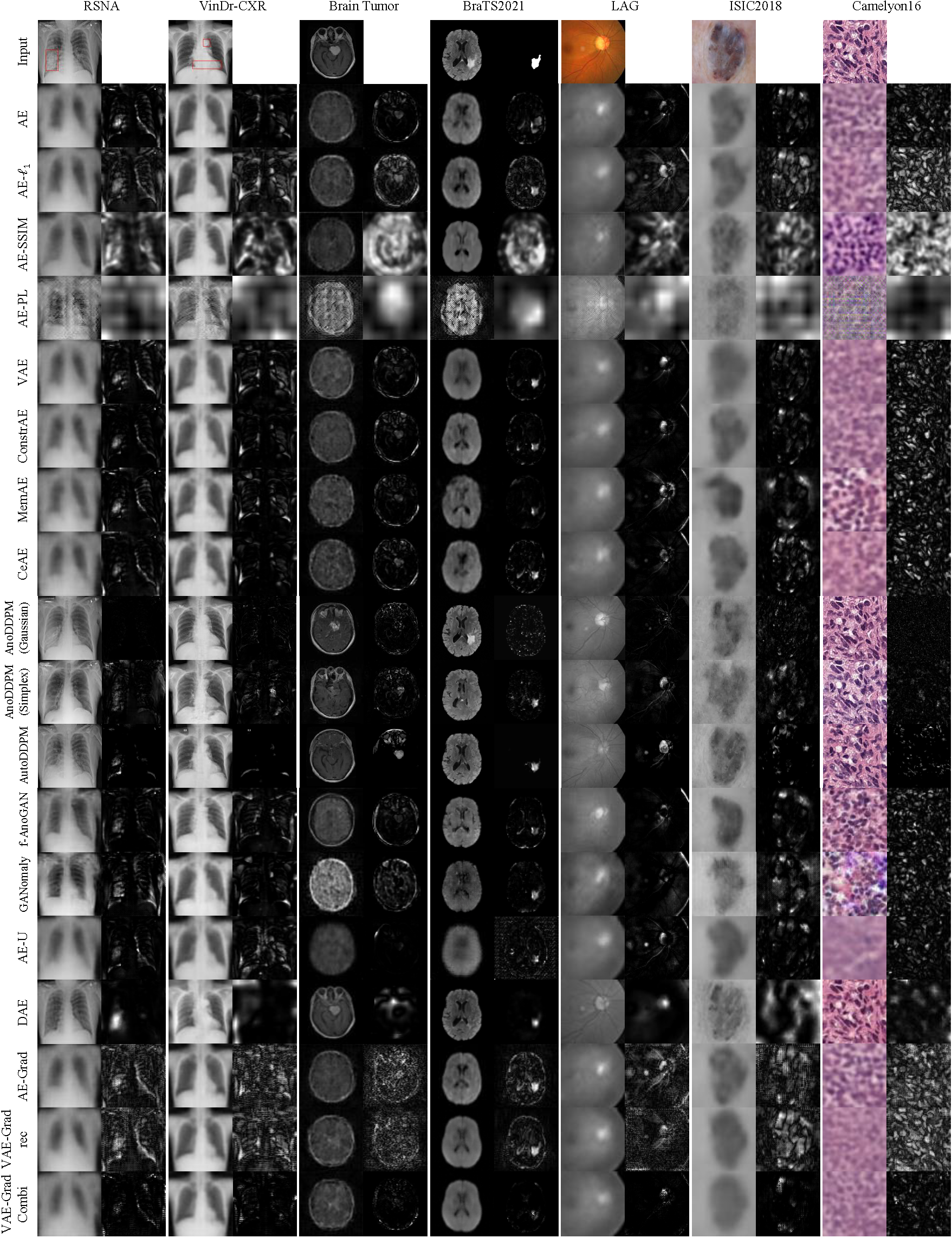
AE (
$\ell_2$ ,$\ell_1$ , SSIM, Perceptual Loss) -
AE-Spatial
-
VAE
-
Constrained AE
-
MemAE
-
CeAE
-
GANomaly
-
AE-U
-
DAE
-
AE-Grad
-
VAE-Grad (
$Grad_{ELBO}$ ,$Grad_{KL}$ ,$Grad_{rec}$ ,$Grad_{Combi}$ )
Train and evaluate these methods via:
cd reconstruction/;
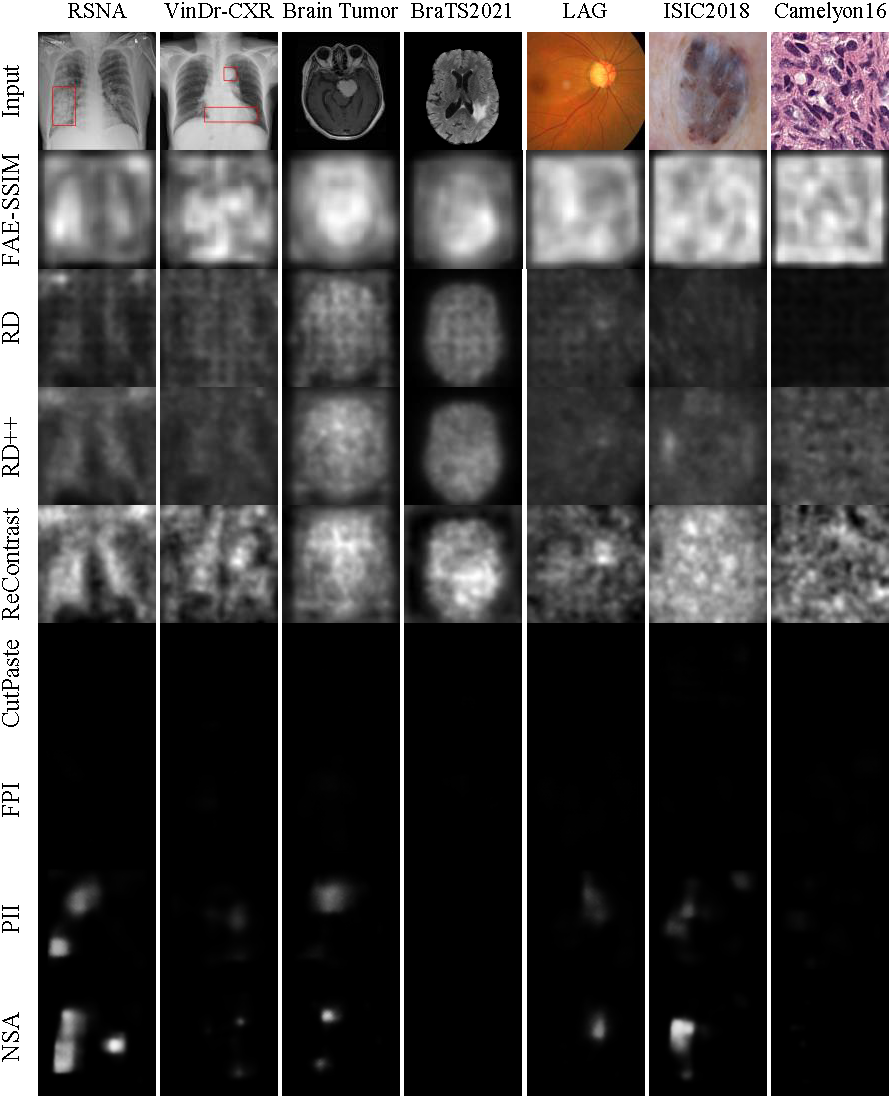
./train_eval.sh- CutPaste
- FPI
- PII
- NSA
Train and evaluate these methods via:
cd ssl/one_stage/;
./train_eval.sh- CutPaste
- AnatPaste
- ResNet18-ImageNet
Train and evaluate these methods via:
cd ssl/two_stage/;
./train_eval.shSome datasets and codes in this repository are based on DDAD-ASR, BMAD, NSA, CutPaste, AnatPaste. We thank the original authors for their excellent work.
If any questions, feel free to contact Yu Cai: yu.cai@connect.ust.hk.