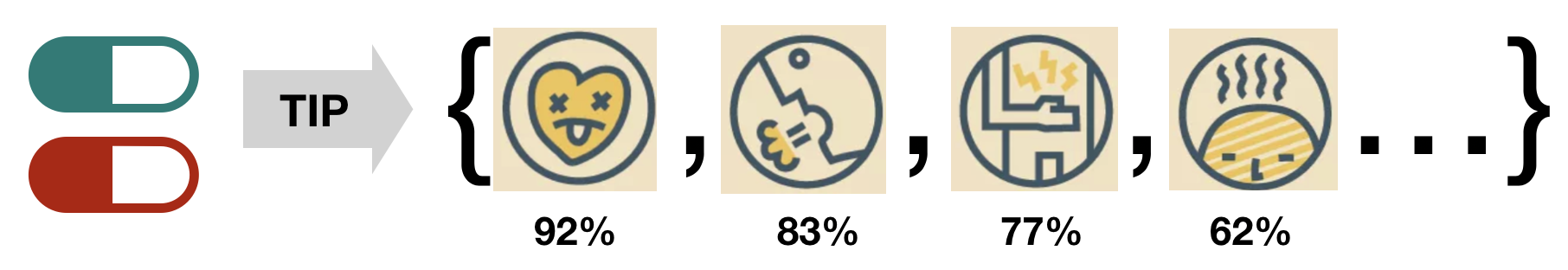
TIP is an efficient general approach for multi-relational link prediction in any multi-modal (i.e. heterogeneous and multi-relational) network with two types of nodes. It can also be applied to the Knowledge Graph Completion and Recommendation task. TIP model is inspired by the Decagon and R-GCN models, motivated by their limitations of high computational cost and memory demand when graph goes really complex. TIP improves their link prediction accuracy, and time and space efficiency of node representation learning. See details on the algorithm in our paper (Xu, Sang, and Lu, 2019).
we are particularly concerned about the safety of polypharmacy, which is the concurrent use of multiple medications by a patient. Given a pair of drug (:pill:,:pill:), the TIP model will predict how many polypharmacy side effects the drug pair will have, and what are the possibilities.
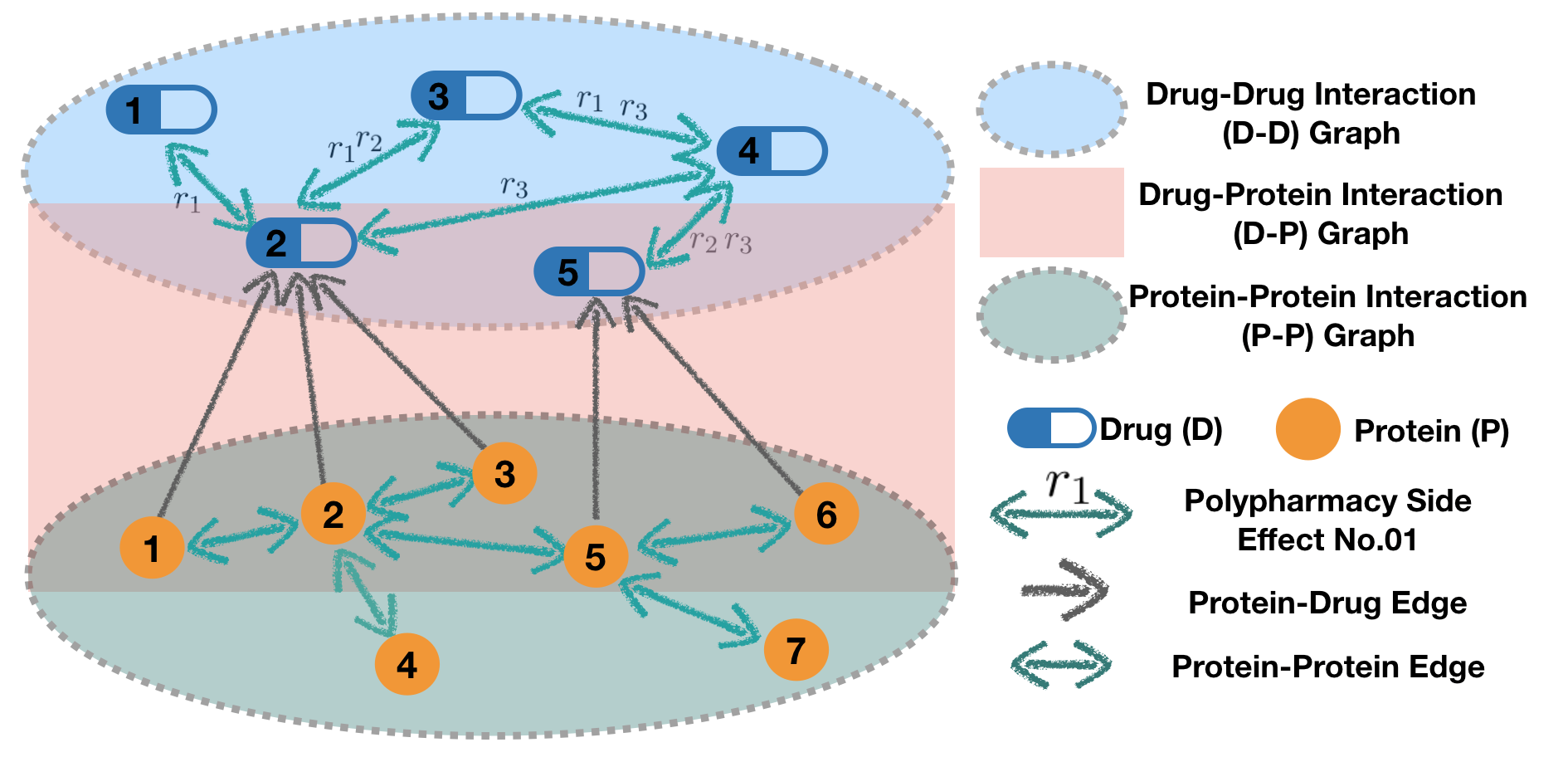
We use POSE clinical records and pharmacological information to construct a multi-modal biomedical graph with two types of nodes: Drug (D) and Protein (P). The graph contains three types of interaction (refer to three subgraphs):
🍪 D-D graph: drug-drug interactions with side effects as edge labels
🍰 P-D graph: protein-drug interactions (with a fixed label)
🍨 P-P graph: protein-protein interactions (with a fixed label)
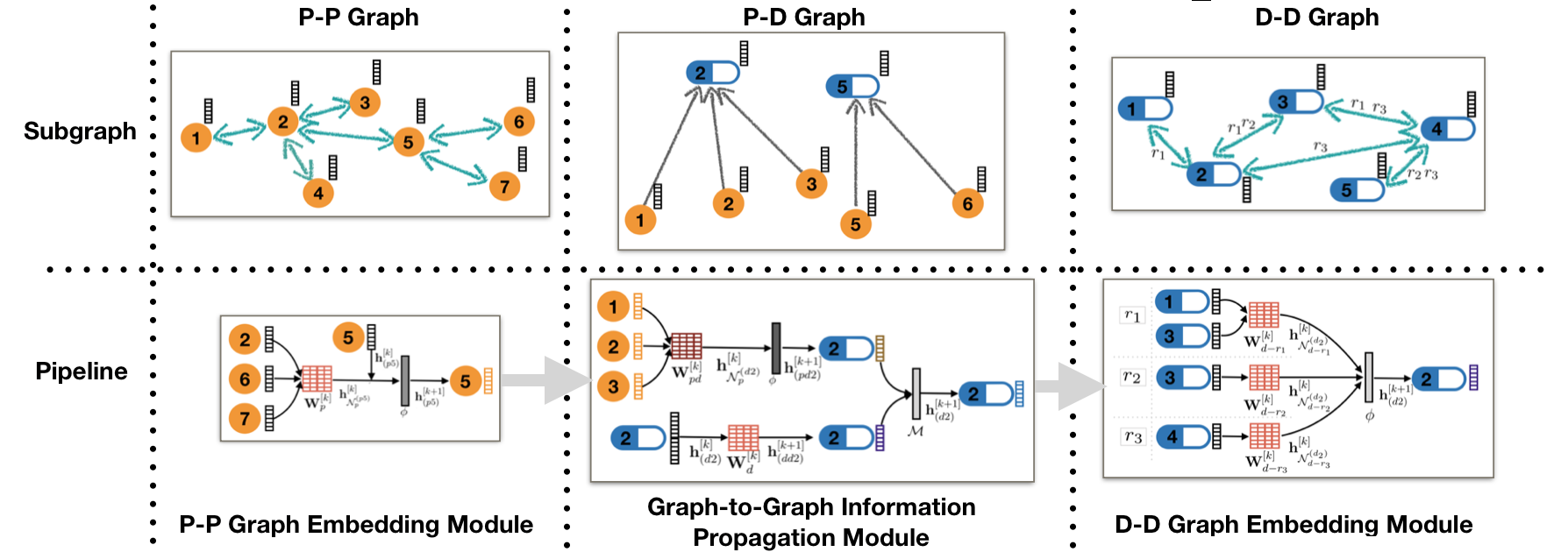
TIP model embeds proteins and drugs into different spaces of possibly different dimensions in the encoder, and predict side effects of drug combinations in the decoder. As shown below, TIP learns the protein embedding firstly on the P-P graph, and passes it to D-D graph via D-P graph. On D-D graph, TIP learns drug embedding and predicts relationships between drugs.
TIP Encoder:
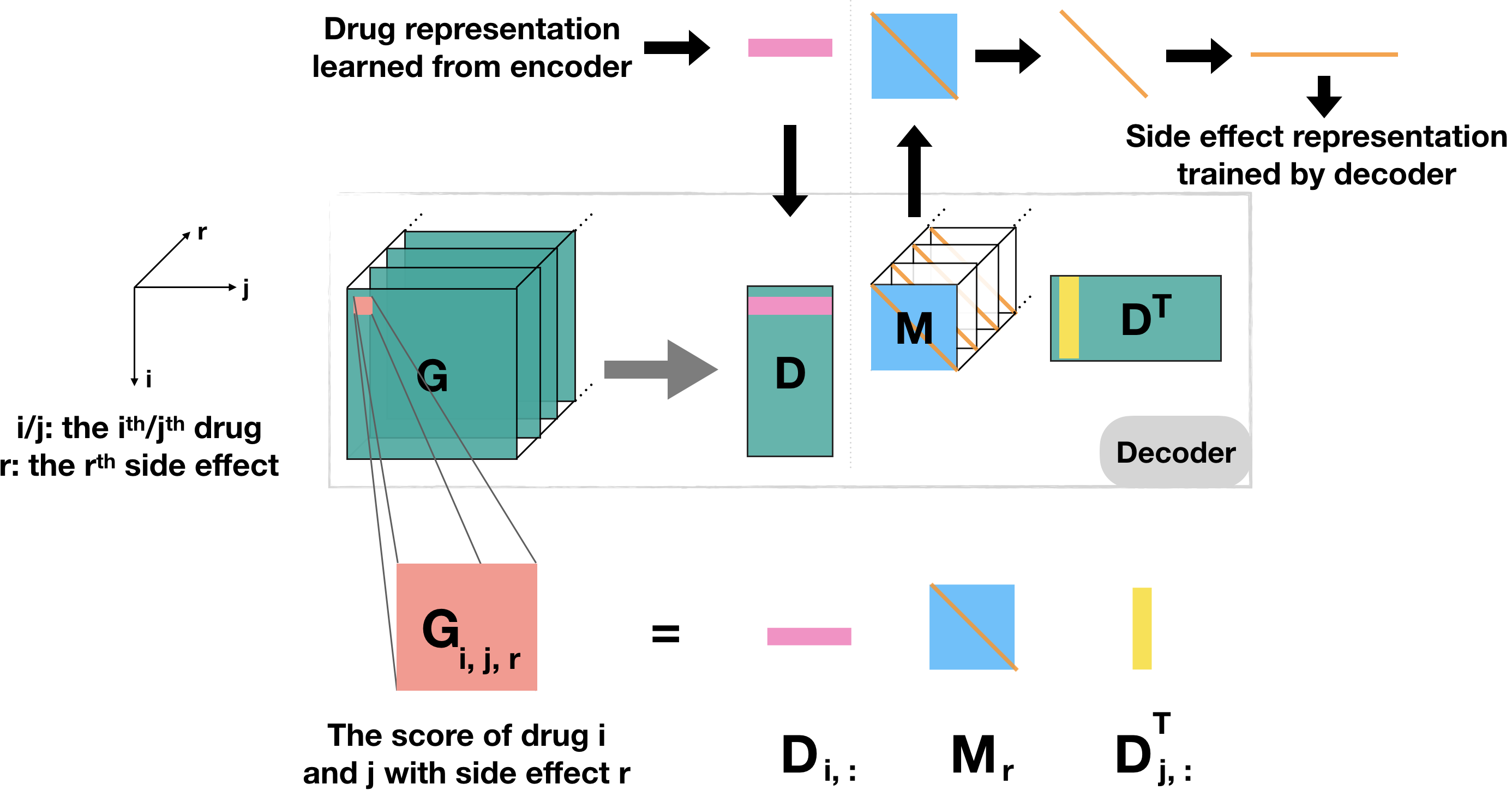
TIP Decoder:
TIP is implemented in PyTorch with PyTorch-Geometric package. It is developed and tested under Python 3.
TIP is trained and tested on GPU. Make sure the essential packages in requirements_gpu.txt has been installed in your environment properly. Use the following command to install all the required packages:
$ pip install -r requirements_gpu.txtIf you do want to run the code with CPU, install the packages in the requirements_cpu.txt. Then, comment or remove @profile before train() function in the python file you want to run.
##################################################
@profile # remove this for training on CPU
##################################################
def train():The processed data and the code for data processing are in the ./data/ folder. The raw datasets are available on the BioSNAP. See ./data.ipynb for the full polypharmacy datasets analysis.
The ./model/ folder contains two TIP implementation examples and four TIP variants. Details on experimental setup can be found in our paper (Xu, Sang, and Lu, 2019). You can run any of them as following:
$ python [model_name].pyBy default, it uses a subdataset with only five side effects. Comment or remove the following code to train and test with the full datasets. It appears at the head part of the file.
#########################################################################
et_list = et_list[:3] # remove this line for full dataset learning
#########################################################################🌚🌒🌓🌔 Please browse/open issues should you have any questions or ideas 🌖🌗🌘🌚
TIP is licensed under the MIT License.