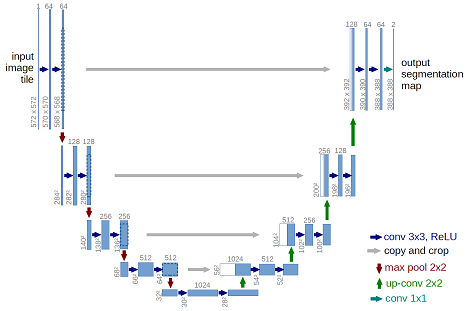
The architecture was inspired by U-Net: Convolutional Networks for Biomedical Image Segmentation.
The original dataset is from isbi challenge, and I've downloaded it and done the pre-processing.
The data for training contains 30 512*512 images, which are far not enough to feed a deep learning neural network. I use a module called ImageDataGenerator in keras.preprocessing.image to do data augmentation.
See dataPrepare.ipynb and data.py for detail.
This deep neural network is implemented with Keras functional API, which makes it extremely easy to experiment with different interesting architectures.
Output from the network is a 512*512 which represents mask that should be learned. Sigmoid activation function makes sure that mask pixels are in [0, 1] range.
The model is trained for 5 epochs.
After 5 epochs, calculated accuracy is about 0.97.
Loss function for the training is basically just a binary crossentropy.
- Tensorflow
- Keras >= 1.0
You will see the predicted results of test image in data/membrane/test