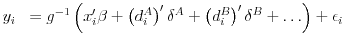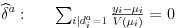The R package glmhdfe allows for the estimation of generalized linear models with high dimensional fixed effects. The package makes use of a convenient property of some combinations of error term distributions and link functions, where the fixed effects have — conditional on all other estimated parameters — an explicit solution.
Consider the following equation that we want to estimate:
The first order conditions for fixed effects can be simplified to
For certain distribution and link combinations this yields explicit solutions for the estimated coefficient for the fixed effects 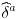 , given estimates for beta and the other deltas. Specifically, this is the case for the Gaussian distribution with identity and log link, and for the Poisson, Gamma and Inverse Gaussian distributions with log link. This makes it possible to update the fixed effects separately from the estimation of the coefficients on variables of interest in every iteration of the IRLS procedure used to estimate beta, dramatically increasing the speed of the estimation procedure.
, given estimates for beta and the other deltas. Specifically, this is the case for the Gaussian distribution with identity and log link, and for the Poisson, Gamma and Inverse Gaussian distributions with log link. This makes it possible to update the fixed effects separately from the estimation of the coefficients on variables of interest in every iteration of the IRLS procedure used to estimate beta, dramatically increasing the speed of the estimation procedure.
For more detail on the inner workings see the technical note. A Stata implementation is coming soon.
The R package glmhdfe implements this "trick" and utilizes the powers of the data.table package for a fast implementation. For smaller datasets or other error term distributions we recommend the feglm command in Amrei Stammann's alpaca package that also allows high-dimensional fixed effects in GLM estimations.
Install from Github via the remotes package:
remotes::install_github("julianhinz/R_glmhdfe")The glmhdfe function has a similar syntax as the felm function from the lfe package and the feglm function in the alpaca package:
glmhdfe(trade ~ fta | iso_o_year + iso_d_year + iso_o_iso_d | iso_o + iso_d + year,
family = poisson(link = "log"),
data = data)The first part of the formula is specified as usual. The second part of the formula specifies the fixed effects dimensions, the third part, which is optional, the clustering of the standard errors.
There are numerous options to tweak the estimation procedure:
formuladescribes dependent variable, right-hand side variables of interest, sets of fixed effects and clustering of standard errors, e.g. asy ~ x | fe1 + fe2 | cluster1 + cluster2dataspecifies thedata.tableordata.framewith data used in the regressionfamilyspecifies the estimator used, currently limited togaussian(link = "identity"),gaussian(link = "log"),poisson(link = "log"),Gamma(link = "log")betaallows to include a vector of starting values, although, interestingly, this does not tend to speed up the estimationtolerancespecifies the minimum change in the deviance at which the iteration breaksmax_iterationsspecifies the maximum number of iterationsacceleratespecifies whether to use an acceleration algorithm, still quite buggyaccelerate_iterationsspecifies the number of iterations before starting acceleration algorithmaccelerate_aux_vectorspecifies whether to include the estimated fixed effects vectors in IRLS, which, interestingly, increases convergence speedcompute_vcovasks whether to compute the variance-covariance matrix. It can also be computed ex-post when data from estimation is provideddemean_variablesif you don't want to compute the variance-covariance matrix right away, do you still want to demean variables to be used in estimation of variance-covariance matrix?demean_iterationsspecifies the number of iterations for the demeaningdemean_tolerancespecifies the minimum change in the diagonal of the Hessian at which the demeaning iteration breaksinclude_feasks whether the estimated fixed effects should be returnedinclude_dataasks whether the data used in the estimation should be returned, which may be useful if the variance-covariance matrix will be computed ex-postinclude_data_vcovreturn data used in variance-covariance matrix estimation?skip_checksspecifies, whether certain data integrity checks should be skipped before starting the procedure. Current option to skip are the detection of separation issues ("separation"), multicollinearity ("multicollinearity"), or missing data ("complete_cases")
traceasks whether to show some information during the estimationverboseasks whether to show a bit more information during estimation for the impatient
There is the usual battery of generic functions, like coef, summary, etc. Furthermore, if for some reason you want to (re-)estimate the variance-covariance matrix afterwards, or change the level of clustering, you can do so with the compute_vcov command:
compute_vcov(data, call, info)You need to specify the data (best in the form of a glmhdfe_data object), call (for information on clustering and variable of interest), and info (for information on degrees of freedom, etc.).
- try
evalon variables that are not part of data, e.g. for something likey ~ log(x) - Inverse Gaussian with log link
- tests using
testthat - parallelization in
Rcppwithomp - detect collinearity with
lmhdfeor otherdata.tableroutine, notfelmbecause this slows it down - run first IRLS with transformed lhs, otherwise beta guess only works for gaussian
- faster checks for separation issues, implement procedure recommended by Correia et al. (2019)
- This package is still in its early stages. Let us know if you find bugs or have suggestions for changes by opening an issue or e-mail to mail@julianhinz.com.