How Expressive are Transformers in Spectral Domain for Graphs?
This repository implements the approach outlined in the paper "How Expressive are Transformers in Spectral Domain for Graphs?". Note: This is the code that was submitted for the review. For purposes of keeping information anonymous the code(variables, function names, comments etc.) may not be very understandable. We are in the process of cleaning the code and will push the changes post this. Please write back in case of any queries.
Short Description about FeTA
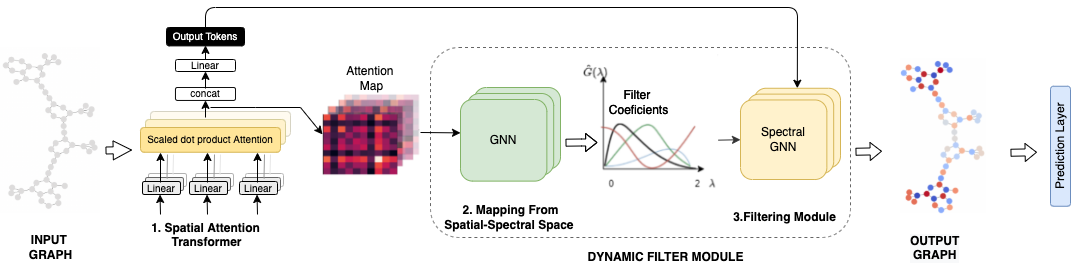
FeTA is a module designed for learning transformations on the spectral components of graph structured data.
Installation
Environment:
numpy=1.18.1
scipy=1.3.2
Cython=0.29.23
scikit-learn=0.22.1
matplotlib=3.4
networkx=2.5
python=3.7
pytorch=1.6
torch-geometric=1.7
rdkit-pypi=2021.03.3
dgl==0.6.1
To begin with, run:
cd FeTA
. s_env
To install GCKN, you also need to run:
make
You also need to create a cache folder to store computed positional encoding
mkdir -p cache/pe
Training FeTA
To train FeTA on MUTAG, run:
python -m experiments.run_transformer_gengcn_cv --gnn_type='ChebConvDynamic' --seed=0 --dataset='MUTAG'To train FeTA on NCI1, run:
python -m experiments.run_transformer_gckn_gengcn_cv --gnn_type='ChebConvDynamic' --seed=0 --dataset='NCI1'To train FeTA on ZINC, run:
python -m experiments.run_transformer_gckn_gengcn --gnn_type='ChebConvDynamic' --seed=0 --dataset='ZINC'To train FeTA on MolHIV, run:
python -m experiments.run_transformer_gengcn_molhiv --gnn_type='ChebConvDynamic' --batch-size=1024 --epochs=10To train FeTA on PATTERN, run:
python -m experiments.run_transformer_gengcn_SBM_cv --gnn_type='ChebConvDynamic' --batch-size=64 --epochs=100 --dataset='PATTERN'To train FeTA on CLUSTER, run:
python -m experiments.run_transformer_gengcn_SBM_cv --gnn_type='ChebConvDynamic' --batch-size=64 --epochs=100 --dataset='CLUSTER'To run the optimized configurations of FeTA (along with the position encodings) the scripts are provided in the folder scripts under the respective dataset.
Here --fold-idx can be varied from 1 to 10 to train on a specified training fold. To test a selected model, just add the --test (this will use complete train + val data) flag.
Configurations from GraphiT paper: To include Laplacian positional encoding into input node features, run:
python run_transformer_gengcn_cv.py --dataset NCI1 --gnn_type='ChebConvDynamic' --fold-idx 1 --pos-enc diffusion --beta 1.0 --lappe --lap-dim 8To include GCKN path features into input node features, run:
python run_transformer_gckn_gengcn_cv.py --dataset NCI1 --gnn_type='ChebConvDynamic' --fold-idx 1 --pos-enc diffusion --beta 1.0 --gckn-path 5Regression
To train FeTA on ZINC, run:
python run_transformer_gengcn.py --gnn_type='ChebConvDynamic' --pos-enc diffusion --beta 1.0To include Laplacian positional encoding into input node features, run:
python run_transformer_gengcn.py --gnn_type='ChebConvDynamic' --pos-enc diffusion --beta 1.0 --lappe --lap-dim 8To include GCKN path features into input node features, run:
python run_transformer_gckn_gengcn.py --gnn_type='ChebConvDynamic' --pos-enc diffusion --beta 1.0 --gckn-path 8Citation
If you use our work kindly consider citing
@article{
bastos2022how,
title={How Expressive are Transformers in Spectral Domain for Graphs?},
author={Anson Bastos and Abhishek Nadgeri and Kuldeep Singh and Hiroki Kanezashi and Toyotaro Suzumura and Isaiah Onando Mulang'},
journal={Transactions on Machine Learning Research},
year={2022},
url={https://openreview.net/forum?id=aRsLetumx1},
note={}
}
Acknowledgements
The code has been adapted from Graph Transformer, GraphiT and SAN repositories.