Benchmarking fastq compression with generic (mature) compression algorithms
This behcmark is motivated by a recent question from Ryan Connor on the µbioinfo Slack group
my impression is that bioinformatics really likes gzip (and only gzip?), but that there are other generic compression algs that are better (for bioinfo data types); assuming you agree (if not, why not?), why haven't the others compression types caught on in bioinformatics?
It kicked off an interesting discussion, which led me to dig into the literature and see what I could find. I'm sure I could search deeper and for longer, but I really couldn't find any benchmarks that satisfied me. Don't get me wrong, there are plenty of benchmarks, but they're always looking at bioinformatics-specific tools for compressing sequencing data. Sure, these perform well, but every repository I went to was untouched in a while. When archiving data, the last thing I want is to try and decompress my data and the tool no longer installs/works on my system. In addition, I want the tool to be ubiquitous and mature. I know this is a lot of constraints, but hey, that's what I am interested in.
This benchmark only covers ubiquitous/mature/generic compression tools
The tools tested in this benchmark are:
These tools were used as they were the main ones that popped up in our discussion. Feel free to raise an issue on this repository if you would like to see another tool included.
All compression level settings were tested for each tool and default settings were used for all other options.
The data used to test each tool are fastqs:
- ERR3152364: Metagenome from https://doi.org/10.1093/gigascience/giz043
- ERR9030439 - Mycobacterium tuberculosis from https://doi.org/10.1016/S2666-5247(22)00301-9
- SRR11038964 - Escherichia coli from https://doi.org/10.1016/j.envpol.2020.115081
- SRR12695179 - Listeria monocytogenes from https://doi.org/10.1128/MRA.01159-20
- SRR24283715 - Salmonella enterica
- ERR10367342 - Klebsiella pneumoniae from https://doi.org/10.1099/mgen.0.000132
- SRR24015950 - S. enterica
- SRR15422103 - Seawater metagenome from https://doi.org/10.1016/j.watres.2022.119282
- SRR13044184 - Mus musculus
- SRR22859778 - Staphylococcus aureus from https://doi.org/10.1101/2023.03.28.534496
- ERR2935805 - Metagenome from https://doi.org/10.1093/gigascience/giz043
- ERR9030317 - M. tuberculosis from https://doi.org/10.1016/S2666-5247(22)00301-9
- SRR11038976 - E. coli from https://doi.org/10.1016/j.envpol.2020.115081
- SRR12695183 - L. monocytogenes from https://doi.org/10.1128/MRA.01159-20
- SRR24283718 - S. enterica
- ERR1023775 - K. pneumoniae from https://doi.org/10.1099/mgen.0.000132
- SRR24015952 - S. enterica
- SRR22859722 - S. aureus from https://doi.org/10.1101/2023.03.28.534496
- SRR098024 - Homo sapiens
- SRR077288 - Maylandia zebra from https://doi.org/10.1038/nature13726
Note: I couldn't find sources for all of these samples. If you can fill in some of the gaps, please raise an issue and I will gladly update the sources.
All data were downloaded with fastq-dl (v2.0.1). Paired Illumina data were
combined into a single fastq file.
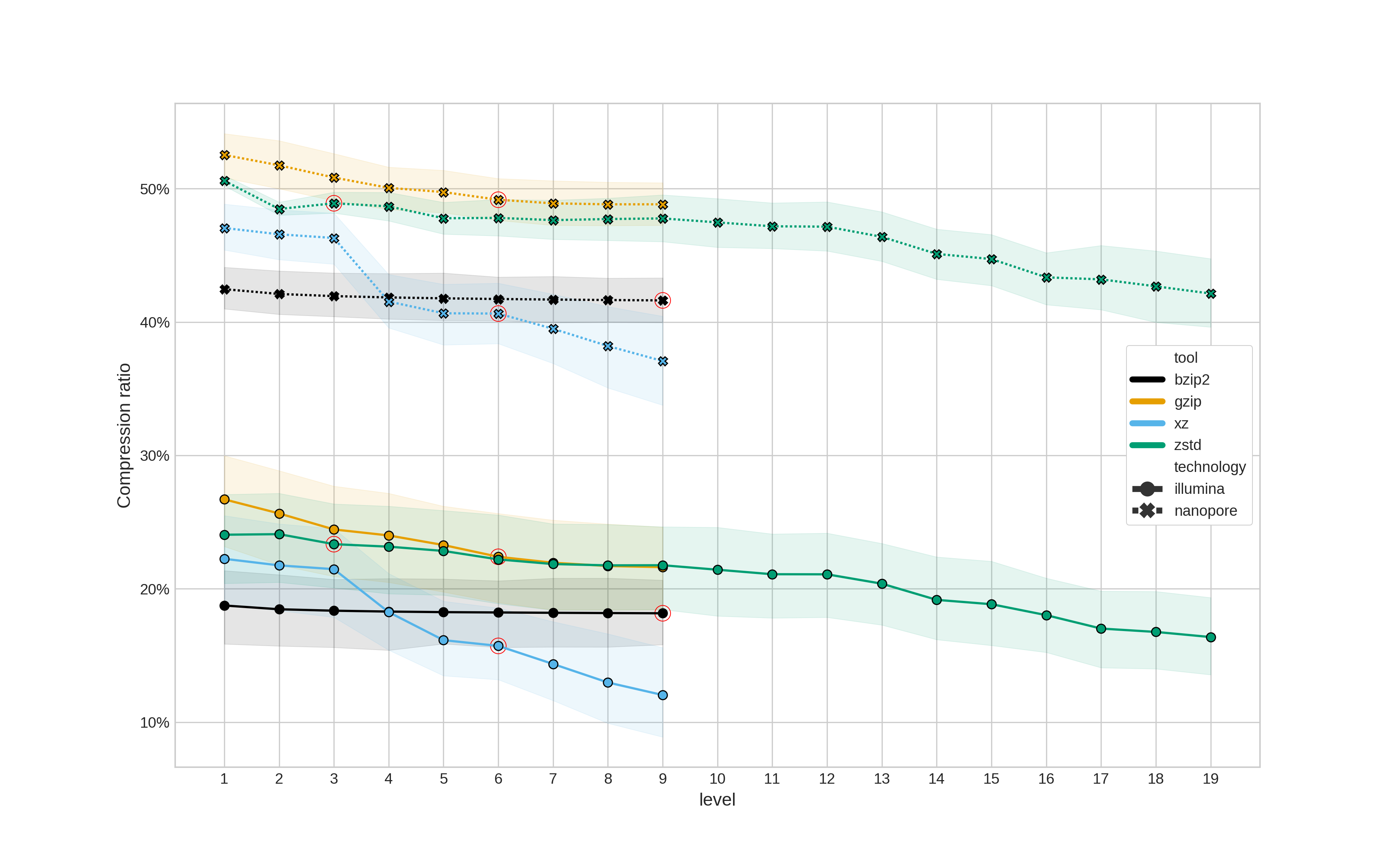
The first question is how much smaller does each compression tool make a fastq file. As this also depends on the compression level selected, all possible levels were tested for each tool (the default being indicated with a red circle).
The compression ratio is a percentage of the original file size - i.e.,
Figure 1: Compression ratio (y-axis) for different compression tools and levels. Compression ratio is a percentage of the original file size. The red circles indicate the default compression level for each tool. Illumina data is represented with a solid line and circular points, whereas Nanopore data is a dashed line with cross points. Translucent error bands represent the 95% confidence interval.
The most striking result here is the noticeable different in compression ratio between Illumina and Nanopore data - regardless of the compression tool used. (If anyone can suggest a reason for this, please raise an issue.)
Update 07/06/2023: Peter Menzel mentioned this is likely due to the noisier quality scores in the Nanopore data. Illumina quality scores are generally quite homogenous, which increases compressability.
Using default settings, zstd and gzip provide similar ratios, as do xz
and bzip2 (however, compression level doesn't seem to actually change the ratio
for bzip2). When using the highest compression level xz provides the best
compression (however, this comes at a cost to runtime as we'll see below).
In many scenarios, the (de)compression rate is just as important as the compression ratio. However, if compressing for archival purposes, rate is probably not as important.
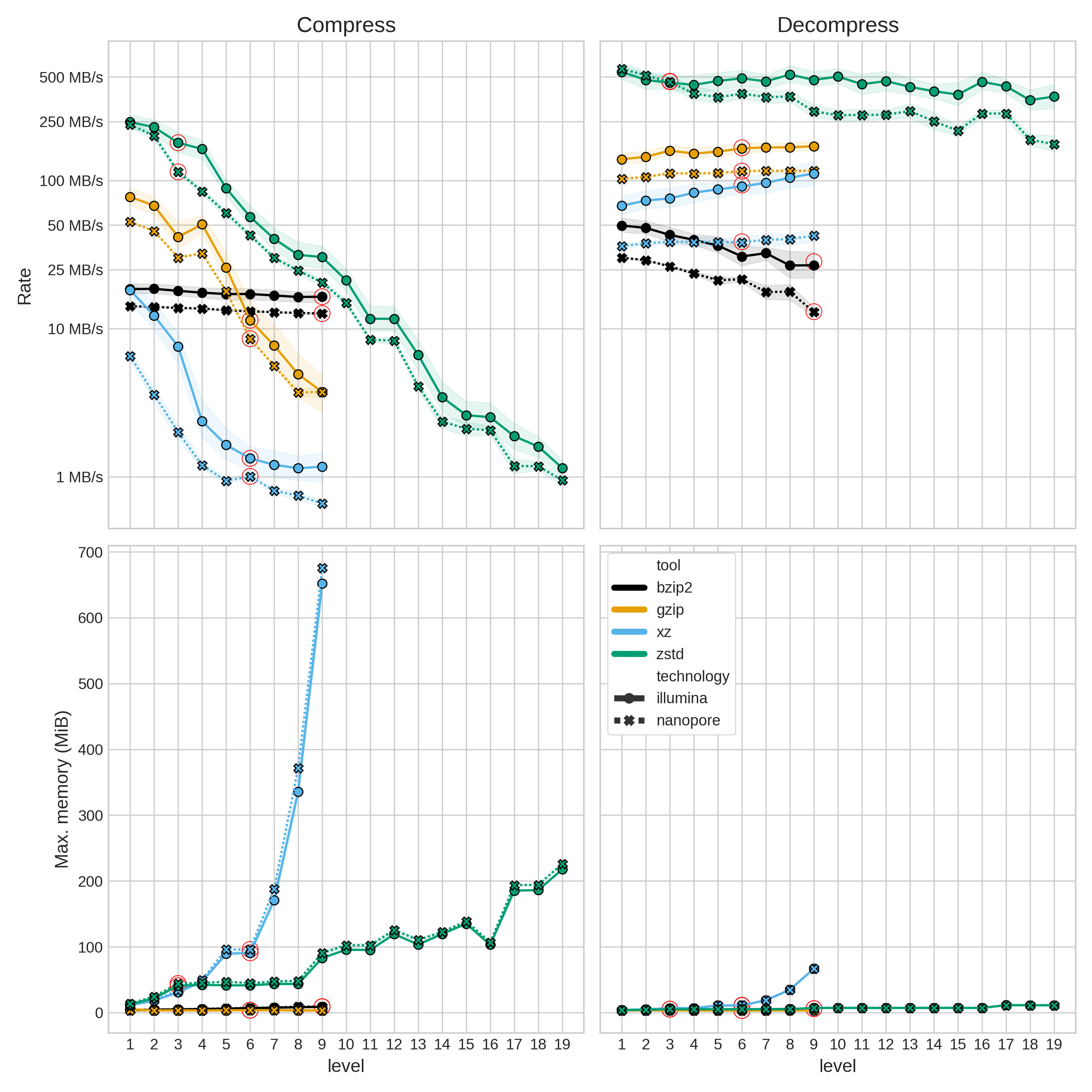
The compression rate is
Figure 2: Compression (left column) and decompression (right column) rate (top row) and peak memory usage (lower row). Note the log scale for rate. The red circles indicate the default compression level for each tool. Illumina data is represented with a solid line and circular points, whereas Nanopore data is a dashed line with cross points. Translucent error bands represent the 95% confidence interval.
As alluded to earlier, xz pays for its fantastic compression ratios by being
orders-of-magnitude slower than the other tools. It also uses a lot more memory than the
other tools when (de)compressing - although in absolute terms, the highest memory usage
is still well below 1GB.
The main take-away from Figure 2 is that zstd (de)compresses much faster than the
other tools (using the default level). Compression level seems to have a big impact in
compression rate (except for bzip2), however, not so much for decompression.
So what tool to use? As most often with benchmarks: it depends on your situation.
If all you care about is compressing your data as small as it will go ,and you don't
mind how long it takes, then xz - with compression level 9 - is the obvious choice.
If you want fast (de)compression, then zstd is the best option - using default
options.
If, like most people, you're contemplating replacing gzip (default options), the
siutation is a little
less clear. As a drop-in replacement, zstd (default options) will give you about the
same compression
ratio with ~10-fold faster compression and ~3-5-fold faster decompression. Another
option is bzip2, which will give you ~1.2-fold better compression ratios than gzip (
and zstd) with a comparable compression rate to gzip. However, bzip2's
decompression rate is ~5-fold slower than gzip.
One final consideration is APIs for various programming languages. If it is difficult to
read/write files that are compressed with a given algorithm, then using that compression
type might cause problems. Most (good) bioinformatics tools support gzip-compressed
input and output. However, support for other compression types shouldn't be too much
work for most software tool developers provided a well-maintained and documented API is
available in the relevant programming language. Here is a list of APIs for the tested
compression tools in a selection of programming languages with an arbitrary grading
system for how "stable" I think they are (feel free to put in a pull request if you want
to contribute other languages).
| gzip | bzip2 | xz | zstd | |
|---|---|---|---|---|
| Python | A | A | A | B+ |
| Rust | A | B+ | B+ | B |
| C/C++ | A | A | A | A |
- A: standard library (i.e. builtin) or library is maintained by the original
developer (note: Rust's
gziplibrary is maintained by rust-lang itself) - B: external library that is actively maintained, well-documented, and has quick response times