wakefield is designed to quickly generate random data sets. The user
passes n (number of rows) and predefined vectors to the r_data_frame
function to produce a dplyr::tbl_df object.
To download the development version of wakefield:
Download the zip
ball or tar
ball, decompress
and run R CMD INSTALL on it, or use the pacman package to install
the development version:
if (!require("pacman")) install.packages("pacman")
pacman::p_load_gh("trinker/wakefield")
pacman::p_load(dplyr, tidyr, ggplot2)
You are welcome to:
- submit suggestions and bug-reports at: https://github.com/trinker/wakefield/issues
- send a pull request on: https://github.com/trinker/wakefield/
- compose a friendly e-mail to: tyler.rinker@gmail.com
The r_data_frame function (random data frame) takes n (the number of
rows) and any number of variables (columns). These columns are typically
produced from a wakefield variable function. Each of these variable
functions has a pre-set behavior that produces a named vector of n
length, allowing the user to lazily pass unnamed functions (optionally,
without call parenthesis). The column name is hidden as a varname
attribute. For example here we see the race variable function:
race(n=10)
## [1] Black White Asian White White Hispanic White
## [8] White Hispanic Native
## Levels: White Hispanic Black Asian Bi-Racial Native Other Hawaiian
attributes(race(n=10))
## $levels
## [1] "White" "Hispanic" "Black" "Asian" "Bi-Racial" "Native"
## [7] "Other" "Hawaiian"
##
## $class
## [1] "variable" "factor"
##
## $varname
## [1] "Race"
When this variable is used inside of r_data_frame the varname is
used as a column name. Additionally, the n argument is not set within
variable functions but is set once in r_data_frame:
r_data_frame(
n = 500,
race
)
## # A tibble: 500 × 1
## Race
## <fctr>
## 1 White
## 2 White
## 3 Black
## 4 White
## 5 White
## 6 White
## 7 Hispanic
## 8 White
## 9 White
## 10 White
## # ... with 490 more rows
The power of r_data_frame is apparent when we use many modular
variable functions:
r_data_frame(
n = 500,
id,
race,
age,
sex,
hour,
iq,
height,
died
)
## # A tibble: 500 × 8
## ID Race Age Sex Hour IQ Height Died
## <chr> <fctr> <int> <fctr> <S3: times> <dbl> <dbl> <lgl>
## 1 001 White 26 Male 00:00:00 98 72 TRUE
## 2 002 White 30 Male 00:00:00 94 74 FALSE
## 3 003 White 22 Female 00:00:00 99 69 TRUE
## 4 004 White 26 Male 00:00:00 103 66 TRUE
## 5 005 White 26 Female 00:00:00 102 71 TRUE
## 6 006 White 26 Female 00:00:00 97 71 FALSE
## 7 007 White 29 Female 00:30:00 94 65 TRUE
## 8 008 Black 22 Male 00:30:00 105 75 TRUE
## 9 009 White 33 Female 00:30:00 105 65 FALSE
## 10 010 White 32 Male 00:30:00 102 65 FALSE
## # ... with 490 more rows
There are 49 wakefield based variable functions to chose from,
spanning R's various data types (see ?variables for details).
| age | dice | hair | military | sex_inclusive |
| animal | dna | height | month | smokes |
| answer | dob | income | name | speed |
| area | dummy | internet_browser | normal | state |
| car | education | iq | political | string |
| children | employment | language | race | upper |
| coin | eye | level | religion | valid |
| color | grade | likert | sat | year |
| date_stamp | grade_level | lorem_ipsum | sentence | zip_code |
| death | group | marital | sex |
Available Variable Functions
However, the user may also pass their own vector producing functions or
vectors to r_data_frame. Those with an n argument can be set by
r_data_frame:
r_data_frame(
n = 500,
id,
Scoring = rnorm,
Smoker = valid,
race,
age,
sex,
hour,
iq,
height,
died
)
## # A tibble: 500 × 10
## ID Scoring Smoker Race Age Sex Hour IQ Height
## <chr> <dbl> <lgl> <fctr> <int> <fctr> <S3: times> <dbl> <dbl>
## 1 001 -1.00172866 TRUE White 32 Male 00:00:00 107 74
## 2 002 -1.22045688 FALSE Hispanic 34 Female 00:00:00 112 69
## 3 003 0.03800246 TRUE White 28 Female 00:00:00 84 65
## 4 004 0.71036400 TRUE White 21 Female 00:00:00 85 65
## 5 005 0.59644996 TRUE White 22 Female 00:00:00 96 69
## 6 006 0.24556053 TRUE White 23 Male 00:00:00 123 79
## 7 007 1.59434567 FALSE White 30 Female 00:00:00 101 65
## 8 008 0.14108265 TRUE White 31 Male 00:00:00 88 71
## 9 009 -0.96799173 TRUE White 29 Male 00:00:00 98 68
## 10 010 -0.06773925 FALSE Hispanic 22 Female 00:30:00 89 70
## # ... with 490 more rows, and 1 more variables: Died <lgl>
r_data_frame(
n = 500,
id,
age, age, age,
grade, grade, grade
)
## # A tibble: 500 × 7
## ID Age_1 Age_2 Age_3 Grade_1 Grade_2 Grade_3
## <chr> <int> <int> <int> <dbl> <dbl> <dbl>
## 1 001 28 26 24 96.1 78.9 83.7
## 2 002 30 22 27 95.4 86.9 89.4
## 3 003 35 27 22 91.8 86.9 89.4
## 4 004 25 27 33 81.7 87.1 91.1
## 5 005 33 35 33 86.7 87.2 82.4
## 6 006 29 20 34 91.1 86.3 92.1
## 7 007 26 29 33 86.2 87.8 81.3
## 8 008 34 33 31 84.5 81.6 89.8
## 9 009 21 33 27 96.0 88.6 94.0
## 10 010 31 29 26 91.4 89.0 87.8
## # ... with 490 more rows
While passing variable functions to r_data_frame without call
parenthesis is handy, the user may wish to set arguments. This can be
done through call parenthesis as we do with data.frame or
dplyr::data_frame:
r_data_frame(
n = 500,
id,
Scoring = rnorm,
Smoker = valid,
`Reading(mins)` = rpois(lambda=20),
race,
age(x = 8:14),
sex,
hour,
iq,
height(mean=50, sd = 10),
died
)
## # A tibble: 500 × 11
## ID Scoring Smoker `Reading(mins)` Race Age Sex
## <chr> <dbl> <lgl> <int> <fctr> <int> <fctr>
## 1 001 1.25315699 FALSE 18 White 9 Male
## 2 002 -0.10451919 FALSE 21 White 8 Female
## 3 003 -0.11401295 TRUE 19 White 14 Female
## 4 004 0.77380822 FALSE 16 White 9 Male
## 5 005 0.36936803 FALSE 18 Hispanic 13 Female
## 6 006 0.72023857 TRUE 24 White 13 Male
## 7 007 0.16074250 FALSE 17 White 10 Male
## 8 008 -0.03576366 FALSE 18 Black 11 Female
## 9 009 0.15264881 TRUE 28 White 10 Male
## 10 010 -0.22782276 FALSE 23 White 10 Female
## # ... with 490 more rows, and 4 more variables: Hour <S3: times>,
## # IQ <dbl>, Height <dbl>, Died <lgl>
Often data contains missing values. wakefield allows the user to add
a proportion of missing values per column/vector via the r_na (random
NA). This works nicely within a dplyr/magrittr %>% then
pipeline:
r_data_frame(
n = 30,
id,
race,
age,
sex,
hour,
iq,
height,
died,
Scoring = rnorm,
Smoker = valid
) %>%
r_na(prob=.4)
## # A tibble: 30 × 10
## ID Race Age Sex Hour IQ Height Died Scoring
## <chr> <fctr> <int> <fctr> <S3: times> <dbl> <dbl> <lgl> <dbl>
## 1 01 White NA Female <NA> 108 NA NA -1.2889925
## 2 02 White NA Male <NA> NA 66 NA NA
## 3 03 White 22 Female 01:00:00 103 76 FALSE NA
## 4 04 White NA NA 01:30:00 94 NA TRUE -1.7087044
## 5 05 Hispanic NA Female 04:30:00 NA 74 NA -0.1776321
## 6 06 NA 27 Female <NA> 101 77 NA 1.0338205
## 7 07 White 34 NA 05:30:00 NA NA TRUE 0.9559290
## 8 08 NA 27 Male <NA> 104 NA FALSE 1.3647700
## 9 09 NA 30 Female 06:30:00 95 66 NA -0.1283762
## 10 10 White 25 NA 06:30:00 104 NA NA -1.3232695
## # ... with 20 more rows, and 1 more variables: Smoker <lgl>
The r_series function allows the user to pass a single wakefield
function and dictate how many columns (j) to produce.
set.seed(10)
r_series(likert, j = 3, n=10)
## # A tibble: 10 × 3
## Likert_1 Likert_2 Likert_3
## * <ord> <ord> <ord>
## 1 Neutral Disagree Strongly Disagree
## 2 Agree Neutral Disagree
## 3 Neutral Strongly Agree Disagree
## 4 Disagree Neutral Agree
## 5 Strongly Agree Agree Neutral
## 6 Agree Neutral Disagree
## 7 Agree Strongly Agree Strongly Disagree
## 8 Agree Agree Agree
## 9 Disagree Agree Disagree
## 10 Neutral Strongly Disagree Agree
Often the user wants a numeric score for Likert type columns and similar
variables. For series with multiple factors the as_integer converts
all columns to integer values. Additionally, we may want to specify
column name prefixes. This can be accomplished via the variable
function's name argument. Both of these features are demonstrated
here.
set.seed(10)
as_integer(r_series(likert, j = 5, n=10, name = "Item"))
## # A tibble: 10 × 5
## Item_1 Item_2 Item_3 Item_4 Item_5
## * <int> <int> <int> <int> <int>
## 1 3 2 1 3 4
## 2 4 3 2 5 4
## 3 3 5 2 5 5
## 4 2 3 4 1 2
## 5 5 4 3 3 4
## 6 4 3 2 2 5
## 7 4 5 1 1 5
## 8 4 4 4 1 3
## 9 2 4 2 2 5
## 10 3 1 4 3 1
r_series can be used within a r_data_frame as well.
set.seed(10)
r_data_frame(n=100,
id,
age,
sex,
r_series(likert, 3, name = "Question")
)
## # A tibble: 100 × 6
## ID Age Sex Question_1 Question_2
## * <chr> <int> <fctr> <ord> <ord>
## 1 001 28 Male Agree Agree
## 2 002 24 Male Neutral Strongly Agree
## 3 003 26 Male Disagree Neutral
## 4 004 31 Male Strongly Disagree Neutral
## 5 005 21 Female Strongly Agree Strongly Disagree
## 6 006 23 Female Disagree Disagree
## 7 007 24 Female Disagree Strongly Agree
## 8 008 24 Male Strongly Disagree Agree
## 9 009 29 Female Agree Strongly Agree
## 10 010 26 Male Strongly Disagree Strongly Disagree
## # ... with 90 more rows, and 1 more variables: Question_3 <ord>
set.seed(10)
r_data_frame(n=100,
id,
age,
sex,
r_series(likert, 5, name = "Item", integer = TRUE)
)
## # A tibble: 100 × 8
## ID Age Sex Item_1 Item_2 Item_3 Item_4 Item_5
## * <chr> <int> <fctr> <int> <int> <int> <int> <int>
## 1 001 28 Male 4 4 1 1 1
## 2 002 24 Male 3 5 2 1 2
## 3 003 26 Male 2 3 2 1 2
## 4 004 31 Male 1 3 2 4 3
## 5 005 21 Female 5 1 1 5 4
## 6 006 23 Female 2 2 4 3 4
## 7 007 24 Female 2 5 1 5 2
## 8 008 24 Male 1 4 4 5 5
## 9 009 29 Female 4 5 5 4 3
## 10 010 26 Male 1 1 4 1 2
## # ... with 90 more rows
The user can also create related series via the relate argument in
r_series. It allows the user to specify the relationship between
columns. relate may be a named list of or a short hand string of the
form of "fM_sd" where:
fis one of (+, -, *, /)Mis a mean valuesdis a standard deviation of the mean value
For example you may use relate = "*4_1". If relate = NULL no
relationship is generated between columns. I will use the short hand
string form here.
r_series(grade, j = 5, n = 100, relate = "+1_6")
## # A tibble: 100 × 5
## Grade_1 Grade_2 Grade_3 Grade_4 Grade_5
## * <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 84.5 92.5 91.6 87.4 76.7
## 2 93.1 85.0 81.8 87.8 91.3
## 3 81.6 67.5 52.6 48.8 56.8
## 4 92.5 89.3 95.3 102.2 94.5
## 5 96.6 95.9 98.7 115.9 114.7
## 6 89.7 88.1 88.8 89.0 86.4
## 7 92.8 91.7 98.3 98.7 101.6
## 8 92.1 92.9 92.6 85.5 93.1
## 9 90.6 96.9 103.9 107.6 106.2
## 10 96.0 94.8 84.3 91.1 106.6
## # ... with 90 more rows
r_series(age, 5, 100, relate = "+5_0")
## # A tibble: 100 × 5
## Age_1 Age_2 Age_3 Age_4 Age_5
## * <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 24 29 34 39 44
## 2 24 29 34 39 44
## 3 27 32 37 42 47
## 4 22 27 32 37 42
## 5 32 37 42 47 52
## 6 27 32 37 42 47
## 7 21 26 31 36 41
## 8 29 34 39 44 49
## 9 35 40 45 50 55
## 10 33 38 43 48 53
## # ... with 90 more rows
r_series(likert, 5, 100, name ="Item", relate = "-.5_.1")
## # A tibble: 100 × 5
## Item_1 Item_2 Item_3 Item_4 Item_5
## * <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 2 1 0 -1 -1
## 2 3 2 1 1 0
## 3 1 1 1 0 0
## 4 4 3 3 2 1
## 5 2 1 1 0 0
## 6 2 1 1 1 0
## 7 1 0 0 -1 -2
## 8 2 2 1 1 0
## 9 2 2 1 0 0
## 10 3 3 3 3 3
## # ... with 90 more rows
r_series(grade, j = 5, n = 100, relate = "*1.05_.1")
## # A tibble: 100 × 5
## Grade_1 Grade_2 Grade_3 Grade_4 Grade_5
## * <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 85.7 94.27 113.124 113.1240 113.1240
## 2 86.4 77.76 77.760 85.5360 85.5360
## 3 90.6 99.66 89.694 98.6634 108.5297
## 4 89.1 89.10 89.100 71.2800 71.2800
## 5 87.0 95.70 114.840 103.3560 113.6916
## 6 93.9 103.29 123.948 136.3428 136.3428
## 7 80.1 72.09 64.881 84.3453 84.3453
## 8 91.7 110.04 132.048 132.0480 145.2528
## 9 87.4 96.14 96.140 105.7540 116.3294
## 10 92.9 92.90 83.610 91.9710 101.1681
## # ... with 90 more rows
Use the sd command to adjust correlations.
round(cor(r_series(grade, 8, 10, relate = "+1_2")), 2)
## Grade_1 Grade_2 Grade_3 Grade_4 Grade_5 Grade_6 Grade_7 Grade_8
## Grade_1 1.00 0.85 0.64 0.39 0.28 0.25 0.28 0.15
## Grade_2 0.85 1.00 0.86 0.68 0.61 0.56 0.56 0.47
## Grade_3 0.64 0.86 1.00 0.77 0.70 0.80 0.86 0.78
## Grade_4 0.39 0.68 0.77 1.00 0.94 0.80 0.65 0.74
## Grade_5 0.28 0.61 0.70 0.94 1.00 0.85 0.69 0.73
## Grade_6 0.25 0.56 0.80 0.80 0.85 1.00 0.92 0.89
## Grade_7 0.28 0.56 0.86 0.65 0.69 0.92 1.00 0.91
## Grade_8 0.15 0.47 0.78 0.74 0.73 0.89 0.91 1.00
round(cor(r_series(grade, 8, 10, relate = "+1_0")), 2)
## Grade_1 Grade_2 Grade_3 Grade_4 Grade_5 Grade_6 Grade_7 Grade_8
## Grade_1 1 1 1 1 1 1 1 1
## Grade_2 1 1 1 1 1 1 1 1
## Grade_3 1 1 1 1 1 1 1 1
## Grade_4 1 1 1 1 1 1 1 1
## Grade_5 1 1 1 1 1 1 1 1
## Grade_6 1 1 1 1 1 1 1 1
## Grade_7 1 1 1 1 1 1 1 1
## Grade_8 1 1 1 1 1 1 1 1
round(cor(r_series(grade, 8, 10, relate = "+1_20")), 2)
## Grade_1 Grade_2 Grade_3 Grade_4 Grade_5 Grade_6 Grade_7 Grade_8
## Grade_1 1.00 0.26 0.27 0.40 0.21 -0.21 -0.36 -0.41
## Grade_2 0.26 1.00 0.77 0.60 0.64 0.50 0.53 0.46
## Grade_3 0.27 0.77 1.00 0.78 0.76 0.66 0.62 0.66
## Grade_4 0.40 0.60 0.78 1.00 0.95 0.76 0.59 0.55
## Grade_5 0.21 0.64 0.76 0.95 1.00 0.82 0.65 0.61
## Grade_6 -0.21 0.50 0.66 0.76 0.82 1.00 0.90 0.82
## Grade_7 -0.36 0.53 0.62 0.59 0.65 0.90 1.00 0.94
## Grade_8 -0.41 0.46 0.66 0.55 0.61 0.82 0.94 1.00
round(cor(r_series(grade, 8, 10, relate = "+15_20")), 2)
## Grade_1 Grade_2 Grade_3 Grade_4 Grade_5 Grade_6 Grade_7 Grade_8
## Grade_1 1.00 -0.10 -0.50 -0.39 -0.25 -0.52 -0.26 -0.31
## Grade_2 -0.10 1.00 0.74 0.50 0.13 0.03 0.36 0.46
## Grade_3 -0.50 0.74 1.00 0.81 0.48 0.41 0.71 0.78
## Grade_4 -0.39 0.50 0.81 1.00 0.75 0.66 0.58 0.75
## Grade_5 -0.25 0.13 0.48 0.75 1.00 0.91 0.70 0.74
## Grade_6 -0.52 0.03 0.41 0.66 0.91 1.00 0.58 0.57
## Grade_7 -0.26 0.36 0.71 0.58 0.70 0.58 1.00 0.78
## Grade_8 -0.31 0.46 0.78 0.75 0.74 0.57 0.78 1.00
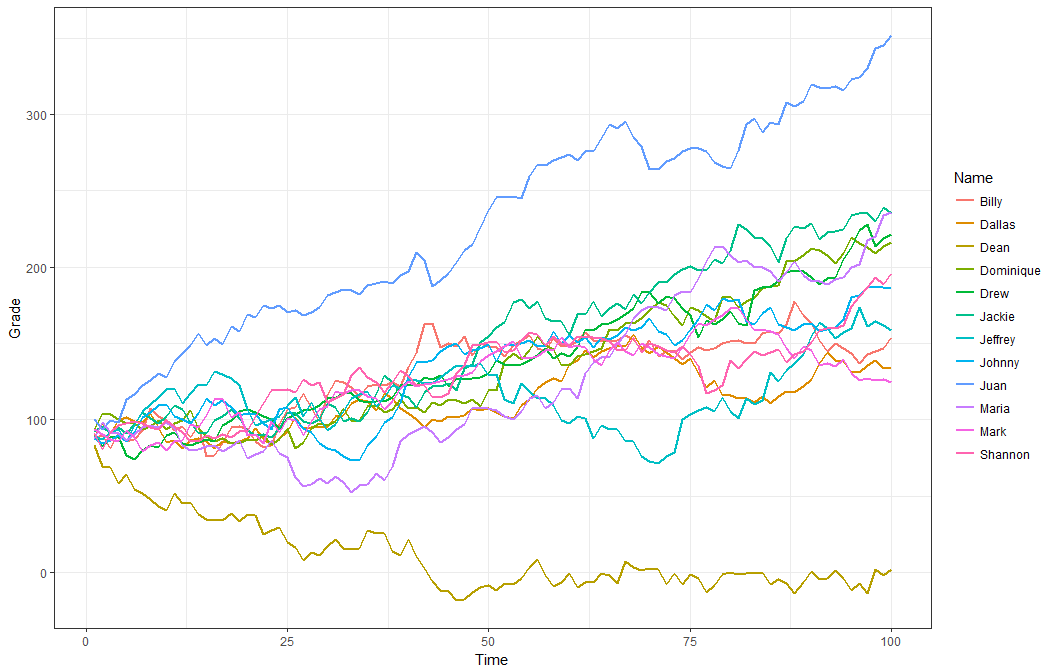
dat <- r_data_frame(12,
name,
r_series(grade, 100, relate = "+1_6")
)
dat %>%
gather(Time, Grade, -c(Name)) %>%
mutate(Time = as.numeric(gsub("\\D", "", Time))) %>%
ggplot(aes(x = Time, y = Grade, color = Name, group = Name)) +
geom_line(size=.8) +
theme_bw()
The user may wish to expand a factor into j dummy coded columns. The
r_dummy function expands a factor into j columns and works similar
to the r_series function. The user may wish to use the original factor
name as the prefix to the j columns. Setting prefix = TRUE within
r_dummy accomplishes this.
set.seed(10)
r_data_frame(n=100,
id,
age,
r_dummy(sex, prefix = TRUE),
r_dummy(political)
)
## # A tibble: 100 × 9
## ID Age Sex_Male Sex_Female Democrat Republican Constitution
## * <chr> <int> <int> <int> <int> <int> <int>
## 1 001 28 1 0 1 0 0
## 2 002 24 1 0 1 0 0
## 3 003 26 1 0 0 1 0
## 4 004 31 1 0 0 1 0
## 5 005 21 0 1 1 0 0
## 6 006 23 0 1 0 1 0
## 7 007 24 0 1 0 1 0
## 8 008 24 1 0 0 0 0
## 9 009 29 0 1 1 0 0
## 10 010 26 1 0 0 1 0
## # ... with 90 more rows, and 2 more variables: Libertarian <int>,
## # Green <int>
It is helpful to see the column types and NAs as a visualization. The
table_heat (also the plot method assigned to tbl_df as well) can
provide visual glimpse of data types and missing cells.
set.seed(10)
r_data_frame(n=100,
id,
dob,
animal,
grade, grade,
death,
dummy,
grade_letter,
gender,
paragraph,
sentence
) %>%
r_na() %>%
plot(palette = "Set1")