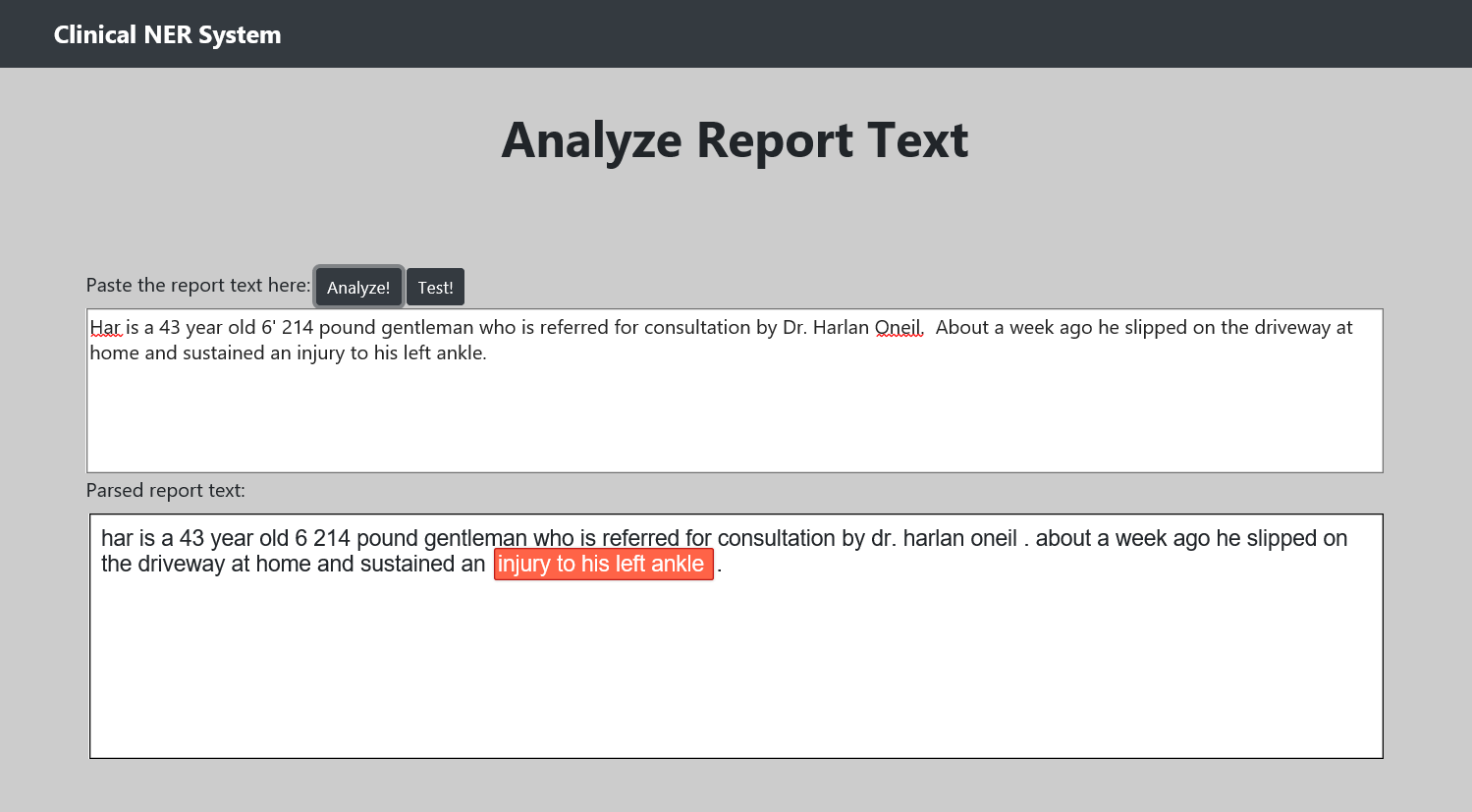
The purpose of this project is to identify the Named Entity Recognition in clinical reports. I used the Unified Medical Language System (UMLS) database for the baseline for disease and symptoms names. The data used in this project is "n2c2 NLP Research Data Sets". For UMLS and n2c2 dataset, we need proper license from the respective sites.
Notebooks are not committed to this repo. due to the n2c2 dataset PHI & policies.
| Name | Github Page | Personal Website |
|---|---|---|
| Asha Ponraj | asha | www.devskrol.com |
- Conditional Random Fields from sklearn_crfsuite
- Machine Learning techniques
- NLP tasks
- Predictive Modeling
- Manual Data Annotation
- Feature Engineering and Extraction
- etc.
- Python
- sklearn_crfsuite
- MySql
- Pandas, jupyter
- HTML
- etc.
- Increase accuracy
- CRF implementation
- Negation detection
- Differentiate Symptoms, Disease names, Exam names.
This file structure is based on the DSSG machine learning pipeline.