- Introduction
- Dirichlet Energy and Laplacian for (Directed) Graphs
- Fractional Graph Laplacian
- Fractional Graph ODE
- Experiments
Let
- the adjacency matrix
$\mathbf{A}\in{0, 1}^{N\times N}$ such that$a_{i, j}=1$ if there is an edge from node$j$ to node$i$ ; - the in-degree matrix
$\mathbf{D}_\text{in} \coloneqq \mathrm{diag}(\mathbf{A}\mathbf{1})$ ; - the out-degree matrix
$\mathbf{D}_\text{out} \coloneqq \mathrm{diag}(\mathbf{A}^\top\mathbf{1})$ ; - the nodes' feature matrix
$\mathbf{x}\in\mathbb{R}^{N\times K}$ .
We define the Dirichlet Energy for directed graphs as
We define the symmetrically normalized adjacency (SNA) as
$$\mathbf{L}\coloneqq \mathbf{D}\text{in}^{-\frac{1}{2}} \mathbf{A} \mathbf{D}\text{out}^{-\frac{1}{2}}.$$
In the paper, we show theoretically that both definitions are extensions of the usual definitions for the undirected case. More specifically,
- the spectrum of the SNA lies in the unit circle, i.e.,
$\lvert\lambda(\mathbf{L})\rvert \leq 1$ ; - the Dirichlet Energy and the SNA are related by
$\mathfrak{E}(\mathbf{x})=\frac{1}{2}\Re\left(\mathrm{trace}\left(\mathbf{x}^\text{H} (\mathbf{I}-\mathbf{L}) \mathbf{x}\right)\right)$
Unlike the undirected case,
We define the
In the paper, we prove that the
We consider the fractional Laplacian ode
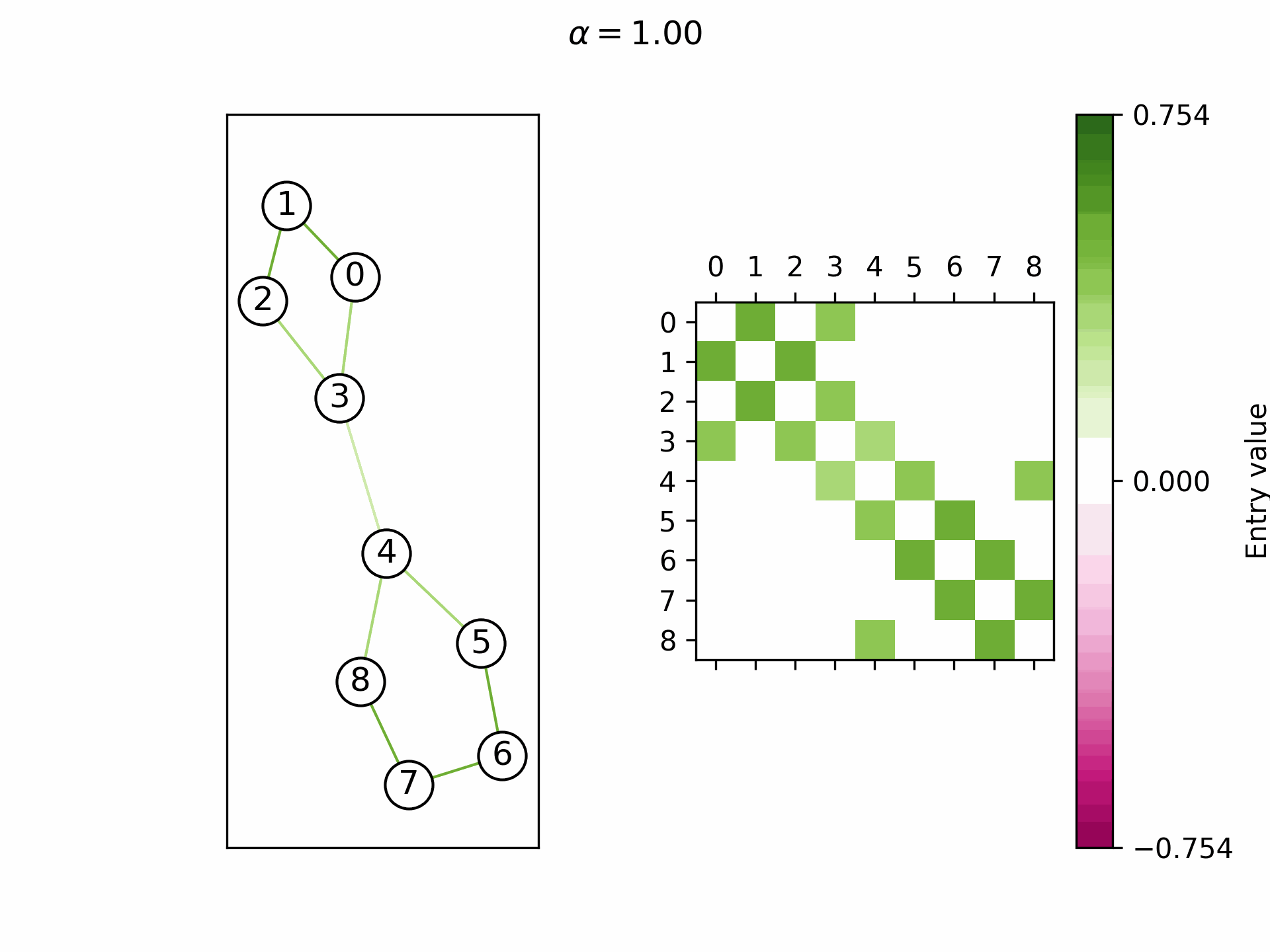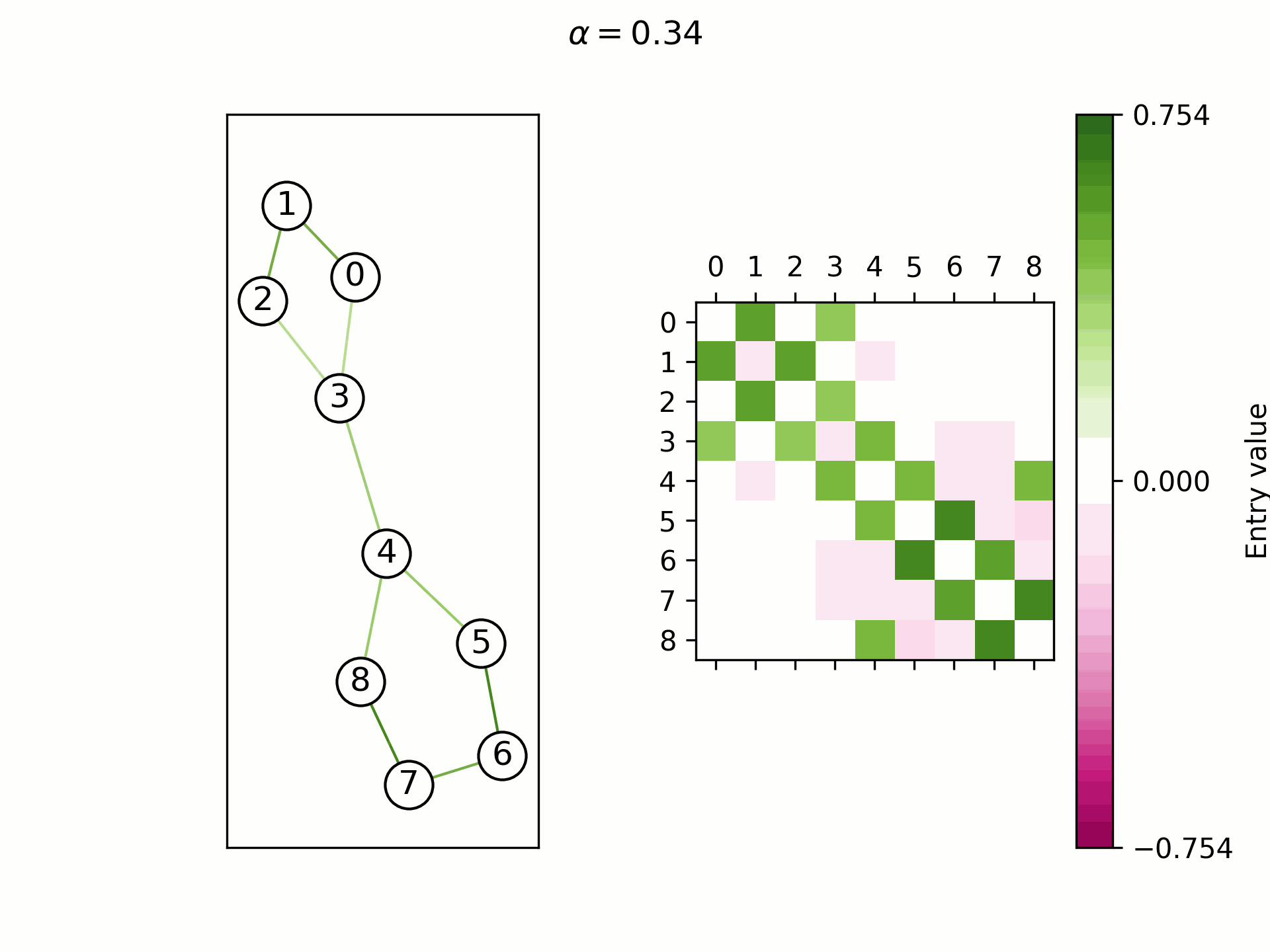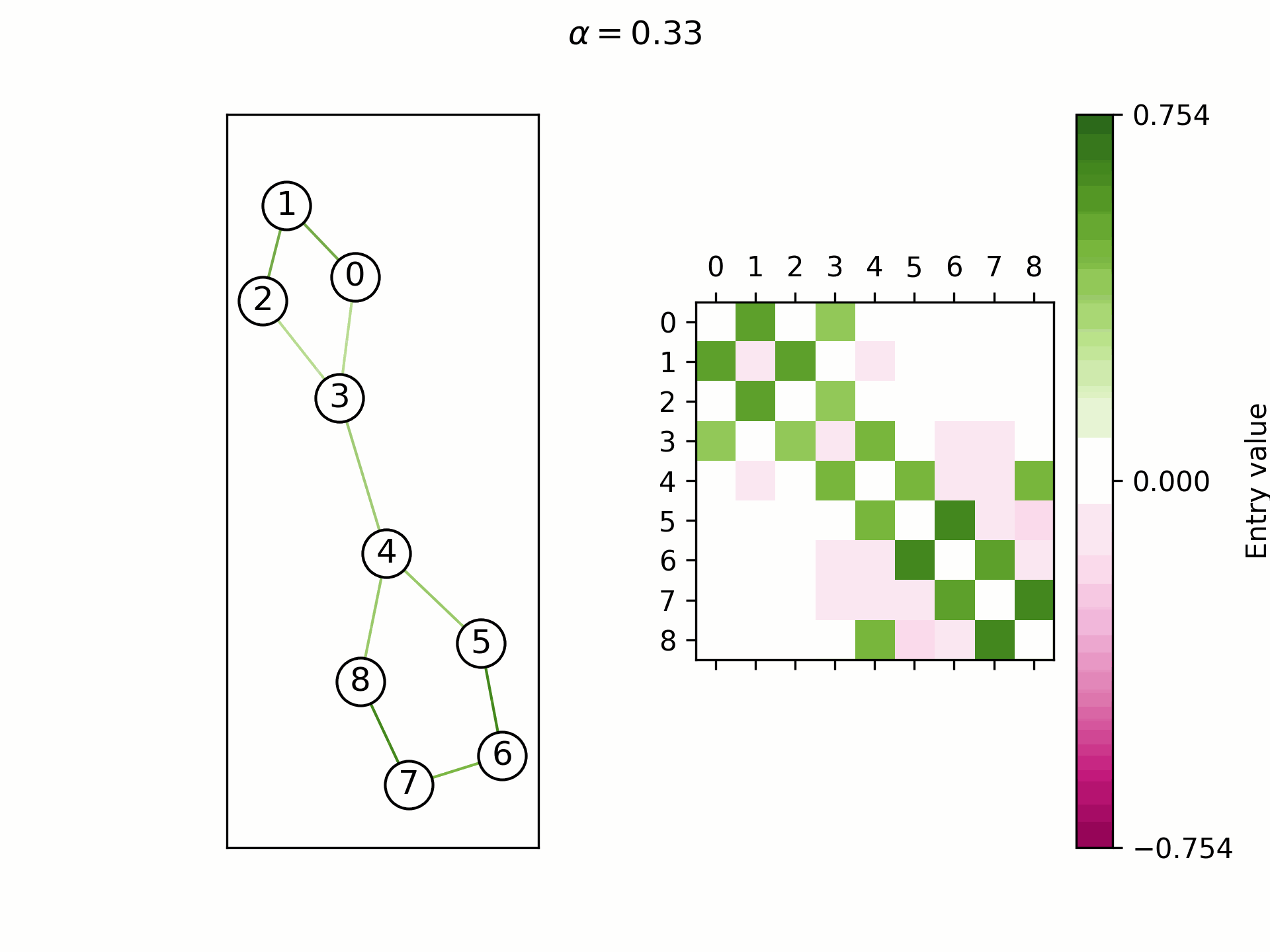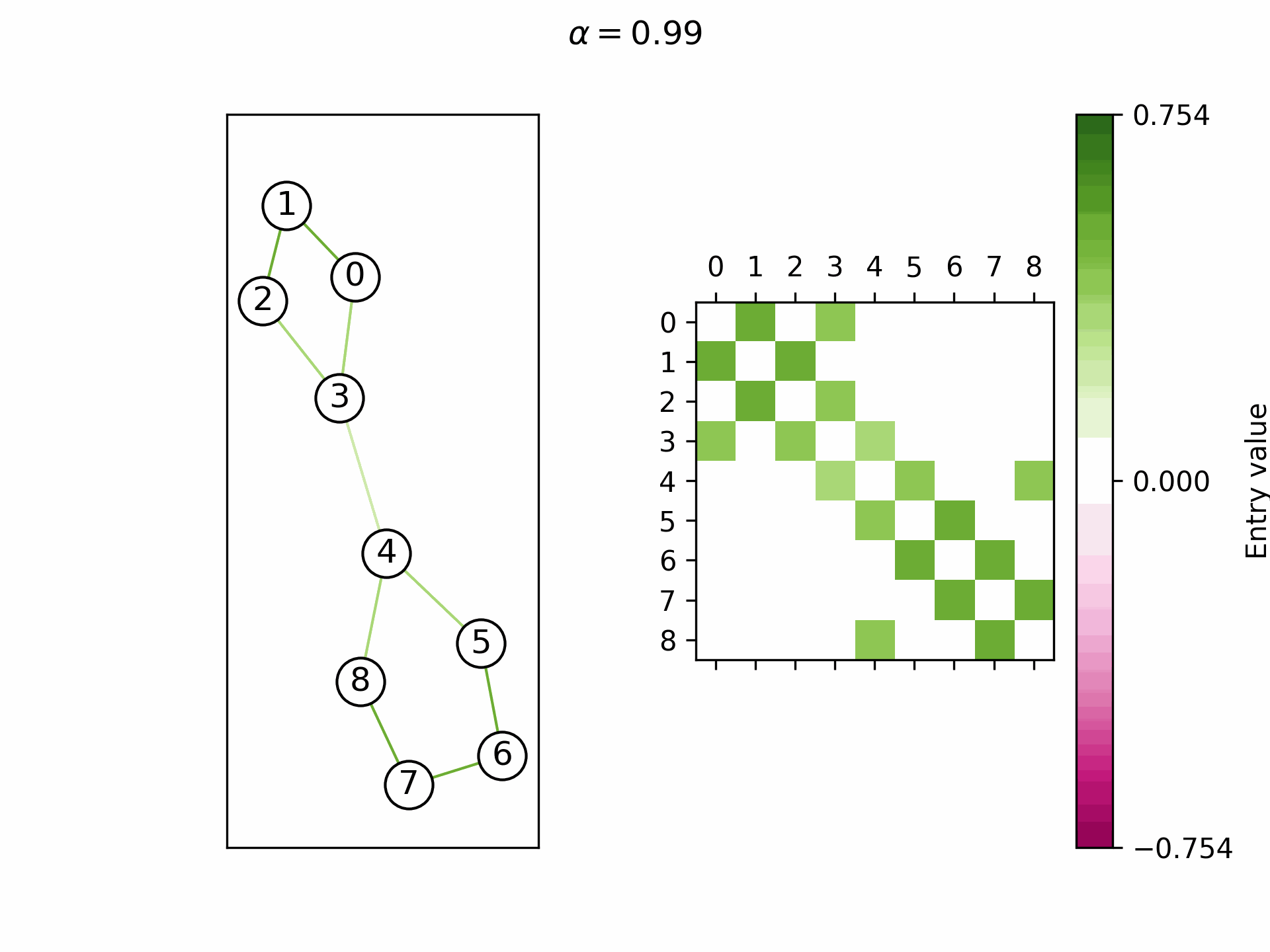
We theoretically show that by selecting a learnable
Real-world graphs are not purely homophilic (bottom right) nor purely heterophilic (bottom left), but lie somewhere in between (bottom center). Hence, the ability to adapt the convergence speed and the limit frequency
Clone the repository:
git clone https://github.com/RPaolino/fLode.git
Please check the dependencies and the required packages, or create a new environment from the environment.yml file
conda env create -f environment.yml
conda activate flode
To run the experiments, for example, on chameleon, type:
python node_classification.py --dataset chameleon
If you want to use the best hyperparams we found, you can use the flag -b: this will overwrite the default values with the values saved in lib.best. To see which argument will be overwritten, please check the dataset in lib.best.
python node_classification.py --dataset chameleon -b
You can specify your own configuration via command line. For a complete list of all arguments and their explanation, type:
python node_classification.py -h
An overview of the results is shown below. In the paper, one can find a comparison with other models. Note that for "Minesweeper", "Tolokers" and "Questions" the evaluation metric is AUC-ROC, while for the other datasets the evaluation metric is accuracy.
| film | squirrel | chameleon | Citeseer | Pubmed | Cora | Minesweeper | Tolokers | Roman-empire | Questions | |
|---|---|---|---|---|---|---|---|---|---|---|
| Undirected | 37.16 ± 1.42 | 64.23 ± 1.84 | 73.60 ± 1.55 | 78.07 ± 1.62 | 89.02 ± 0.38 | 86.44 ± 1.17 | 92.43 ± 0.51 | 84.17 ± 0.58 | 74.97 ± 0.53 | 78.39 ± 1.22 |
| Directed | 37.41 ± 1.06 | 74.03 ± 1.58 | 77.98 ± 1.05 | - | - | - | - | - | - | - |
In order to give a rough idea of the computational time, we report some statistics. The GPU is a NVIDIA TITANRTX with 24 GB of memory. Moreover, for Pubmed, Roman-empire and Questions we compute only 30% of the singular values due to memory (and time) limitations.
| film | squirrel | chameleon | Citeseer | Pubmed | Cora | Minesweeper | Tolokers | Roman-empire | Questions | |
|---|---|---|---|---|---|---|---|---|---|---|
| #Nodes | 7,600 | 5,201 | 2,277 | 3,327 | 18,717 | 2,708 | 10,000 | 11,758 | 22,662 | 48,921 |
| #Edges | 26,752 | 198,493 | 31,421 | 4,676 | 44,327 | 5,278 | 39,402 | 519,000 | 32,927 | 153,540 |
| SVD [mm:ss] | 02:55 | 01:30 | 00:03 | 00:03 | 07:46 | 00:04 | 04:22 | 12:53 | 10:26 | 26:15 |
| Training [iters/sec] | 5 | 4 | 10 | 8 | 4 | 15 | <1 | <1 | <1 | <1 |